Usage examples
CENSO can be used for several applications / target quantities. Some are listed below:
Hint
CENSO has sorting “parts” which can be turned on and off. The parts are run in sequence and the sorting-results are influenced by the choice of sorting-parts employed. If for example the optimization part (part2) is not performed, then all subsequent parts will calculate free energies or properties on the input SQM/FF geometries (not DFT optimized geometries)! Each part contains thresholds (see Threshold) and choosing to low (free) energy windows (in the early sorting parts) will affect your final ensemble / averaged free energy / property.
Note
For the demonstration purpose it is assumed that all parts are turned off in the global configuration file of the user!
Calculate fast DFT(B97-D3(0)/def2-SV(P)+gcp) single-point energies on GFNn-xTB input geometries
Hint
Useful in case of large structure ensembles (SE). Very efficient (fast) improvement on the electronic energy description compared to the initial SQM/FF energies. High lying conformers are quickly sorted out.
$ censo -inp ensemble.xyz -part0 on -chrg 1 -solvent h2o > censo.out &
______________________________________________________________
| |
| |
| CENSO - Commandline ENSO |
| v 1.0.3 |
| energetic sorting of CREST Conformer Rotamer Ensembles |
| University of Bonn, MCTC |
| Feb 2021 |
| based on ENSO version 2.0.1 |
| F. Bohle and S. Grimme |
| |
|______________________________________________________________|
This program is distributed in the hope that it will be useful,
but WITHOUT ANY WARRANTY; without even the implied warranty of
MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE.
----------------------------------------------------------------------------------------------------
PARAMETERS
----------------------------------------------------------------------------------------------------
The configuration file .censorc is read from /home/bohle/1projects/from_tmp1/CENSO/documentation-calcs/1-part0/.censorc.
Reading conformer rotamer ensemble from: /home/bohle/1projects/from_tmp1/CENSO/documentation-calcs/1-part0/ensemble.xyz.
Reading file: censo_solvents.json
--------------------------------------------------
CRE SORTING SETTINGS
--------------------------------------------------
number of atoms in system: 25
number of considered conformers: 22
number of all conformers from input: 22
charge: 1
unpaired: 0
solvent: h2o
temperature: 298.15
evaluate at different temperatures: on
temperature range: 273.15, 278.15, 283.15, 288.15, ...
calculate mRRHO contribution: on
consider symmetry for mRRHO contribution: off
cautious checking for error and failed calculations: on
checking the DFT-ensemble using CREST: off
maxthreads: 2
omp: 2
--------------------------------------------------
CRE CHEAP-PRESCREENING - PART0
--------------------------------------------------
part0: on
starting number of considered conformers: 22
program for part0: tm
functional for fast single-point: b97-d
basis set for fast single-point: def2-SV(P)
threshold g_thr(0) for sorting in part0: 4.0
Solvent model used with xTB: alpb
short-notation:
b97-d-D3/def2-SV(P) // GFNn-xTB (Input geometry)
END of parameters
------------------------------------------------------------
PATHS of external QM programs
------------------------------------------------------------
The following program paths are used:
xTB: /home/abt-grimme/AK-bin/xtb
TURBOMOLE: /home/abt-grimme/TURBOMOLE.7.5.1/bin/em64t-unknown-linux-gnu_smp
Using cefine from /home/bohle/bin/cefine
PARNODES for TM or COSMO-RS calculation was set to 2
----------------------------------------------------------------------------------------------------
Processing data from previous run (enso.json)
----------------------------------------------------------------------------------------------------
INFORMATION: No restart information exists and is created during this run!
----------------------------------------------------------------------------------------------------
CRE CHEAP-PRESCREENING - PART0
----------------------------------------------------------------------------------------------------
program: tm
functional for part0: b97-d
basis set for part0: def2-SV(P)
threshold g_thr(0): 4.0
starting number of considered conformers: 22
Calculating efficient gas-phase single-point energies:
The efficient gas-phase single-point is calculated for:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF10, CONF11
CONF12, CONF13, CONF14, CONF15, CONF16, CONF17, CONF18, CONF19, CONF20, CONF21, CONF22
Constructed folders!
Starting 22 ALPB-Gsolv calculations
Running single-point in CONF1/part0_sp
Running single-point in CONF2/part0_sp
Running ALPB_GSOLV calculation in 1-part0/CONF1/part0_sp
Running ALPB_GSOLV calculation in 1-part0/CONF2/part0_sp
Running single-point in CONF3/part0_sp
Running single-point in CONF4/part0_sp
Running ALPB_GSOLV calculation in 1-part0/CONF3/part0_sp
Running ALPB_GSOLV calculation in 1-part0/CONF4/part0_sp
Running single-point in CONF5/part0_sp
Running single-point in CONF6/part0_sp
Running ALPB_GSOLV calculation in 1-part0/CONF5/part0_sp
Running ALPB_GSOLV calculation in 1-part0/CONF6/part0_sp
Running single-point in CONF7/part0_sp
Running single-point in CONF8/part0_sp
Running ALPB_GSOLV calculation in 1-part0/CONF7/part0_sp
Running ALPB_GSOLV calculation in 1-part0/CONF8/part0_sp
Running single-point in CONF9/part0_sp
Running single-point in CONF10/part0_sp
Running ALPB_GSOLV calculation in 1-part0/CONF9/part0_sp
Running ALPB_GSOLV calculation in 1-part0/CONF10/part0_sp
Running single-point in CONF11/part0_sp
Running single-point in CONF12/part0_sp
Running ALPB_GSOLV calculation in 1-part0/CONF11/part0_sp
Running ALPB_GSOLV calculation in 1-part0/CONF12/part0_sp
Running single-point in CONF13/part0_sp
Running single-point in CONF14/part0_sp
Running ALPB_GSOLV calculation in 1-part0/CONF13/part0_sp
Running ALPB_GSOLV calculation in 1-part0/CONF14/part0_sp
Running single-point in CONF15/part0_sp
Running single-point in CONF16/part0_sp
Running ALPB_GSOLV calculation in 1-part0/CONF15/part0_sp
Running ALPB_GSOLV calculation in 1-part0/CONF16/part0_sp
Running single-point in CONF17/part0_sp
Running single-point in CONF18/part0_sp
Running ALPB_GSOLV calculation in 1-part0/CONF17/part0_sp
Running ALPB_GSOLV calculation in 1-part0/CONF18/part0_sp
Running single-point in CONF19/part0_sp
Running single-point in CONF20/part0_sp
Running ALPB_GSOLV calculation in 1-part0/CONF19/part0_sp
Running ALPB_GSOLV calculation in 1-part0/CONF20/part0_sp
Running single-point in CONF21/part0_sp
Running single-point in CONF22/part0_sp
Running ALPB_GSOLV calculation in 1-part0/CONF21/part0_sp
Running ALPB_GSOLV calculation in 1-part0/CONF22/part0_sp
Tasks completed!
The efficient gas-phase single-point was successful for CONF1/part0_sp: E(DFT) = -496.55110270 Gsolv = -0.12279245
The efficient gas-phase single-point was successful for CONF2/part0_sp: E(DFT) = -496.55473079 Gsolv = -0.12241044
The efficient gas-phase single-point was successful for CONF3/part0_sp: E(DFT) = -496.55551554 Gsolv = -0.12099969
The efficient gas-phase single-point was successful for CONF4/part0_sp: E(DFT) = -496.55384665 Gsolv = -0.12156811
The efficient gas-phase single-point was successful for CONF5/part0_sp: E(DFT) = -496.55341065 Gsolv = -0.12251050
The efficient gas-phase single-point was successful for CONF6/part0_sp: E(DFT) = -496.55368828 Gsolv = -0.12175678
The efficient gas-phase single-point was successful for CONF7/part0_sp: E(DFT) = -496.54759208 Gsolv = -0.12365913
The efficient gas-phase single-point was successful for CONF8/part0_sp: E(DFT) = -496.55118920 Gsolv = -0.12200372
The efficient gas-phase single-point was successful for CONF9/part0_sp: E(DFT) = -496.54970422 Gsolv = -0.12167549
The efficient gas-phase single-point was successful for CONF10/part0_sp: E(DFT) = -496.55157272 Gsolv = -0.12001541
The efficient gas-phase single-point was successful for CONF11/part0_sp: E(DFT) = -496.54991969 Gsolv = -0.12296934
The efficient gas-phase single-point was successful for CONF12/part0_sp: E(DFT) = -496.55212770 Gsolv = -0.11751864
The efficient gas-phase single-point was successful for CONF13/part0_sp: E(DFT) = -496.55077718 Gsolv = -0.11854291
The efficient gas-phase single-point was successful for CONF14/part0_sp: E(DFT) = -496.55028792 Gsolv = -0.12090071
The efficient gas-phase single-point was successful for CONF15/part0_sp: E(DFT) = -496.54987374 Gsolv = -0.12311773
The efficient gas-phase single-point was successful for CONF16/part0_sp: E(DFT) = -496.55360018 Gsolv = -0.11274054
The efficient gas-phase single-point was successful for CONF17/part0_sp: E(DFT) = -496.55289531 Gsolv = -0.11378052
The efficient gas-phase single-point was successful for CONF18/part0_sp: E(DFT) = -496.50423172 Gsolv = -0.16468285
The efficient gas-phase single-point was successful for CONF19/part0_sp: E(DFT) = -496.52384365 Gsolv = -0.14390846
The efficient gas-phase single-point was successful for CONF20/part0_sp: E(DFT) = -496.50481476 Gsolv = -0.16519450
The efficient gas-phase single-point was successful for CONF21/part0_sp: E(DFT) = -496.53860751 Gsolv = -0.13212346
The efficient gas-phase single-point was successful for CONF22/part0_sp: E(DFT) = -496.50938650 Gsolv = -0.15826546
----------------------------------------------------------------------------------------------------
Removing high lying conformers by improved energy description
----------------------------------------------------------------------------------------------------
CONF# G [Eh] ΔG [kcal/mol] E [Eh] Gsolv [Eh] Gtot ΔE(DFT) ΔGsolv ΔGtot
GFN2-xTB GFN2-xTB b97-d-D3(0)/def2-SV(P) alpb [Eh] [kcal/mol] [kcal/mol] [kcal/mol]
[ALPB] [ALPB] [gfn2]
CONF1 -34.1746819 0.00 -496.5511027 -0.1227924 -496.6738951 2.28 -0.24 2.04
CONF2 -34.1743917 0.18 -496.5547308 -0.1224104 -496.6771412 0.00 0.00 0.00 <------
CONF3 -34.1742792 0.25 -496.5555155 -0.1209997 -496.6765152 -0.49 0.89 0.39
CONF4 -34.1739821 0.44 -496.5538467 -0.1215681 -496.6754148 0.55 0.53 1.08
CONF5 -34.1740224 0.41 -496.5534106 -0.1225105 -496.6759212 0.83 -0.06 0.77
CONF6 -34.1736309 0.66 -496.5536883 -0.1217568 -496.6754451 0.65 0.41 1.06
CONF7 -34.1725555 1.33 -496.5475921 -0.1236591 -496.6712512 4.48 -0.78 3.70
CONF8 -34.1721876 1.57 -496.5511892 -0.1220037 -496.6731929 2.22 0.26 2.48
CONF9 -34.1719439 1.72 -496.5497042 -0.1216755 -496.6713797 3.15 0.46 3.62
CONF10 -34.1714765 2.01 -496.5515727 -0.1200154 -496.6715881 1.98 1.50 3.48
CONF11 -34.1712638 2.14 -496.5499197 -0.1229693 -496.6728890 3.02 -0.35 2.67
CONF12 -34.1704841 2.63 -496.5521277 -0.1175186 -496.6696463 1.63 3.07 4.70
CONF13 -34.1703687 2.71 -496.5507772 -0.1185429 -496.6693201 2.48 2.43 4.91
CONF14 -34.1694191 3.30 -496.5502879 -0.1209007 -496.6711886 2.79 0.95 3.74
CONF15 -34.1691431 3.48 -496.5498737 -0.1231177 -496.6729915 3.05 -0.44 2.60
CONF16 -34.1664418 5.17 -496.5536002 -0.1127405 -496.6663407 0.71 6.07 6.78
CONF17 -34.1661652 5.34 -496.5528953 -0.1137805 -496.6666758 1.15 5.42 6.57
CONF18 -34.1660003 5.45 -496.5042317 -0.1646828 -496.6689146 31.69 -26.53 5.16
CONF19 -34.1664801 5.15 -496.5238437 -0.1439085 -496.6677521 19.38 -13.49 5.89
CONF20 -34.1658387 5.55 -496.5048148 -0.1651945 -496.6700093 31.32 -26.85 4.48
CONF21 -34.1670164 4.81 -496.5386075 -0.1321235 -496.6707310 10.12 -6.10 4.02
CONF22 -34.1656180 5.69 -496.5093865 -0.1582655 -496.6676520 28.45 -22.50 5.95
----------------------------------------------------------------------------------------------------
--------------------------------------------------
Conformers considered further
--------------------------------------------------
These conformers are below the 4.000 kcal/mol g_thr(0) threshold.
