Introduction to ENSO
What is ENSO ?
enso is a utility program to create and refine Conformer, Rotamer-Ensembles and finally prepare all files necessary to generate NMR spectra using the anmr program. enso and additional necessary resources can be found at Github ENSO and Github Cefine.
The Workflow to get the final spectrum is detailed in the following graphic.
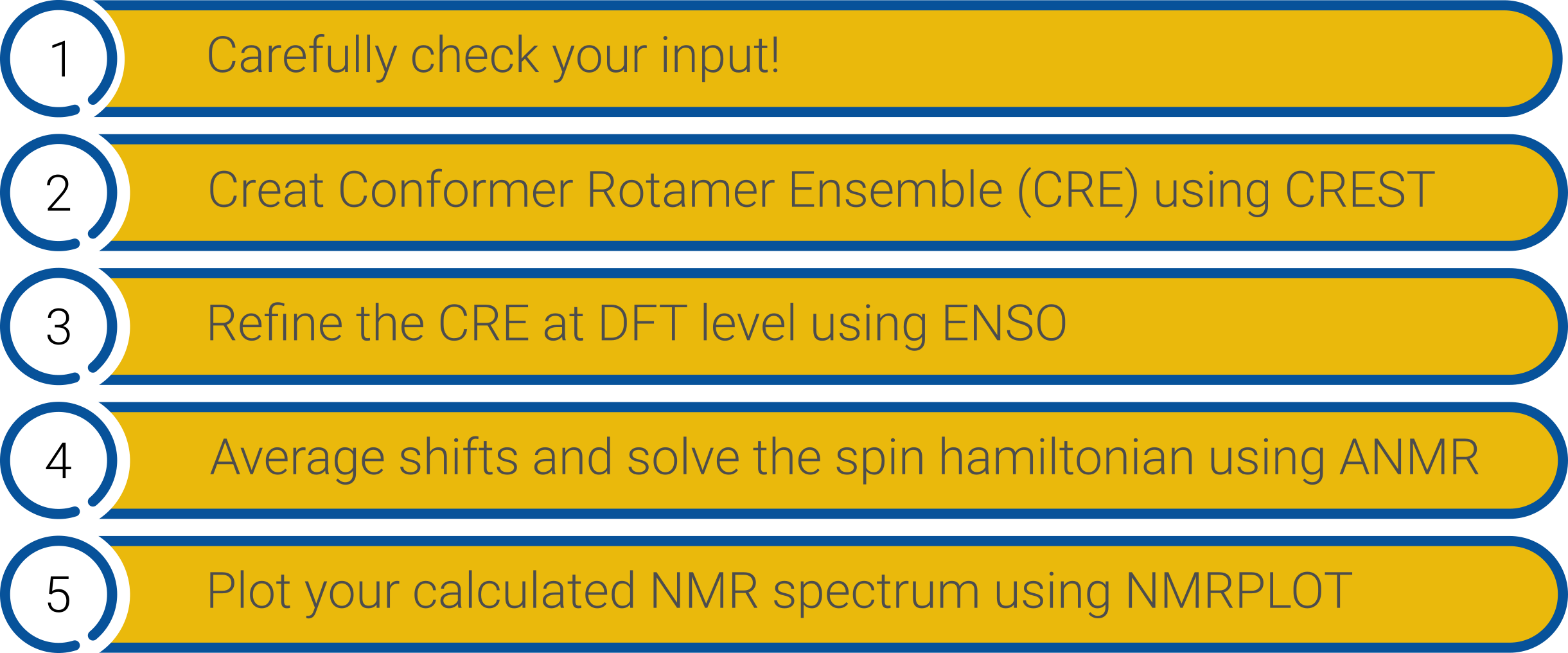

NMR spectrum calculation workflow.
Short summary
The enso program is designed to rerank the Conformer Rotamer Ensemble (CRE) generated by CREST (Conformer Rotamer Ensemble Search Tool) at DFT level and perform subsequent NMR-property calculations. enso interfaces to different QM-CODES (TURBOMOLE, ORCA, xtb) and processes the data. The ENSO is subdivided into four parts.