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF10, CONF11
CONF14, CONF15
Calculating Boltzmann averaged (free) energy of ensemble on input geometries (not DFT optimized)!
temperature /K: avE(T) /a.u. avG(T) /a.u.
----------------------------------------------------------------------------------------------------
298.15 -496.5544580 -496.6764445 <<==part0==
----------------------------------------------------------------------------------------------------
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>END of Part0<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
Ran part0 in 32.8727 seconds
Part : #conf time
--------------------------------------------------
Input : 22 -
Part0_all : 13 32.87s
--------------------------------------------------
All parts : 32.87s
CENSO all done!
$CENSO global configuration file: .censorc
$VERSION:1.0.3
ORCA: /home/$USER/orca_4_2_1_linux_x86-64_openmpi216
ORCA version: 4.2.1
GFN-xTB: /home/$USER/bin/xtb
CREST: /home/$USER/bin/crest
mpshift: /home/$USER/TURBOMOLE.7.5/bin/em64t-unknown-linux-gnu/mpshift
escf: /home/$USER/TURBOMOLE.7.5/bin/em64t-unknown-linux-gnu/escf
#COSMO-RS
ctd = BP_TZVPD_FINE_C30_1601.ctd cdir = "/home/USER/COSMOlogic/COSMOthermX19/COSMOtherm/CTDATA-FILES" ldir = "/home/$USER/COSMOlogic/COSMOthermX19/COSMOtherm/CTDATA-FILES"
cosmothermversion: 19
$ENDPROGRAMS
$CRE SORTING SETTINGS:
$GENERAL SETTINGS:
nconf: all # ['all', 'number e.g. 10 up to all conformers']
charge: 0 # ['number e.g. 0']
unpaired: 0 # ['number e.g. 0']
solvent: gas # ['gas', 'acetone', 'chcl3', 'acetonitrile', 'ch2cl2', 'dmso', 'h2o', 'methanol', 'thf', '...']
prog_rrho: xtb # ['xtb', 'prog']
temperature: 298.15 # ['temperature in K e.g. 298.15']
trange: [273.15, 378.15, 5] # ['temperature range [start, end, step]']
multitemp: on # ['on', 'off']
evaluate_rrho: on # ['on', 'off']
consider_sym: off # ['on', 'off']
bhess: on # ['on', 'off']
imagthr: automatic # ['automatic or e.g., -100 # in cm-1']
sthr: automatic # ['automatic or e.g., 50 # in cm-1']
scale: automatic # ['automatic or e.g., 1.0 ']
rmsdbias: off # ['on', 'off']
sm_rrho: alpb # ['alpb', 'gbsa']
check: on # ['on', 'off']
prog: tm # ['tm', 'orca']
func: r2scan-3c # ['pbe', 'b97-d', 'pbeh-3c', 'tpss', 'b97-d3', 'r2scan-3c', 'b97-3c']
basis: automatic # ['automatic', 'def2-mSVP', 'def2-mTZVP', 'def2-mTZVP', 'def2-TZVP', '...']
maxthreads: 2 # ['number of threads e.g. 2']
omp: 2 # ['number cores per thread e.g. 4']
cosmorsparam: automatic # ['automatic', '12-fine', '12-normal', '13-fine', '13-normal', '14-fine', '...']
$PART0 - CHEAP-PRESCREENING - SETTINGS:
part0: off # ['on', 'off']
func0: b97-d # ['pbeh-3c', 'b97-3c', 'b97-d3', 'pbe', 'r2scan-3c', 'tpss', 'b97-d']
basis0: def2-SV(P) # ['automatic', 'def2-mSVP', 'def2-mTZVP', 'def2-mTZVP', 'def2-TZVP', '...']
part0_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
part0_threshold: 4.0 # ['number e.g. 4.0']
$PART1 - PRESCREENING - SETTINGS:
# func and basis is set under GENERAL SETTINGS
part1: off # ['on', 'off']
smgsolv1: cosmors # ['alpb_gsolv', 'dcosmors', 'cosmo', 'smd', 'cosmors-fine', 'cpcm', 'gbsa_gsolv', '...']
part1_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
part1_threshold: 3.5 # ['number e.g. 5.0']
$PART2 - OPTIMIZATION - SETTINGS:
# func and basis is set under GENERAL SETTINGS
part2: off # ['on', 'off']
opt_limit: 2.5 # ['number e.g. 4.0']
sm2: dcosmors # ['cosmo', 'cpcm', 'default', 'smd', 'dcosmors']
smgsolv2: cosmors # ['cosmors-fine', 'gbsa_gsolv', 'cosmo', 'cpcm', 'smd_gsolv', 'alpb_gsolv', 'smd', '...']
part2_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
ancopt: on # ['on']
hlow: 0.01 # ['lowest force constant in ANC generation, e.g. 0.01']
opt_spearman: on # ['on', 'off']
part2_threshold: 99 # ['Boltzmann sum threshold in %. e.g. 95 (between 1 and 100)']
optlevel2: automatic # ['crude', 'sloppy', 'loose', 'lax', 'normal', 'tight', 'vtight', 'extreme', '...']
optcycles: 8 # ['number e.g. 5 or 10']
spearmanthr: -4.0 # ['value between -1 and 1, if outside set automatically']
radsize: 10 # ['number e.g. 8 or 10']
crestcheck: off # ['on', 'off']
$PART3 - REFINEMENT - SETTINGS:
part3: off # ['on', 'off']
prog3: prog # ['tm', 'orca', 'prog']
func3: pw6b95 # ['dsd-blyp', 'pw6b95', 'b97-d3', 'r2scan-3c', 'pbe0', 'wb97x']
basis3: def2-TZVPD # ['SVP', 'SV(P)', 'TZVP', 'TZVPP', 'QZVP', 'QZVPP', 'def2-SV(P)', 'def2-mSVP', '...']
smgsolv3: cosmors # ['alpb_gsolv', 'dcosmors', 'cosmo', 'smd', 'cosmors-fine', 'cpcm', 'gbsa_gsolv', '...']
part3_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
part3_threshold: 99 # ['Boltzmann sum threshold in %. e.g. 95 (between 1 and 100)']
$NMR PROPERTY SETTINGS:
$PART4 SETTINGS:
part4: off # ['on', 'off']
couplings: on # ['on', 'off']
progJ: prog # ['tm', 'orca', 'adf', 'prog']
funcJ: pbe0 # ['tpss', 'pbe0', 'pbeh-3c']
basisJ: def2-TZVP # ['SVP', 'SV(P)', 'TZVP', 'TZVPP', 'QZVP', 'QZVPP', 'def2-SV(P)', 'def2-mSVP', '...']
sm4J: default # ['dcosmors', 'smd', 'cosmo', 'cpcm']
shieldings: on # ['on', 'off']
progS: prog # ['tm', 'orca', 'adf', 'prog']
funcS: pbe0 # ['dsd-blyp', 'pbeh-3c', 'tpss', 'pbe0', 'kt2']
basisS: def2-TZVP # ['SVP', 'SV(P)', 'TZVP', 'TZVPP', 'QZVP', 'QZVPP', 'def2-SV(P)', 'def2-mSVP', '...']
sm4S: default # ['dcosmors', 'smd', 'cosmo', 'cpcm']
reference_1H: TMS # ['TMS']
reference_13C: TMS # ['TMS']
reference_19F: CFCl3 # ['CFCl3']
reference_29Si: TMS # ['TMS']
reference_31P: TMP # ['TMP', 'PH3']
1H_active: on # ['on', 'off']
13C_active: on # ['on', 'off']
19F_active: off # ['on', 'off']
29Si_active: off # ['on', 'off']
31P_active: off # ['on', 'off']
resonance_frequency: 300.0 # ['MHz number of your experimental spectrometer setup']
$OPTICAL ROTATION PROPERTY SETTINGS:
$PART5 SETTINGS:
optical_rotation: off # ['on', 'off']
funcOR: pbe # ['functional for opt_rot e.g. pbe']
funcOR_SCF: r2scan-3c # ['functional for SCF in opt_rot e.g. r2scan-3c']
basisOR: def2-SVPD # ['basis set for opt_rot e.g. def2-SVPD']
frequency_optical_rot: [589.0] # ['list of freq in nm to evaluate opt rot at e.g. [589, 700]']
$END CENSORC
25
-34.15995484
O -2.7404108553 -0.9210263756 0.0505546155
O -0.6532148483 -1.7303331963 0.1381884568
N -2.1811530948 1.4662136641 -0.3280604715
N 1.9999042791 -1.6115147219 -0.1737452561
C 1.2832974514 1.7494376339 0.2478643443
C 0.0382242755 1.0547713107 0.8044626435
C 2.1972182535 0.8461961224 -0.5852172042
C -0.9624505804 0.6276788621 -0.2713633850
C 2.7304045776 -0.3664138808 0.1827513585
C -1.5037598584 -0.8390454546 0.0048901917
H 1.8561477225 2.1402514306 1.0892838382
H 0.9852377848 2.6009114549 -0.3642611073
H -0.4518270956 1.7282098745 1.5089809935
H 0.3322545004 0.1775252924 1.3825123935
H 1.6820235078 0.5059218506 -1.4832357051
H 3.0380251642 1.4546200515 -0.9165284484
H -0.4868586610 0.6089411942 -1.2530945560
H 2.6388482788 -0.2021893439 1.2562535001
H 3.7853986884 -0.5188774904 -0.0473087348
H -2.2380899515 2.1426216511 0.4407329422
H -2.3218532286 1.9466721688 -1.2205963738
H 2.3394829642 -2.3992859458 0.3882351555
H 0.9350292280 -1.5547588441 -0.0165809300
H 2.1475890354 -1.8368760327 -1.1636223077
H -2.9328940343 0.6971914121 -0.1921053926
25
-34.15944839
O -2.5910945174 -0.7526707388 0.4737473250
O -0.5258609320 -1.6055473558 0.4259946366
N -2.1883047577 1.2429283468 -0.9276187078
N 2.0221224751 -1.5482527990 -0.3635876013
C 1.0959612422 0.9269941773 1.0928268792
C -0.1118183900 1.5287127427 0.3646185610
C 2.3883955950 0.8232150730 0.2771806877
C -0.9137052232 0.5859776609 -0.5447048684
C 2.3984857923 -0.1999516224 -0.8542391926
C -1.3796771114 -0.7342464594 0.2052799197
H 0.8202702309 -0.0346561349 1.5230052762
H 1.3150554091 1.5881707649 1.9329336393
H 0.2252898792 2.3837647705 -0.2251981715
H -0.7970460540 1.8894152433 1.1356879771
H 2.6174364896 1.7989347868 -0.1517789136
H 3.1942064704 0.5825472961 0.9724285437
H -0.3499369029 0.3059081761 -1.4335569624
H 3.4048136146 -0.2402683769 -1.2749539435
H 1.7122153210 0.0852818535 -1.6492945642
H -2.2438155060 2.2226460139 -0.6308218074
H -2.4094139539 1.1700691932 -1.9248068681
H 0.9927667520 -1.5675551657 -0.0428313977
H 2.1377830502 -2.2542373686 -1.0973459319
H 2.6054467969 -1.8083924261 0.4393088723
H -2.8796747565 0.6310313638 -0.3580585669
25
-34.15925304
O -2.6152400314 -1.0465052003 0.2378304729
O -0.5362082855 -1.4663149505 -0.4593671015
N -2.4557715395 1.4039918868 0.4362491058
N 2.1447268012 -1.5911295291 -0.0908488427
C 1.1849792140 1.1291603967 -0.7123125066
C 0.0549239500 1.3190958463 0.3109988160
C 2.5516538580 0.8614163999 -0.0796419105
C -1.2770304521 0.8481909298 -0.2699298296
C 2.6278492228 -0.4408272994 0.7143702153
C -1.4798499370 -0.7275247362 -0.1594413730
H 1.2638776944 2.0288103522 -1.3231621409
H 0.9216680773 0.3153070715 -1.3864974308
H -0.0132519646 2.3752873127 0.5749134480
H 0.2604586805 0.7576942571 1.2223482952
H 3.2969117377 0.8423750312 -0.8755852689
H 2.8115908685 1.6810094994 0.5898815386
H -1.3353089850 1.1014685130 -1.3310936133
H 2.0240292821 -0.3842292575 1.6193674755
H 3.6638250279 -0.6160286136 1.0070988283
H -2.2160156909 1.7709567167 1.3631094761
H -2.9723151833 2.1102297821 -0.0929227069
H 2.6253531272 -1.6216026098 -0.9974699479
H 2.3064470721 -2.4761725884 0.4009650983
H 1.0998172314 -1.5111491960 -0.2664030290
H -3.0342449423 0.4895941432 0.5473101358
25
-34.15909539
O -2.5859531813 -0.8098009525 0.4685129327
O -0.4816245052 -1.5436026390 0.3102868940
N -2.3339104715 1.3007756781 -0.7836622820
N 2.0492564794 -1.5871472251 -0.5900906398
C 1.1052196056 0.9870949301 0.9271737021
C -0.2140772115 1.5925038037 0.4451868362
C 2.1667497593 0.8464686753 -0.1771749747
C -1.0154302176 0.6861215046 -0.4984780600
C 2.9073785151 -0.4823039670 -0.0859386105
C -1.3840762559 -0.7076318190 0.1707141950
H 0.8920163207 0.0201435075 1.3815317896
H 1.4930774855 1.6249540613 1.7213050551
H -0.0067258320 2.5368714102 -0.0630175124
H -0.8282372023 1.7972548662 1.3249938991
H 1.7135886317 0.9350726379 -1.1636100949
H 2.8915967677 1.6540129472 -0.0882838850
H -0.4677225963 0.5007855037 -1.4227063242
H 3.1688824777 -0.6847294678 0.9524589517
H 3.8237876908 -0.4492687854 -0.6762385697
H -2.5870605138 1.3036833133 -1.7755142934
H -2.9689761577 0.6014426661 -0.2449843511
H 1.9723283573 -1.5417442554 -1.6118847971
H 2.4360322309 -2.5007606995 -0.3327999527
H 1.0548817542 -1.5294780724 -0.1881461707
H -2.4296444696 2.2452869585 -0.3976639893
25
-34.15892503
O -2.4565204045 -0.8423454324 0.7070928712
O -0.5980888346 -1.6084799899 -0.2741310513
N -2.2081439307 1.5499049911 0.0797136762
N 2.0665657616 -1.5738488309 0.1665385721
C 0.8489992162 1.1021214800 0.5754910780
C 0.1319038004 1.1643519245 -0.7806607655
C 2.3611287197 0.8895018711 0.4504901307
C -1.3084663819 0.6750096988 -0.7114456668
C 2.7727042231 -0.3622083435 -0.3226992137
C -1.4583757981 -0.7537800183 -0.0274928276
H 0.4155006230 0.3036523983 1.1797441237
H 0.7020515096 2.0340597672 1.1228295970
H 0.6304477123 0.5210588124 -1.5026223849
H 0.1602968624 2.1778026423 -1.1819014190
H 2.8058653217 1.7473421346 -0.0537220547
H 2.7810957945 0.8489865537 1.4562868383
H -1.7106345304 0.5638398621 -1.7216356404
H 3.8478441803 -0.5048945341 -0.2050768697
H 2.5630334022 -0.2516058916 -1.3860041489
H -2.8269415029 2.1396743466 -0.4823076953
H -2.7642911303 0.7978509993 0.6164252281
H 2.4375926491 -2.4160896797 -0.2859521821
H 2.1793786728 -1.6693140839 1.1823811296
H 1.0245787819 -1.5392035408 -0.0507409777
H -1.6858363886 2.1297513428 0.7452520673
25
-34.15874648
O -2.3605981473 -0.8116042758 0.8659409274
O -0.5843274484 -1.6230194998 -0.2244865945
N -2.2109877661 1.5364887021 0.0742080589
N 2.0757147084 -1.4672125485 -0.4830865528
C 0.8597549995 1.2262238823 0.4517476395
C 0.0874032105 1.1513231513 -0.8771132949
C 2.3512317789 0.8985689618 0.2982329522
C -1.3361657088 0.6347191585 -0.7132778341
C 2.6824356905 -0.5712975375 0.5355813092
C -1.4234260659 -0.7566236862 0.0528412632
H 0.4221004807 0.5356789162 1.1753502177
H 0.7733707657 2.2309740830 0.8650565898
H 0.5827203837 0.4802075195 -1.5756071724
H 0.0631682297 2.1335262804 -1.3514203271
H 2.7003072358 1.2129278451 -0.6855204577
H 2.9172351560 1.4667500260 1.0360745366
H -1.7833398578 0.4616073570 -1.6955611361
H 2.3142140534 -0.8770131145 1.5154931358
H 3.7656889000 -0.7005750162 0.5151167425
H -1.6671444199 2.1735125324 0.6661186569
H -2.8827343086 2.0695764162 -0.4835506438
H 2.3643871815 -1.1879653946 -1.4273853498
H 2.3725094498 -2.4360036549 -0.3237631449
H 1.0071911744 -1.4659435853 -0.4227446705
H -2.7090441201 0.8070062032 0.6949537917
25
-34.15780648
O -0.6998712600 -1.7595821198 0.3315266042
O -2.7564980797 -0.8922468771 0.1127087169
N -2.0912345729 1.3897515697 -0.6207661035
N 1.9218668343 -1.5538051846 -0.0911139226
C 1.2724611881 1.7665596691 0.2614262584
C -0.0126245071 1.0963347719 0.7686474620
C 2.5091659413 0.8642624497 0.2746224048
C -0.9098130242 0.5407794733 -0.3415020783
C 2.4929985644 -0.3196472927 -0.6935656161
C -1.5185682699 -0.8616584773 0.0830057999
H 1.4961397665 2.6156227165 0.9073501428
H 1.1219744652 2.1603919352 -0.7434483203
H -0.5894557289 1.8123867185 1.3552290309
H 0.2456564204 0.2915541326 1.4587568868
H 3.3629575544 1.4905499194 0.0148013745
H 2.6790096911 0.5030846620 1.2896168914
H -0.3423105707 0.3842564461 -1.2596402044
H 3.5203242755 -0.5424656780 -0.9864282797
H 1.9321769951 -0.0751719518 -1.5947809502
H -2.8797605313 0.6907420298 -0.3760235139
H -2.1503924665 2.2120232522 -0.0104200923
H 2.3803263882 -1.7503210865 0.8055549093
H 0.8561607870 -1.5268128372 0.0892049642
H 2.0800598725 -2.3543999415 -0.7123089418
H -2.1774862389 1.6805852373 -1.5986092278
25
-34.15712409
O -0.5169415866 -1.4677953372 -0.4155166026
O -2.6718105717 -1.1029442546 0.0532698902
N -2.5041882719 1.3222450456 0.5356135683
N 2.0547500815 -1.4660199090 0.3952674841
C 1.1460672076 1.2350620947 -0.5905524477
C 0.0199991306 1.2471137062 0.4571994904
C 2.5324119546 0.9449216441 0.0010319703
C -1.3028765754 0.8219664596 -0.1753086903
C 2.9759584940 -0.4928036507 -0.2449354813
C -1.5027299259 -0.7570468322 -0.1889883845
H 1.1567284127 2.2014747982 -1.0932584224
H 0.9206483903 0.4869950226 -1.3509226784
H -0.0730555237 2.2557886152 0.8619704582
H 0.2503077610 0.5750867956 1.2847618985
H 3.2724883861 1.5930094341 -0.4660504111
H 2.5411017268 1.1667575642 1.0680525607
H -1.3439398378 1.1490834108 -1.2170635960
H 3.9809942022 -0.6416689206 0.1522940223
H 2.9906906878 -0.6917352586 -1.3167852841
H -3.1147358709 0.4302326474 0.4978236161
H -2.3055244586 1.5552316017 1.5147727778
H 1.0589542040 -1.3722826387 0.0237179616
H 2.3537713222 -2.4274392985 0.2004063016
H 2.0384435533 -1.3281897829 1.4119124615
H -2.9671283162 2.1123945703 0.0792841186
25
-34.15703719
O -0.4840204654 -1.5908390855 -0.1977054306
O -2.6424172363 -1.0172969494 -0.2313294170
N -2.3824391815 1.4118161237 0.1129756179
N 2.0591606020 -1.4080559447 0.6444993748
C 1.3255949204 1.5932768744 0.1316443482
C 0.0292286137 1.1405631347 0.8180158808
C 1.9611296764 0.5914767293 -0.8382657525
C -1.0774245534 0.7702791240 -0.1744053472
C 2.8498636045 -0.4645206094 -0.1922799795
C -1.4218007565 -0.7850501416 -0.2021503198
H 2.0511714191 1.8626404605 0.8996124027
H 1.1017826165 2.5033885185 -0.4265728164
H -0.3083479528 1.9763847128 1.4332069953
H 0.2033356421 0.3027453082 1.4918974312
H 1.1939462785 0.0852292002 -1.4240400771
H 2.5871303840 1.1474547523 -1.5362113415
H -0.7775416723 1.0312499156 -1.1919311777
H 3.6205966803 0.0055444316 0.4194317100
H 3.3361661233 -1.0318279746 -0.9856062704
H -2.6284628403 2.1753990573 -0.5216811303
H -3.0325627338 0.5540981187 -0.0284710758
H 2.4910941867 -2.3367192092 0.6680136974
H 1.9781398838 -1.0663385144 1.6071566991
H 1.0702227114 -1.5059322479 0.2455456256
H -2.4592842029 1.7302943098 1.0848423546
25
-34.15632171
O -0.6377330316 -1.5602972801 -0.5925407457
O -1.9438226066 -0.8962122015 1.0968963302
N -2.0691310397 1.5115608178 0.5520085904
N 2.0161394685 -1.6864701960 -0.6808935380
C 1.0245208700 1.5678424724 0.3114580637
C 0.0268814843 1.3822638313 -0.8376423667
C 1.7125418505 0.2918072015 0.8062089597
C -1.3255436976 0.7602094261 -0.4861917907
C 2.5798867778 -0.3735189736 -0.2699515146
C -1.2821941821 -0.7269243894 0.0579815663
H 0.5483025277 2.0449086772 1.1692286093
H 1.7966540418 2.2561425086 -0.0359536522
H 0.4649670843 0.7632830938 -1.6196837604
H -0.1661732017 2.3619672870 -1.2796881790
H 2.3322565126 0.5621906956 1.6605054657
H 0.9655697921 -0.4140620317 1.1690705974
H -1.9201614836 0.7061105980 -1.4020292431
H 3.5936748701 -0.5312821922 0.0979019662
H 2.6369135707 0.2686143194 -1.1490685384
H -2.9086496435 1.9840429736 0.2094100180
H -2.3313717130 0.6888337551 1.2084854943
H 2.1820919942 -2.3913736082 0.0460188498
H 0.9581287874 -1.6201065231 -0.7926007692
H 2.4297604389 -2.0163864988 -1.5591512145
H -1.4715716162 2.1835379113 1.0453989384
25
-34.15592507
O -2.1033969149 -0.8344078612 0.9487088877
O -0.5699612374 -1.6680274471 -0.4517081377
N -1.9569275917 1.5465619838 0.2627686808
N 2.0837337252 -1.3633283870 -0.5773762363
C 1.1075416509 1.5419562581 -0.0103918263
C 0.1209945172 1.1686244794 -1.1251953409
C 1.4352918321 0.4570859531 1.0214669735
C -1.2503428799 0.6557309102 -0.6867527508
C 2.5057372966 -0.5407868185 0.5902823997
C -1.2948162683 -0.7745608298 0.0075959523
H 0.7274518351 2.4141375453 0.5251162579
H 2.0330446007 1.8691474761 -0.4864359338