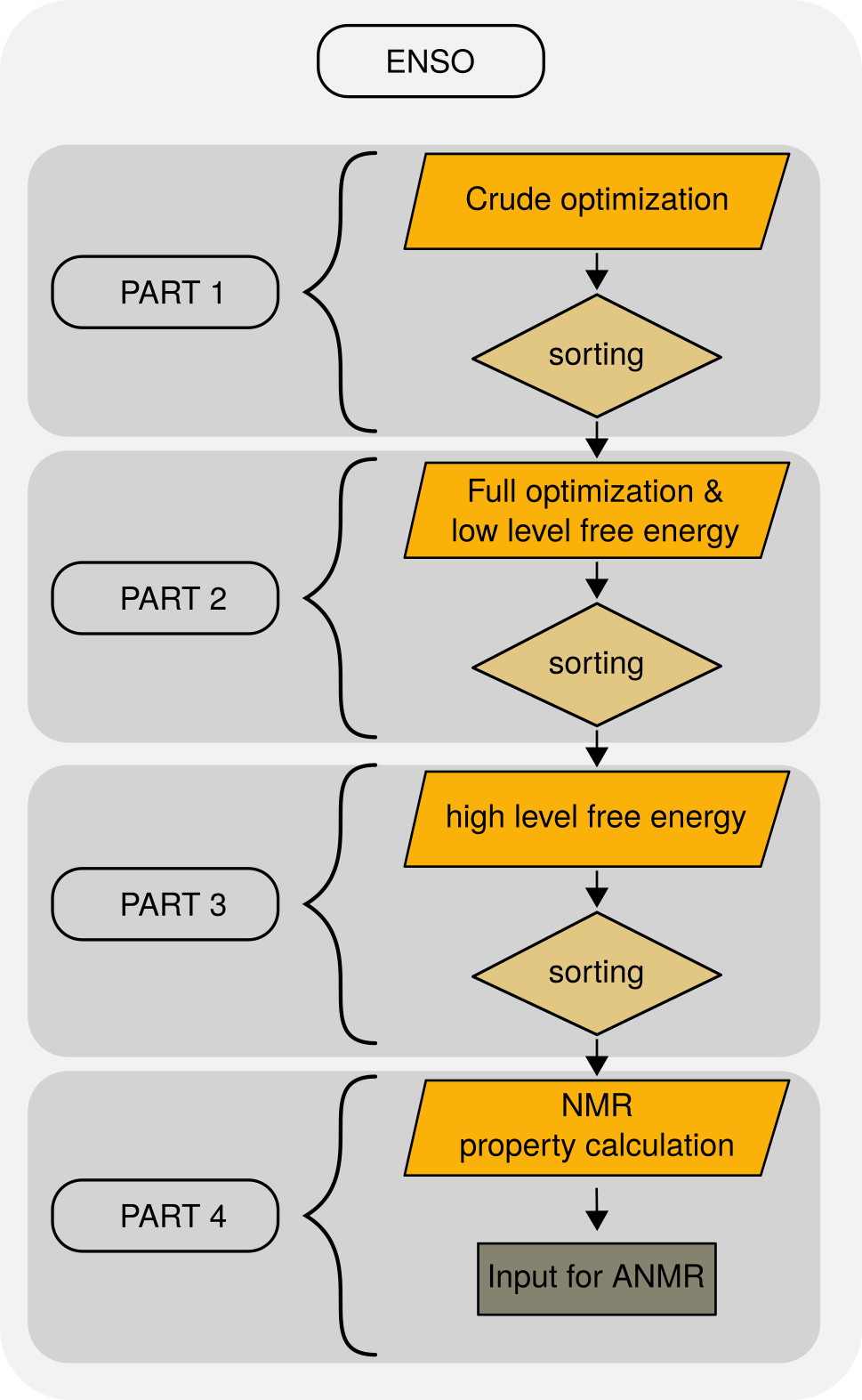
ENSO flowchart.
In part1 crude optimizations are performed at DFT level. Because of crude thresholds only a very small number of optimization cycles are perfomed at DFT level, and the upside is that bondlength are more or less correct. All geometries with a corresponding DFT energy below an energy thresold (thr-part1) are considered for the full optimization in part2.
In part2 the structures are fully optimized at DFT level. On these optimized geometries solvation contributions and thermostatistical contributions to free energy are calculated. The free energy is obtained as the sum of the ‘low-level’ DFT energy \(E\), \(G_{RRHO}\) (thermostatistical contribution to free energy) and \(δG_{solv}\) (optional solvation contribution to free energy). Using a second threshold (this time a free energy based threshold) only conformers with a free energy below the threshold are considered for the final free energy evaluation in part3.
In part3 the ‘high-level’ free energy calculations are performed by calculating high level DFT single-point energies and recalculating the free energy. At this stage Boltzmann weights for the remaining conformers are calculated. All conformers with a Boltzmann contribution below 1% are neglected and the Bolzmann weights are recalculated. The Boltzmann weights are summed up to 98% and all conformers within these 98% are considered for the NMR property calculations.
In part4 the couplings and shieldings calculations are performed. Additionally all information which is required by anmr is written by the enso.
After refining the CREST-ensemble at DFT level and performing the NMR property calculations, anmr correctly averages the shifts and solves the spin hamiltonian for the final spectrum generation. Then anmr generates a file ‘anmr.dat’ which can be plotted to visualize the final spectrum.
Getting started
To perform an ENSO calculation first run:
./enso.py [-options]
This generates a ‘flags.dat’ configuration file which stores most information relevant to ENSO.
vim flags.dat # adjust and save the settings in the configuration file to your needs.
After adjusting the settings in the ‘flags.dat’, the input can be automatically checked (for typos and setting-combinations) by enso. This way settings or combinations of settings which are not possible are detected before you actually start the program. The check is performed with:
./enso.py --checkinput
When the input is correct the program can be started by typing:
export PYTHONUNBUFFERED=1
./enso.py --run > enso.out 2> enso.error &
To save the output of enso it is necessary to pipe the output into a file.
Using this export PYTHONUNBUFFERED=1 the progress of the calculations can be
tracked immediatly and the user doesn’t need to wait until the python buffer is
written to the file. For the program to work correctly the information of the
absolute paths to the programs employed in ENSO has to be available in the
global configuration file .ensorc.