H 0.5312965142 0.4086384701 -1.7869917742
H -0.0324587227 2.0621064981 -1.7347668750
H 1.8169519144 0.9463147999 1.9183948838
H 0.5367641743 -0.0848292604 1.3149274400
H -1.8634330878 0.5270669166 -1.5839659846
H 2.7015999487 -1.2099014501 1.4282318085
H 3.4296803314 -0.0167769980 0.3414844052
H -1.2992225514 2.1272618717 0.7958603647
H -2.6755557698 2.1370116562 -0.1630903688
H 2.2813676133 -0.8782194673 -1.4586367594
H 2.5735263109 -2.2635096262 -0.5875520766
H 1.0325521121 -1.5556317222 -0.5353874345
H -2.3935764356 0.8063761382 0.9145561672
25
-34.15547499
O -2.3288681540 -0.9689367039 0.5974852723
O -0.4008216845 -1.3559960891 -0.4559450313
N -2.6333104330 1.3901145964 0.0148791472
N 2.2688734235 -1.7836122490 -0.3135245436
C 1.2045970266 1.2147830910 -0.3207104722
C -0.1766849998 1.7049883679 0.1388725454
C 1.9421007389 0.4104322018 0.7578355455
C -1.3241367595 0.8966054975 -0.4749078790
C 2.9601024995 -0.5430248114 0.1410496733
C -1.3285933693 -0.6410895869 -0.0712604222
H 1.8194085679 2.0702167267 -0.5988277282
H 1.0710671602 0.6065579215 -1.2146344993
H -0.3076575365 2.7461839050 -0.1612866699
H -0.2378627475 1.6551531995 1.2275331138
H 2.4468113264 1.0960018700 1.4370167059
H 1.2297127637 -0.1683684162 1.3468389979
H -1.2704988015 0.9491259299 -1.5640455987
H 3.7310656000 -0.8072491298 0.8640696447
H 3.4371256948 -0.0632845904 -0.7133472266
H -2.5432643621 2.1520849835 0.6930123940
H -3.2834926874 1.6599628672 -0.7262042831
H 2.7178100064 -2.1982719631 -1.1361624210
H 2.2482990815 -2.4828183187 0.4368503414
H 1.2567750610 -1.5723871751 -0.5319861416
H -2.9757946227 0.4721100866 0.5082732629
25
-34.15545846
O -2.3777023451 -0.9177733836 0.6489622803
O -0.4009151018 -1.4460423055 -0.2363506290
N -2.5928675119 1.3702602408 -0.1958788709
N 2.2705863538 -1.5552728060 -0.6959671399
C 1.2626408325 1.2650767695 -0.1999508619
C -0.1676900608 1.6997857440 0.1433859727
C 1.8576282855 0.3222355626 0.8537184709
C -1.2484356318 0.8255651567 -0.5054674146
C 2.9084111559 -0.6168744122 0.2764230940
C -1.3258577703 -0.6732783629 0.0271901706
H 1.8935551201 2.1501893106 -0.2772016053
H 1.2458750287 0.7908735642 -1.1798122422
H -0.3171466901 2.7186794233 -0.2201486160
H -0.2967658760 1.7007241110 1.2272150872
H 2.3089550370 0.9081814099 1.6535209776
H 1.0686425282 -0.2818531316 1.3016188748
H -1.0876822336 0.7817119785 -1.5847014431
H 3.3524799930 -1.1924945383 1.0873745985
H 3.6963304583 -0.0519426265 -0.2212122945
H -2.9913275573 0.5162412164 0.3615908966
H -2.5589278358 2.2025641690 0.3999047288
H 2.6187881911 -2.5117770389 -0.5815509942
H 1.2217357961 -1.5619680721 -0.5321789754
H 2.4345460024 -1.2660811775 -1.6650051770
H -3.1696789754 1.5558016135 -1.0193997348
25
-34.15372845
O -0.8759513206 -1.0507714945 0.7395130487
O -1.0471783147 -1.2210634731 -1.4857197061
N -2.0100713682 1.1638582833 0.9733772433
N 1.7020599065 -1.7447966827 0.8365171560
C 0.9224701988 1.3977552196 0.1299066020
C -0.3034955554 1.7474745141 -0.7290850754
C 1.8313865205 0.3371413309 -0.5152183962
C -1.5397536849 0.8954837973 -0.4124258872
C 2.5286316072 -0.5462459399 0.5204181162
C -1.1337894558 -0.6241211220 -0.4224393440
H 0.5999134506 1.0447379113 1.1088504940
H 1.5075448530 2.3032957331 0.2891927248
H -0.0603736240 1.6138572758 -1.7832639404
H -0.5689030950 2.7975685720 -0.5933476217
H 1.2520636483 -0.2949117808 -1.1892484363
H 2.5821478388 0.8362393886 -1.1261103610
H -2.3194229349 1.0797505602 -1.1497937252
H 2.6951475316 0.0179430508 1.4383444706
H 3.4942793529 -0.8824584649 0.1428332329
H -1.6366862386 2.0406897131 1.3566364027
H -3.0319635647 1.1643591957 1.0502533535
H 2.0123953855 -2.1956055335 1.7033301803
H 0.6617699349 -1.4988916853 0.9180060433
H 1.7620357216 -2.4244612968 0.0700935319
H -1.6155014744 0.3110635636 1.4848864268
25
-34.15296366
O -0.9806508876 -0.9847570656 0.9016095316
O -1.0342961735 -1.4975369515 -1.2790158618
N -2.0600699488 1.3003330160 0.6598555012
N 1.6483681534 -1.4770759046 1.0193012553
C 0.8930197778 1.4409262402 0.3162871289
C -0.1018287155 1.4892580056 -0.8622608316
C 2.2127299908 0.7309357778 -0.0043863659
C -1.4370888955 0.7837090011 -0.5889439421
C 2.1074033787 -0.7768274559 -0.2134156979
C -1.1447588094 -0.7374998177 -0.3265126418
H 0.4424872625 0.9591479756 1.1822965353
H 1.1476108940 2.4604379775 0.6056686014
H 0.3344753521 1.0203527521 -1.7446629435
H -0.3104542151 2.5259312846 -1.1308040482
H 2.6400205950 1.1617275571 -0.9101356051
H 2.9157320587 0.9267783500 0.8074716570
H -2.0965188879 0.8923814409 -1.4486238017
H 3.0903725751 -1.1592546293 -0.4905246456
H 1.4081882505 -1.0149227036 -1.0156071158
H -3.0831759004 1.3363322990 0.6000761811
H -1.7780292676 0.5566802205 1.3629478428
H 0.5935972201 -1.3552723672 1.1310943683
H 1.8230458958 -2.4842696303 0.9418471118
H 2.1352867113 -1.1124232113 1.8467861942
H -1.6994055067 2.2241075607 0.9291898687
25
-34.15141381
O -1.3943785432 -1.0307567325 0.9025777408
O -0.4225106153 -1.2766835982 -1.0942963862
N -2.1382332497 1.3072100696 0.6820859684
N 1.8303685962 -1.9710495392 0.0707587666
C 0.9607624662 1.6168605590 0.1707894371
C -0.1959862014 1.7114505222 -0.8393306352
C 1.9799050629 0.5013843169 -0.1348429740
C -1.4247901716 0.8500833461 -0.5368218294
C 1.9371661639 -0.6960029627 0.8273160062
C -1.0467465663 -0.6453837331 -0.2300826075
H 0.5821582777 1.4936196592 1.1855414824
H 1.4853467761 2.5717069600 0.1437592453
H 0.1684852019 1.4328271659 -1.8279890567
H -0.5295558410 2.7487716363 -0.9068101296
H 1.7990108513 0.1423863706 -1.1479579769
H 2.9805578856 0.9303497506 -0.1217137710
H -2.0885910031 0.8601550715 -1.4024758281
H 1.0694995988 -0.6377417602 1.4853144468
H 2.8334350246 -0.7245407799 1.4470296049
H -1.6564311817 2.0816802979 1.1506799283
H -3.1156981590 1.5603768219 0.5197984898
H 2.6238854975 -2.0888513426 -0.5678832194
H 1.7803639034 -2.7755595157 0.7040003308
H 0.9218328585 -1.9204346441 -0.4990801476
H -2.0745975566 0.3957317695 1.2728068192
25
-34.15122546
O -1.4776298354 -0.9779162840 0.9689911790
O -0.4925234496 -1.3628551244 -1.0036507954
N -2.1639329473 1.3683833598 0.5828976748
N 1.8796097534 -1.9018199765 -0.0324662357
C 0.9212842989 1.5787487906 0.2022882964
C -0.1575522111 1.5949891498 -0.8983466380
C 2.0807266384 0.5903317386 -0.0312863938
C -1.4336452672 0.8124987799 -0.5833640723
C 2.0372506913 -0.6914975753 0.8153963432
C -1.1077721831 -0.6712596326 -0.1784565029
H 0.4772061571 1.3762504059 1.1769048440
H 1.3491352329 2.5796760629 0.2516191005
H 0.2519128515 1.1866703660 -1.8221067218
H -0.4443786804 2.6264492546 -1.1098247151
H 2.1069094344 0.3214704229 -1.0870338715
H 3.0133169149 1.1098047495 0.1839278312
H -2.0690635873 0.7917157945 -1.4696202320
H 1.1994207751 -0.6693189818 1.5132782369
H 2.9569334194 -0.7921592267 1.3922309558
H -1.6800036187 2.1679532791 1.0053931771
H -3.1334683512 1.6233161070 0.3791665015
H 2.6232513605 -1.9579134554 -0.7362257317
H 1.8912989959 -2.7525757838 0.5392212719
H 0.9273933706 -1.8444066710 -0.5257136187
H -2.1333692549 0.5050567581 1.2388793063
25
-34.15102491
O -2.7007099152 -1.6852260154 -0.4665848832
O -3.5578446821 0.0350414384 0.6950996250
N -1.7249631955 1.6316108345 0.4298941943
N 4.8503432460 -0.2765578553 0.0049566510
C 1.1047501969 0.4501828237 -0.3379055216
C -0.1445140391 -0.2886936630 0.1459845288
C 2.3741940866 -0.2828979224 0.0965187068
C -1.4339016427 0.3793165613 -0.3028003047
C 3.6160850376 0.4539254738 -0.3951671902
C -2.6934727541 -0.5488189533 -0.0039847520
H 1.1202182595 1.4641232269 0.0631636060
H 1.0744507790 0.5160807326 -1.4261342120
H -0.1466884742 -0.3764861277 1.2347858649
H -0.1614173698 -1.3033571375 -0.2567289346
H 2.3826839006 -0.3550960319 1.1849100754
H 2.3458274825 -1.2937888279 -0.3120785959
H -1.4193517541 0.5529473168 -1.3796978179
H 3.6476836700 1.4575725004 0.0303995802
H 3.5908364898 0.5356651831 -1.4823627548
H -1.7888195298 2.4639789876 -0.1581990321
H -2.7206446817 1.3385249836 0.8116492773
H 5.6915565940 0.2174183391 -0.3259194480
H 4.8962264202 -0.3605243899 1.0314916029
H 4.8443782262 -1.2261178281 -0.3963638456
H -1.0847775954 1.7904079765 1.2125175138
25
-34.15096274
O -3.0664396656 -0.6811514259 0.4732552569
O -1.1830540829 -1.6357953168 -0.2743493731
N -2.5677177347 1.6455787239 -0.0357930901
N 3.4161836207 -1.4947231742 0.0043983586
C 1.0526738400 0.4198619079 -0.3271341828
C -0.1392962419 1.1756334823 0.2601668795
C 2.3301787116 0.6870764296 0.4694516416
C -1.4374925629 0.7826136622 -0.4432938226
C 3.5433351448 -0.0155439223 -0.1263696890
C -1.9232732343 -0.6849210780 -0.0466890927
H 1.1914397265 0.7321338619 -1.3635085567
H 0.7858921189 -0.6373779302 -0.3204311762
H 0.0218209785 2.2505182304 0.1468292069
H -0.2266241551 0.9455046686 1.3247226359
H 2.5278796928 1.7590277621 0.4727119199
H 2.1882724928 0.3782271446 1.5058219746
H -1.3025118280 0.8032883597 -1.5259809861
H 4.4479425383 0.3027435209 0.3921583620
H 3.6346463266 0.2349775607 -1.1836334244
H -3.0211573542 2.1410074345 -0.8045922540
H -3.2218629847 0.8371482423 0.3587042034
H 4.2253281871 -1.9726822174 -0.4155872948
H 3.3489651634 -1.7581308116 0.9983748528
H 2.5570336567 -1.8144640232 -0.4679619280
H -2.3203269015 2.3008181314 0.7098687755
25
-34.15085064
O -3.5353488917 0.0322108098 0.4084906022
O -2.4254579214 -1.9084093375 0.6284254091
N -1.7744403653 1.3652775888 -0.6438663864
N 4.1843308757 -0.1227935311 -1.0149931633
C 1.2054477125 0.3338345853 -0.2573250215
C -0.0723683835 -0.0922253390 0.4701395235
C 2.4067626747 0.2476116899 0.6871054133
C -1.3095998224 -0.0201188789 -0.4106994862
C 3.6989393857 0.7735207317 0.0732627710
C -2.5481677099 -0.7347193516 0.2919952398
H 1.1039179513 1.3593179418 -0.6150772671
H 1.3305507433 -0.3269971917 -1.1156830038
H -0.2323641779 0.5179203592 1.3619095074
H 0.0148258366 -1.1272987239 0.8068454593
H 2.1939049448 0.8432977651 1.5748900270
H 2.5356727482 -0.7824695024 1.0211816116
H -1.1430786561 -0.5391980852 -1.3555500184
H 4.4681954117 0.8303339300 0.8434799998
H 3.5410323285 1.7707403176 -0.3387001734
H -2.7959421798 1.2370421526 -0.2418926774
H -1.2663143192 2.0543977636 -0.0827359543
H 4.3101055621 -1.0795234870 -0.6518667702
H 3.4997443822 -0.1538144186 -1.7841531054
H 5.0842467659 0.2120203831 -1.3873716549
H -1.8037773550 1.6476905491 -1.6249441216
25
-34.15048917
O -1.9774584219 -0.8822136242 -1.3447844876
O -0.7353222350 -1.3040874260 0.4712779300
N -1.2018711229 0.8204001109 1.6759883927
N 1.9464051556 -1.6923491374 0.3496868798
C 0.8058411616 1.0462579377 -0.9468506072
C -0.4754178942 1.7785998054 -0.5275051890
C 1.8226970853 0.7789686777 0.1752315710
C -1.5019161161 0.9155320919 0.2220611780
C 2.6576744598 -0.4659344426 -0.1177787426
C -1.4255171488 -0.5796126398 -0.2987743324
H 1.2927227489 1.6467822241 -1.7161034319
H 0.5115436171 0.1136959367 -1.4299162738
H -0.9626535770 2.1244807715 -1.4395462399
H -0.2313854006 2.6631384020 0.0636669169
H 2.4836824282 1.6401220787 0.2631946203
H 1.3379895832 0.6415415675 1.1385387246
H -2.5044279955 1.3042479639 0.0511624497
H 3.6220420553 -0.4105315490 0.3870533340
H 2.8346807725 -0.5412209492 -1.1903941633
H -0.5537202797 1.5469435382 2.0029686770
H -2.0510266978 0.8448716653 2.2497725116
H 0.8939724574 -1.6086997720 0.2334441992
H 2.2630899289 -2.5235198565 -0.1606712473
H 2.1155048181 -1.8450780227 1.3513380594
H -0.7798223195 -0.1709536166 1.7101046448
25
-34.15043307
O -3.3078583007 -0.3518099727 0.5480566241
O -1.8441149941 -2.0073969491 0.1397595703
N -1.9672328473 1.5403727742 -0.2372000800
N 3.6044925126 -0.8358440764 -0.6659647320
C 1.1684565146 1.0446332415 -0.4943940738
C 0.1082467096 0.2177955398 0.2395309885
C 2.4785715426 1.1657908669 0.2896724500
C -1.2458520269 0.2676289576 -0.4513144135
C 3.1696429470 -0.1592315605 0.5890169973
C -2.2263428078 -0.8414226742 0.1422529421
H 0.7932826990 2.0569151989 -0.6482792542
H 1.3354698371 0.6191303209 -1.4843783659
H -0.0031461344 0.5661490514 1.2686433794
H 0.3791631733 -0.8378892566 0.2841197320
H 3.1579098277 1.8188328559 -0.2597110183
H 2.2713923792 1.6478155935 1.2450536588
H -1.1432114953 0.0569418065 -1.5170262136
H 2.5004236326 -0.8240708177 1.1341470919
H 4.0493855213 0.0280822082 1.2049984895
H -1.5070190112 2.1488324010 0.4454597884
H -2.1837208522 2.0584218679 -1.0901715209
H 4.2094106392 -0.2068158824 -1.2153183037
H 4.1256865653 -1.6973771969 -0.4514692989
H 2.7834959501 -1.0854874671 -1.2358687369
H -2.8845837684 1.1083956166 0.2016053653
Calculate free energies in solution phase (CHCl3) on GFNn-xTB geometries
Note
There are two options available:
Using part1 (prescreening) or
only using part3 and do not calculate part2 (optimization)
The difference between the two approaches is that Part3 applies tighter thresholds in the SCF.
$ censo -inp ensemble.xyz -part1 on -chrg 1 -solvent chcl3 -smgsolv1 cosmors -func r2scan-3c -basis automatic -P 4 -O 2 > censo.out &
______________________________________________________________
| |
| |
| CENSO - Commandline ENSO |
| v 1.0.3 |
| energetic sorting of CREST Conformer Rotamer Ensembles |
| University of Bonn, MCTC |
| Feb 2021 |
| based on ENSO version 2.0.1 |
| F. Bohle and S. Grimme |
| |
|______________________________________________________________|
This program is distributed in the hope that it will be useful,
but WITHOUT ANY WARRANTY; without even the implied warranty of
MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE.
----------------------------------------------------------------------------------------------------
PARAMETERS
----------------------------------------------------------------------------------------------------
The configuration file .censorc is read from /home/bohle/1projects/from_tmp1/CENSO/documentation-calcs/2-part1/.censorc.
Reading conformer rotamer ensemble from: /home/bohle/1projects/from_tmp1/CENSO/documentation-calcs/2-part1/ensemble.xyz.
Reading file: censo_solvents.json
--------------------------------------------------
CRE SORTING SETTINGS
--------------------------------------------------
number of atoms in system: 25
number of considered conformers: 22
number of all conformers from input: 22
charge: 1
unpaired: 0
solvent: chcl3
temperature: 298.15
evaluate at different temperatures: on
temperature range: 273.15, 278.15, 283.15, 288.15, ...
calculate mRRHO contribution: on
consider symmetry for mRRHO contribution: off
cautious checking for error and failed calculations: on
checking the DFT-ensemble using CREST: off
maxthreads: 4
omp: 2
--------------------------------------------------
CRE PRESCREENING - PART1
--------------------------------------------------
part1: on
starting number of considered conformers: 22
program for part1: tm
functional for initial evaluation: r2scan-3c
basis set for initial evaluation: def2-mTZVPP
calculate mRRHO contribution: on
program for mRRHO contribution: xtb
GFN version for mRRHO and/or GBSA_Gsolv: gfn2
Apply constraint to input geometry during mRRHO calculation: on
solvent model applied with xTB: alpb
evaluate at different temperatures: off
threshold g_thr(1) and G_thr(1) for sorting in part1: 3.5
solvent model for Gsolv contribution of part1: cosmors
short-notation:
r2scan-3c + COSMORS[chcl3] + GmRRHO(GFN2[alpb]-bhess) // GFNn-xTB (Input geometry)
END of parameters
------------------------------------------------------------
PATHS of external QM programs
------------------------------------------------------------
The following program paths are used:
xTB: /home/abt-grimme/AK-bin/xtb
TURBOMOLE: /home/abt-grimme/TURBOMOLE.7.5.1/bin/em64t-unknown-linux-gnu_smp
Setup of COSMO-RS:
ctd = BP_TZVPD_FINE_C30_1601.ctd
cdir = "/home/bohle/COSMOlogic/COSMOthermX19/COSMOtherm/CTDATA-FILES"
ldir = "/home/bohle/COSMOlogic/COSMOthermX19/COSMOtherm/CTDATA-FILES"
Using /home/bohle/COSMOlogic/COSMOthermX19/COSMOtherm/DATABASE-COSMO/BP-TZVP-COSMO
as path to the COSMO-RS NORMAL DATABASE.
Using cefine from /home/bohle/bin/cefine
PARNODES for TM or COSMO-RS calculation was set to 2
Using COSMOtherm from /home/bohle/COSMOlogic/COSMOthermX19/COSMOtherm/BIN-LINUX/cosmotherm
----------------------------------------------------------------------------------------------------
Processing data from previous run (enso.json)
----------------------------------------------------------------------------------------------------
INFORMATION: No restart information exists and is created during this run!
----------------------------------------------------------------------------------------------------
CRE PRESCREENING - PART1
----------------------------------------------------------------------------------------------------
program: tm
functional for part1 and 2: r2scan-3c
basis set for part1 and 2: def2-mTZVPP
Solvent: chcl3
solvent model for Gsolv contribution: cosmors
threshold g_thr(1) and G_thr(1): 3.5
starting number of considered conformers: 22
calculate mRRHO contribution: on
program for mRRHO contribution: xtb
GFN version for mRRHO and/or GBSA_Gsolv: gfn2
Apply constraint to input geometry during mRRHO calculation: on
temperature: 298.15
Calculating single-point energies and solvation contribution (G_solv):
The prescreening COSMO-RS is calculated for:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF10, CONF11
CONF12, CONF13, CONF14, CONF15, CONF16, CONF17, CONF18, CONF19, CONF20, CONF21, CONF22
Constructed folders!
Constructed folders!
Starting 22 COSMO-RS-Gsolv calculations.
Running COSMO-RS calculation in CONF1/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF2/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF3/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF4/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF5/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF6/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF7/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF8/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF9/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF10/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF11/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF12/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF13/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF14/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF15/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF16/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF17/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF18/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF19/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF20/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF21/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF22/r2scan-3c/COSMO
Tasks completed!
prescreening COSMO-RS calculation was successful for CONF1/r2scan-3c/COSMO: -0.09391084
prescreening COSMO-RS calculation was successful for CONF2/r2scan-3c/COSMO: -0.09344716
prescreening COSMO-RS calculation was successful for CONF3/r2scan-3c/COSMO: -0.09189166
prescreening COSMO-RS calculation was successful for CONF4/r2scan-3c/COSMO: -0.09269361
prescreening COSMO-RS calculation was successful for CONF5/r2scan-3c/COSMO: -0.09328873
prescreening COSMO-RS calculation was successful for CONF6/r2scan-3c/COSMO: -0.09278491
prescreening COSMO-RS calculation was successful for CONF7/r2scan-3c/COSMO: -0.09457155
prescreening COSMO-RS calculation was successful for CONF8/r2scan-3c/COSMO: -0.09275254
prescreening COSMO-RS calculation was successful for CONF9/r2scan-3c/COSMO: -0.09316162
prescreening COSMO-RS calculation was successful for CONF10/r2scan-3c/COSMO: -0.09116989
prescreening COSMO-RS calculation was successful for CONF11/r2scan-3c/COSMO: -0.09348580
prescreening COSMO-RS calculation was successful for CONF12/r2scan-3c/COSMO: -0.08900263
prescreening COSMO-RS calculation was successful for CONF13/r2scan-3c/COSMO: -0.08998048
prescreening COSMO-RS calculation was successful for CONF14/r2scan-3c/COSMO: -0.09122770
prescreening COSMO-RS calculation was successful for CONF15/r2scan-3c/COSMO: -0.09272288
prescreening COSMO-RS calculation was successful for CONF16/r2scan-3c/COSMO: -0.08509833
prescreening COSMO-RS calculation was successful for CONF17/r2scan-3c/COSMO: -0.08598277
prescreening COSMO-RS calculation was successful for CONF18/r2scan-3c/COSMO: -0.13224220
prescreening COSMO-RS calculation was successful for CONF19/r2scan-3c/COSMO: -0.11057302
prescreening COSMO-RS calculation was successful for CONF20/r2scan-3c/COSMO: -0.13223689
prescreening COSMO-RS calculation was successful for CONF21/r2scan-3c/COSMO: -0.09915029
prescreening COSMO-RS calculation was successful for CONF22/r2scan-3c/COSMO: -0.12536341
--------------------------------------------------
Removing high lying conformers
--------------------------------------------------
CONF# E(GFNn-xTB) ΔE(GFNn-xTB) E [Eh] Gsolv [Eh] Gtot ΔGtot
[a.u.] [kcal/mol] r2scan-3c COSMO-RS-normal [Eh] [kcal/mol]
[r2scan-3c]
CONF1 -34.1599548 0.00 -497.3021467 -0.0939108 -497.3960575 1.82
CONF2 -34.1594484 0.32 -497.3049956 -0.0934472 -497.3984428 0.33
CONF3 -34.1592530 0.44 -497.3064930 -0.0918917 -497.3983847 0.36
CONF4 -34.1590954 0.54 -497.3049058 -0.0926936 -497.3975994 0.86
CONF5 -34.1589250 0.65 -497.3056747 -0.0932887 -497.3989634 0.00 <------
CONF6 -34.1587465 0.76 -497.3049670 -0.0927849 -497.3977519 0.76
CONF7 -34.1578065 1.35 -497.2983644 -0.0945716 -497.3929360 3.78
CONF8 -34.1571241 1.78 -497.3029570 -0.0927525 -497.3957096 2.04
CONF9 -34.1570372 1.83 -497.3008248 -0.0931616 -497.3939864 3.12
CONF10 -34.1563217 2.28 -497.3038986 -0.0911699 -497.3950685 2.44
CONF11 -34.1559251 2.53 -497.3008250 -0.0934858 -497.3943108 2.92
CONF12 -34.1554750 2.81 -497.3045828 -0.0890026 -497.3935854 3.37
CONF13 -34.1554585 2.82 -497.3036749 -0.0899805 -497.3936554 3.33
CONF14 -34.1537285 3.91 -497.3022176 -0.0912277 -497.3934453 3.46
CONF15 -34.1529637 4.39 -497.3009381 -0.0927229 -497.3936610 3.33
CONF16 -34.1514138 5.36 -497.3063638 -0.0850983 -497.3914621 4.71
CONF17 -34.1512255 5.48 -497.3047147 -0.0859828 -497.3906975 5.19
CONF18 -34.1510249 5.60 -497.2564102 -0.1322422 -497.3886524 6.47
CONF19 -34.1509627 5.64 -497.2746319 -0.1105730 -497.3852050 8.63
CONF20 -34.1508506 5.71 -497.2550494 -0.1322369 -497.3872863 7.33
CONF21 -34.1504892 5.94 -497.2896633 -0.0991503 -497.3888135 6.37
CONF22 -34.1504331 5.98 -497.2596139 -0.1253634 -497.3849773 8.78
--------------------------------------------------
Conformers considered further
--------------------------------------------------
Below the g_thr(1) threshold of 3.5 kcal/mol.
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF8, CONF9, CONF10, CONF11, CONF12
CONF13, CONF14, CONF15
--------------------------------------------------
Calculating prescreening G_mRRHO with implicit solvation!
The prescreening G_mRRHO calculation is now performed for:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF8, CONF9, CONF10, CONF11, CONF12
CONF13, CONF14, CONF15
Constructed folders!
Starting 14 G_RRHO calculations.
Running GFN2-xTB mRRHO in CONF1/rrho_part1
Running GFN2-xTB mRRHO in CONF2/rrho_part1
Running GFN2-xTB mRRHO in CONF3/rrho_part1
Running GFN2-xTB mRRHO in CONF4/rrho_part1
Running GFN2-xTB mRRHO in CONF5/rrho_part1
Running GFN2-xTB mRRHO in CONF6/rrho_part1
Running GFN2-xTB mRRHO in CONF8/rrho_part1
Running GFN2-xTB mRRHO in CONF9/rrho_part1
Running GFN2-xTB mRRHO in CONF10/rrho_part1
Running GFN2-xTB mRRHO in CONF11/rrho_part1
Running GFN2-xTB mRRHO in CONF12/rrho_part1
Running GFN2-xTB mRRHO in CONF13/rrho_part1
Running GFN2-xTB mRRHO in CONF14/rrho_part1
Running GFN2-xTB mRRHO in CONF15/rrho_part1
Tasks completed!
The prescreening G_mRRHO calculation @ c1 was successful for CONF1/rrho_part1: 0.18236493
The prescreening G_mRRHO calculation @ c1 was successful for CONF2/rrho_part1: 0.18246714
The prescreening G_mRRHO calculation @ c1 was successful for CONF3/rrho_part1: 0.18235782
The prescreening G_mRRHO calculation @ c1 was successful for CONF4/rrho_part1: 0.18239049
The prescreening G_mRRHO calculation @ c1 was successful for CONF5/rrho_part1: 0.18241907
The prescreening G_mRRHO calculation @ c1 was successful for CONF6/rrho_part1: 0.18225430
The prescreening G_mRRHO calculation @ c1 was successful for CONF8/rrho_part1: 0.18193879
The prescreening G_mRRHO calculation @ c1 was successful for CONF9/rrho_part1: 0.18214342
The prescreening G_mRRHO calculation @ c1 was successful for CONF10/rrho_part1: 0.18217367
The prescreening G_mRRHO calculation @ c1 was successful for CONF11/rrho_part1: 0.18267361
The prescreening G_mRRHO calculation @ c1 was successful for CONF12/rrho_part1: 0.18167483
The prescreening G_mRRHO calculation @ c1 was successful for CONF13/rrho_part1: 0.18145764
The prescreening G_mRRHO calculation @ c1 was successful for CONF14/rrho_part1: 0.18268845
The prescreening G_mRRHO calculation @ c1 was successful for CONF15/rrho_part1: 0.18320604
--------------------------------------------------
* Gibbs free energies of part1 *
--------------------------------------------------
CONF# G(GFNn-xTB) ΔG(GFNn-xTB) E [Eh] Gsolv [Eh] GmRRHO [Eh] Gtot ΔGtot
[a.u.] [kcal/mol] r2scan-3c COSMO-RS-normal GFN2 [Eh] [kcal/mol]
[r2scan-3c] [alpb]-bhess
CONF1 -33.9775899 0.00 -497.3021467 -0.0939108 0.1823649 -497.2136926 1.79
CONF2 -33.9769813 0.38 -497.3049956 -0.0934472 0.1824671 -497.2159757 0.36
CONF3 -33.9768952 0.44 -497.3064930 -0.0918917 0.1823578 -497.2160268 0.32
CONF4 -33.9767049 0.56 -497.3049058 -0.0926936 0.1823905 -497.2152089 0.84
CONF5 -33.9765060 0.68 -497.3056747 -0.0932887 0.1824191 -497.2165444 0.00 <------
CONF6 -33.9764922 0.69 -497.3049670 -0.0927849 0.1822543 -497.2154976 0.66
CONF8 -33.9751853 1.51 -497.3029570 -0.0927525 0.1819388 -497.2137708 1.74
CONF9 -33.9748938 1.69 -497.3008248 -0.0931616 0.1821434 -497.2118430 2.95
CONF10 -33.9741480 2.16 -497.3038986 -0.0911699 0.1821737 -497.2128948 2.29
CONF11 -33.9732515 2.72 -497.3008250 -0.0934858 0.1826736 -497.2116372 3.08
CONF12 -33.9738002 2.38 -497.3045828 -0.0890026 0.1816748 -497.2119106 2.91
CONF13 -33.9740008 2.25 -497.3036749 -0.0899805 0.1814576 -497.2121977 2.73
CONF14 -33.9710400 4.11 -497.3022176 -0.0912277 0.1826885 -497.2107568 3.63
CONF15 -33.9697576 4.91 -497.3009381 -0.0927229 0.1832060 -497.2104549 3.82
Additional global 'fuzzy-threshold' based on the standard deviation of (G_mRRHO):
Std_dev(G_mRRHO) = 0.272 kcal/mol
Fuzzythreshold = 0.309 kcal/mol
Final sorting threshold G_thr(1) = 3.500 + 0.309 = 3.809 kcal/mol
Spearman correlation coefficient between (E + Solv) and (E + Solv + mRRHO) = 0.908
--------------------------------------------------
Conformers considered further
--------------------------------------------------
Considered CONF14 because of increased fuzzythr.
These conformers are below the 3.809 kcal/mol G_thr(1) threshold.
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF8, CONF9, CONF10, CONF11, CONF12
CONF13, CONF14
Calculating Boltzmann averaged free energy of ensemble on input geometries (not DFT optimized)!
temperature /K: avE(T) /a.u. avGmRRHO(T) /a.u. avGsolv(T) /a.u. avG(T) /a.u.
----------------------------------------------------------------------------------------------------
298.15 -497.3054054 0.1823775 -0.0928881 -497.2159160 <<==part1==
----------------------------------------------------------------------------------------------------
Calculating unbiased GFNn-xTB energy
Constructed folders!
Starting 13 xTB - single-point calculations.
gfn2-xTB energy for CONF1/GFN_unbiased = -34.1515395
gfn2-xTB energy for CONF2/GFN_unbiased = -34.1509941
gfn2-xTB energy for CONF3/GFN_unbiased = -34.1515326
gfn2-xTB energy for CONF4/GFN_unbiased = -34.1508491
gfn2-xTB energy for CONF5/GFN_unbiased = -34.1514366
gfn2-xTB energy for CONF6/GFN_unbiased = -34.1512893
gfn2-xTB energy for CONF8/GFN_unbiased = -34.1493072
gfn2-xTB energy for CONF9/GFN_unbiased = -34.1491025
gfn2-xTB energy for CONF10/GFN_unbiased = -34.1491972
gfn2-xTB energy for CONF11/GFN_unbiased = -34.1487085
gfn2-xTB energy for CONF12/GFN_unbiased = -34.1484725
gfn2-xTB energy for CONF13/GFN_unbiased = -34.1480733
gfn2-xTB energy for CONF14/GFN_unbiased = -34.1461266
Tasks completed!
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>END of Part1<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
Ran part1 in 277.0194 seconds
Part : #conf time
--------------------------------------------------
Input : 22 -
Part1_initial_sort : 14 -
Part1_all : 14 277.02s
--------------------------------------------------
All parts : 277.02s
CENSO all done!
$CENSO global configuration file: .censorc
$VERSION:1.0.3
ORCA: /home/$USER/orca_4_2_1_linux_x86-64_openmpi216
ORCA version: 4.2.1
GFN-xTB: /home/$USER/bin/xtb
CREST: /home/$USER/bin/crest
mpshift: /home/$USER/TURBOMOLE.7.5/bin/em64t-unknown-linux-gnu/mpshift
escf: /home/$USER/TURBOMOLE.7.5/bin/em64t-unknown-linux-gnu/escf
#COSMO-RS
ctd = BP_TZVPD_FINE_C30_1601.ctd cdir = "/home/$USER/COSMOlogic/COSMOthermX19/COSMOtherm/CTDATA-FILES" ldir = "/home/$USER/COSMOlogic/COSMOthermX19/COSMOtherm/CTDATA-FILES"
cosmothermversion: 19
$ENDPROGRAMS
$CRE SORTING SETTINGS:
$GENERAL SETTINGS:
nconf: all # ['all', 'number e.g. 10 up to all conformers']
charge: 0 # ['number e.g. 0']
unpaired: 0 # ['number e.g. 0']
solvent: gas # ['gas', 'acetone', 'chcl3', 'acetonitrile', 'ch2cl2', 'dmso', 'h2o', 'methanol', 'thf', '...']
prog_rrho: xtb # ['xtb', 'prog']
temperature: 298.15 # ['temperature in K e.g. 298.15']
trange: [273.15, 378.15, 5] # ['temperature range [start, end, step]']
multitemp: on # ['on', 'off']
evaluate_rrho: on # ['on', 'off']
consider_sym: off # ['on', 'off']
bhess: on # ['on', 'off']
imagthr: automatic # ['automatic or e.g., -100 # in cm-1']
sthr: automatic # ['automatic or e.g., 50 # in cm-1']
scale: automatic # ['automatic or e.g., 1.0 ']
rmsdbias: off # ['on', 'off']
sm_rrho: alpb # ['alpb', 'gbsa']
check: on # ['on', 'off']
prog: tm # ['tm', 'orca']
func: r2scan-3c # ['pbe', 'b97-d', 'pbeh-3c', 'tpss', 'b97-d3', 'r2scan-3c', 'b97-3c']
basis: automatic # ['automatic', 'def2-mSVP', 'def2-mTZVP', 'def2-mTZVP', 'def2-TZVP', '...']
maxthreads: 2 # ['number of threads e.g. 2']
omp: 2 # ['number cores per thread e.g. 4']
cosmorsparam: automatic # ['automatic', '12-fine', '12-normal', '13-fine', '13-normal', '14-fine', '...']
$PART0 - CHEAP-PRESCREENING - SETTINGS:
part0: off # ['on', 'off']
func0: b97-d # ['pbeh-3c', 'b97-3c', 'b97-d3', 'pbe', 'r2scan-3c', 'tpss', 'b97-d']
basis0: def2-SV(P) # ['automatic', 'def2-mSVP', 'def2-mTZVP', 'def2-mTZVP', 'def2-TZVP', '...']
part0_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
part0_threshold: 4.0 # ['number e.g. 4.0']
$PART1 - PRESCREENING - SETTINGS:
# func and basis is set under GENERAL SETTINGS
part1: off # ['on', 'off']
smgsolv1: cosmors # ['alpb_gsolv', 'dcosmors', 'cosmo', 'smd', 'cosmors-fine', 'cpcm', 'gbsa_gsolv', '...']
part1_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
part1_threshold: 3.5 # ['number e.g. 5.0']
$PART2 - OPTIMIZATION - SETTINGS:
# func and basis is set under GENERAL SETTINGS
part2: off # ['on', 'off']
opt_limit: 2.5 # ['number e.g. 4.0']
sm2: dcosmors # ['cosmo', 'cpcm', 'default', 'smd', 'dcosmors']
smgsolv2: cosmors # ['cosmors-fine', 'gbsa_gsolv', 'cosmo', 'cpcm', 'smd_gsolv', 'alpb_gsolv', 'smd', '...']
part2_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
ancopt: on # ['on']
hlow: 0.01 # ['lowest force constant in ANC generation, e.g. 0.01']
opt_spearman: on # ['on', 'off']
part2_threshold: 99 # ['Boltzmann sum threshold in %. e.g. 95 (between 1 and 100)']
optlevel2: automatic # ['crude', 'sloppy', 'loose', 'lax', 'normal', 'tight', 'vtight', 'extreme', '...']
optcycles: 8 # ['number e.g. 5 or 10']
spearmanthr: -4.0 # ['value between -1 and 1, if outside set automatically']
radsize: 10 # ['number e.g. 8 or 10']
crestcheck: off # ['on', 'off']
$PART3 - REFINEMENT - SETTINGS:
part3: off # ['on', 'off']
prog3: prog # ['tm', 'orca', 'prog']
func3: pw6b95 # ['dsd-blyp', 'pw6b95', 'b97-d3', 'r2scan-3c', 'pbe0', 'wb97x']
basis3: def2-TZVPD # ['SVP', 'SV(P)', 'TZVP', 'TZVPP', 'QZVP', 'QZVPP', 'def2-SV(P)', 'def2-mSVP', '...']
smgsolv3: cosmors # ['alpb_gsolv', 'dcosmors', 'cosmo', 'smd', 'cosmors-fine', 'cpcm', 'gbsa_gsolv', '...']
part3_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
part3_threshold: 99 # ['Boltzmann sum threshold in %. e.g. 95 (between 1 and 100)']
$NMR PROPERTY SETTINGS:
$PART4 SETTINGS:
part4: off # ['on', 'off']
couplings: on # ['on', 'off']
progJ: prog # ['tm', 'orca', 'adf', 'prog']
funcJ: pbe0 # ['tpss', 'pbe0', 'pbeh-3c']
basisJ: def2-TZVP # ['SVP', 'SV(P)', 'TZVP', 'TZVPP', 'QZVP', 'QZVPP', 'def2-SV(P)', 'def2-mSVP', '...']
sm4J: default # ['dcosmors', 'smd', 'cosmo', 'cpcm']
shieldings: on # ['on', 'off']
progS: prog # ['tm', 'orca', 'adf', 'prog']
funcS: pbe0 # ['dsd-blyp', 'pbeh-3c', 'tpss', 'pbe0', 'kt2']
basisS: def2-TZVP # ['SVP', 'SV(P)', 'TZVP', 'TZVPP', 'QZVP', 'QZVPP', 'def2-SV(P)', 'def2-mSVP', '...']
sm4S: default # ['dcosmors', 'smd', 'cosmo', 'cpcm']
reference_1H: TMS # ['TMS']
reference_13C: TMS # ['TMS']
reference_19F: CFCl3 # ['CFCl3']
reference_29Si: TMS # ['TMS']
reference_31P: TMP # ['TMP', 'PH3']
1H_active: on # ['on', 'off']
13C_active: on # ['on', 'off']
19F_active: off # ['on', 'off']
29Si_active: off # ['on', 'off']
31P_active: off # ['on', 'off']
resonance_frequency: 300.0 # ['MHz number of your experimental spectrometer setup']
$OPTICAL ROTATION PROPERTY SETTINGS:
$PART5 SETTINGS:
optical_rotation: off # ['on', 'off']
funcOR: pbe # ['functional for opt_rot e.g. pbe']
funcOR_SCF: r2scan-3c # ['functional for SCF in opt_rot e.g. r2scan-3c']
basisOR: def2-SVPD # ['basis set for opt_rot e.g. def2-SVPD']
frequency_optical_rot: [589.0] # ['list of freq in nm to evaluate opt rot at e.g. [589, 700]']
$END CENSORC
25
-34.15995484
O -2.7404108553 -0.9210263756 0.0505546155
O -0.6532148483 -1.7303331963 0.1381884568
N -2.1811530948 1.4662136641 -0.3280604715
N 1.9999042791 -1.6115147219 -0.1737452561
C 1.2832974514 1.7494376339 0.2478643443
C 0.0382242755 1.0547713107 0.8044626435
C 2.1972182535 0.8461961224 -0.5852172042
C -0.9624505804 0.6276788621 -0.2713633850
C 2.7304045776 -0.3664138808 0.1827513585
C -1.5037598584 -0.8390454546 0.0048901917
H 1.8561477225 2.1402514306 1.0892838382
H 0.9852377848 2.6009114549 -0.3642611073
H -0.4518270956 1.7282098745 1.5089809935
H 0.3322545004 0.1775252924 1.3825123935
H 1.6820235078 0.5059218506 -1.4832357051
H 3.0380251642 1.4546200515 -0.9165284484
H -0.4868586610 0.6089411942 -1.2530945560
H 2.6388482788 -0.2021893439 1.2562535001
H 3.7853986884 -0.5188774904 -0.0473087348
H -2.2380899515 2.1426216511 0.4407329422
H -2.3218532286 1.9466721688 -1.2205963738
H 2.3394829642 -2.3992859458 0.3882351555
H 0.9350292280 -1.5547588441 -0.0165809300
H 2.1475890354 -1.8368760327 -1.1636223077
H -2.9328940343 0.6971914121 -0.1921053926
25
-34.15944839
O -2.5910945174 -0.7526707388 0.4737473250
O -0.5258609320 -1.6055473558 0.4259946366
N -2.1883047577 1.2429283468 -0.9276187078
N 2.0221224751 -1.5482527990 -0.3635876013
C 1.0959612422 0.9269941773 1.0928268792
C -0.1118183900 1.5287127427 0.3646185610
C 2.3883955950 0.8232150730 0.2771806877
C -0.9137052232 0.5859776609 -0.5447048684
C 2.3984857923 -0.1999516224 -0.8542391926
C -1.3796771114 -0.7342464594 0.2052799197
H 0.8202702309 -0.0346561349 1.5230052762
H 1.3150554091 1.5881707649 1.9329336393
H 0.2252898792 2.3837647705 -0.2251981715
H -0.7970460540 1.8894152433 1.1356879771
H 2.6174364896 1.7989347868 -0.1517789136
H 3.1942064704 0.5825472961 0.9724285437
H -0.3499369029 0.3059081761 -1.4335569624
H 3.4048136146 -0.2402683769 -1.2749539435
H 1.7122153210 0.0852818535 -1.6492945642
H -2.2438155060 2.2226460139 -0.6308218074
H -2.4094139539 1.1700691932 -1.9248068681
H 0.9927667520 -1.5675551657 -0.0428313977
H 2.1377830502 -2.2542373686 -1.0973459319
H 2.6054467969 -1.8083924261 0.4393088723
H -2.8796747565 0.6310313638 -0.3580585669
25
-34.15925304
O -2.6152400314 -1.0465052003 0.2378304729
O -0.5362082855 -1.4663149505 -0.4593671015
N -2.4557715395 1.4039918868 0.4362491058
N 2.1447268012 -1.5911295291 -0.0908488427
C 1.1849792140 1.1291603967 -0.7123125066
C 0.0549239500 1.3190958463 0.3109988160
C 2.5516538580 0.8614163999 -0.0796419105
C -1.2770304521 0.8481909298 -0.2699298296
C 2.6278492228 -0.4408272994 0.7143702153
C -1.4798499370 -0.7275247362 -0.1594413730
H 1.2638776944 2.0288103522 -1.3231621409
H 0.9216680773 0.3153070715 -1.3864974308
H -0.0132519646 2.3752873127 0.5749134480
H 0.2604586805 0.7576942571 1.2223482952
H 3.2969117377 0.8423750312 -0.8755852689
H 2.8115908685 1.6810094994 0.5898815386
H -1.3353089850 1.1014685130 -1.3310936133
H 2.0240292821 -0.3842292575 1.6193674755
H 3.6638250279 -0.6160286136 1.0070988283
H -2.2160156909 1.7709567167 1.3631094761
H -2.9723151833 2.1102297821 -0.0929227069
H 2.6253531272 -1.6216026098 -0.9974699479
H 2.3064470721 -2.4761725884 0.4009650983
H 1.0998172314 -1.5111491960 -0.2664030290
H -3.0342449423 0.4895941432 0.5473101358
25
-34.15909539
O -2.5859531813 -0.8098009525 0.4685129327
O -0.4816245052 -1.5436026390 0.3102868940
N -2.3339104715 1.3007756781 -0.7836622820
N 2.0492564794 -1.5871472251 -0.5900906398
C 1.1052196056 0.9870949301 0.9271737021
C -0.2140772115 1.5925038037 0.4451868362
C 2.1667497593 0.8464686753 -0.1771749747
C -1.0154302176 0.6861215046 -0.4984780600
C 2.9073785151 -0.4823039670 -0.0859386105
C -1.3840762559 -0.7076318190 0.1707141950
H 0.8920163207 0.0201435075 1.3815317896
H 1.4930774855 1.6249540613 1.7213050551
H -0.0067258320 2.5368714102 -0.0630175124
H -0.8282372023 1.7972548662 1.3249938991
H 1.7135886317 0.9350726379 -1.1636100949
H 2.8915967677 1.6540129472 -0.0882838850
H -0.4677225963 0.5007855037 -1.4227063242
H 3.1688824777 -0.6847294678 0.9524589517
H 3.8237876908 -0.4492687854 -0.6762385697
H -2.5870605138 1.3036833133 -1.7755142934
H -2.9689761577 0.6014426661 -0.2449843511
H 1.9723283573 -1.5417442554 -1.6118847971
H 2.4360322309 -2.5007606995 -0.3327999527
H 1.0548817542 -1.5294780724 -0.1881461707
H -2.4296444696 2.2452869585 -0.3976639893
25
-34.15892503
O -2.4565204045 -0.8423454324 0.7070928712
O -0.5980888346 -1.6084799899 -0.2741310513
N -2.2081439307 1.5499049911 0.0797136762
N 2.0665657616 -1.5738488309 0.1665385721
C 0.8489992162 1.1021214800 0.5754910780
C 0.1319038004 1.1643519245 -0.7806607655
C 2.3611287197 0.8895018711 0.4504901307
C -1.3084663819 0.6750096988 -0.7114456668
C 2.7727042231 -0.3622083435 -0.3226992137
C -1.4583757981 -0.7537800183 -0.0274928276
H 0.4155006230 0.3036523983 1.1797441237
H 0.7020515096 2.0340597672 1.1228295970
H 0.6304477123 0.5210588124 -1.5026223849
H 0.1602968624 2.1778026423 -1.1819014190
H 2.8058653217 1.7473421346 -0.0537220547
H 2.7810957945 0.8489865537 1.4562868383
H -1.7106345304 0.5638398621 -1.7216356404
H 3.8478441803 -0.5048945341 -0.2050768697
H 2.5630334022 -0.2516058916 -1.3860041489
H -2.8269415029 2.1396743466 -0.4823076953
H -2.7642911303 0.7978509993 0.6164252281
H 2.4375926491 -2.4160896797 -0.2859521821
H 2.1793786728 -1.6693140839 1.1823811296
H 1.0245787819 -1.5392035408 -0.0507409777
H -1.6858363886 2.1297513428 0.7452520673
25
-34.15874648
O -2.3605981473 -0.8116042758 0.8659409274
O -0.5843274484 -1.6230194998 -0.2244865945
N -2.2109877661 1.5364887021 0.0742080589
N 2.0757147084 -1.4672125485 -0.4830865528
C 0.8597549995 1.2262238823 0.4517476395
C 0.0874032105 1.1513231513 -0.8771132949
C 2.3512317789 0.8985689618 0.2982329522
C -1.3361657088 0.6347191585 -0.7132778341
C 2.6824356905 -0.5712975375 0.5355813092
C -1.4234260659 -0.7566236862 0.0528412632
H 0.4221004807 0.5356789162 1.1753502177
H 0.7733707657 2.2309740830 0.8650565898
H 0.5827203837 0.4802075195 -1.5756071724
H 0.0631682297 2.1335262804 -1.3514203271
H 2.7003072358 1.2129278451 -0.6855204577
H 2.9172351560 1.4667500260 1.0360745366
H -1.7833398578 0.4616073570 -1.6955611361
H 2.3142140534 -0.8770131145 1.5154931358
H 3.7656889000 -0.7005750162 0.5151167425
H -1.6671444199 2.1735125324 0.6661186569
H -2.8827343086 2.0695764162 -0.4835506438
H 2.3643871815 -1.1879653946 -1.4273853498
H 2.3725094498 -2.4360036549 -0.3237631449
H 1.0071911744 -1.4659435853 -0.4227446705
H -2.7090441201 0.8070062032 0.6949537917
25
-34.15780648
O -0.6998712600 -1.7595821198 0.3315266042
O -2.7564980797 -0.8922468771 0.1127087169
N -2.0912345729 1.3897515697 -0.6207661035
N 1.9218668343 -1.5538051846 -0.0911139226
C 1.2724611881 1.7665596691 0.2614262584
C -0.0126245071 1.0963347719 0.7686474620
C 2.5091659413 0.8642624497 0.2746224048
C -0.9098130242 0.5407794733 -0.3415020783
C 2.4929985644 -0.3196472927 -0.6935656161
C -1.5185682699 -0.8616584773 0.0830057999
H 1.4961397665 2.6156227165 0.9073501428
H 1.1219744652 2.1603919352 -0.7434483203
H -0.5894557289 1.8123867185 1.3552290309
H 0.2456564204 0.2915541326 1.4587568868
H 3.3629575544 1.4905499194 0.0148013745
H 2.6790096911 0.5030846620 1.2896168914
H -0.3423105707 0.3842564461 -1.2596402044
H 3.5203242755 -0.5424656780 -0.9864282797
H 1.9321769951 -0.0751719518 -1.5947809502
H -2.8797605313 0.6907420298 -0.3760235139
H -2.1503924665 2.2120232522 -0.0104200923
H 2.3803263882 -1.7503210865 0.8055549093
H 0.8561607870 -1.5268128372 0.0892049642
H 2.0800598725 -2.3543999415 -0.7123089418
H -2.1774862389 1.6805852373 -1.5986092278
25
-34.15712409
O -0.5169415866 -1.4677953372 -0.4155166026
O -2.6718105717 -1.1029442546 0.0532698902
N -2.5041882719 1.3222450456 0.5356135683
N 2.0547500815 -1.4660199090 0.3952674841
C 1.1460672076 1.2350620947 -0.5905524477
C 0.0199991306 1.2471137062 0.4571994904
C 2.5324119546 0.9449216441 0.0010319703
C -1.3028765754 0.8219664596 -0.1753086903
C 2.9759584940 -0.4928036507 -0.2449354813
C -1.5027299259 -0.7570468322 -0.1889883845
H 1.1567284127 2.2014747982 -1.0932584224
H 0.9206483903 0.4869950226 -1.3509226784
H -0.0730555237 2.2557886152 0.8619704582
H 0.2503077610 0.5750867956 1.2847618985
H 3.2724883861 1.5930094341 -0.4660504111
H 2.5411017268 1.1667575642 1.0680525607
H -1.3439398378 1.1490834108 -1.2170635960
H 3.9809942022 -0.6416689206 0.1522940223
H 2.9906906878 -0.6917352586 -1.3167852841
H -3.1147358709 0.4302326474 0.4978236161
H -2.3055244586 1.5552316017 1.5147727778
H 1.0589542040 -1.3722826387 0.0237179616
H 2.3537713222 -2.4274392985 0.2004063016
H 2.0384435533 -1.3281897829 1.4119124615
H -2.9671283162 2.1123945703 0.0792841186
25
-34.15703719
O -0.4840204654 -1.5908390855 -0.1977054306
O -2.6424172363 -1.0172969494 -0.2313294170
N -2.3824391815 1.4118161237 0.1129756179
N 2.0591606020 -1.4080559447 0.6444993748
C 1.3255949204 1.5932768744 0.1316443482
C 0.0292286137 1.1405631347 0.8180158808
C 1.9611296764 0.5914767293 -0.8382657525
C -1.0774245534 0.7702791240 -0.1744053472
C 2.8498636045 -0.4645206094 -0.1922799795
C -1.4218007565 -0.7850501416 -0.2021503198
H 2.0511714191 1.8626404605 0.8996124027
H 1.1017826165 2.5033885185 -0.4265728164
H -0.3083479528 1.9763847128 1.4332069953
H 0.2033356421 0.3027453082 1.4918974312
H 1.1939462785 0.0852292002 -1.4240400771
H 2.5871303840 1.1474547523 -1.5362113415
H -0.7775416723 1.0312499156 -1.1919311777
H 3.6205966803 0.0055444316 0.4194317100
H 3.3361661233 -1.0318279746 -0.9856062704
H -2.6284628403 2.1753990573 -0.5216811303
H -3.0325627338 0.5540981187 -0.0284710758
H 2.4910941867 -2.3367192092 0.6680136974
H 1.9781398838 -1.0663385144 1.6071566991
H 1.0702227114 -1.5059322479 0.2455456256
H -2.4592842029 1.7302943098 1.0848423546
25
-34.15632171
O -0.6377330316 -1.5602972801 -0.5925407457
O -1.9438226066 -0.8962122015 1.0968963302
N -2.0691310397 1.5115608178 0.5520085904
N 2.0161394685 -1.6864701960 -0.6808935380
C 1.0245208700 1.5678424724 0.3114580637
C 0.0268814843 1.3822638313 -0.8376423667
C 1.7125418505 0.2918072015 0.8062089597
C -1.3255436976 0.7602094261 -0.4861917907
C 2.5798867778 -0.3735189736 -0.2699515146
C -1.2821941821 -0.7269243894 0.0579815663
H 0.5483025277 2.0449086772 1.1692286093
H 1.7966540418 2.2561425086 -0.0359536522
H 0.4649670843 0.7632830938 -1.6196837604
H -0.1661732017 2.3619672870 -1.2796881790
H 2.3322565126 0.5621906956 1.6605054657
H 0.9655697921 -0.4140620317 1.1690705974
H -1.9201614836 0.7061105980 -1.4020292431
H 3.5936748701 -0.5312821922 0.0979019662
H 2.6369135707 0.2686143194 -1.1490685384
H -2.9086496435 1.9840429736 0.2094100180
H -2.3313717130 0.6888337551 1.2084854943
H 2.1820919942 -2.3913736082 0.0460188498
H 0.9581287874 -1.6201065231 -0.7926007692
H 2.4297604389 -2.0163864988 -1.5591512145
H -1.4715716162 2.1835379113 1.0453989384
25
-34.15592507
O -2.1033969149 -0.8344078612 0.9487088877
O -0.5699612374 -1.6680274471 -0.4517081377
N -1.9569275917 1.5465619838 0.2627686808
N 2.0837337252 -1.3633283870 -0.5773762363
C 1.1075416509 1.5419562581 -0.0103918263
C 0.1209945172 1.1686244794 -1.1251953409
C 1.4352918321 0.4570859531 1.0214669735
C -1.2503428799 0.6557309102 -0.6867527508
C 2.5057372966 -0.5407868185 0.5902823997
C -1.2948162683 -0.7745608298 0.0075959523
H 0.7274518351 2.4141375453 0.5251162579
H 2.0330446007 1.8691474761 -0.4864359338
H 0.5312965142 0.4086384701 -1.7869917742
H -0.0324587227 2.0621064981 -1.7347668750
H 1.8169519144 0.9463147999 1.9183948838
H 0.5367641743 -0.0848292604 1.3149274400
H -1.8634330878 0.5270669166 -1.5839659846
H 2.7015999487 -1.2099014501 1.4282318085
H 3.4296803314 -0.0167769980 0.3414844052
H -1.2992225514 2.1272618717 0.7958603647
H -2.6755557698 2.1370116562 -0.1630903688
H 2.2813676133 -0.8782194673 -1.4586367594
H 2.5735263109 -2.2635096262 -0.5875520766
H 1.0325521121 -1.5556317222 -0.5353874345
H -2.3935764356 0.8063761382 0.9145561672
25
-34.15547499
O -2.3288681540 -0.9689367039 0.5974852723
O -0.4008216845 -1.3559960891 -0.4559450313
N -2.6333104330 1.3901145964 0.0148791472
N 2.2688734235 -1.7836122490 -0.3135245436
C 1.2045970266 1.2147830910 -0.3207104722
C -0.1766849998 1.7049883679 0.1388725454
C 1.9421007389 0.4104322018 0.7578355455
C -1.3241367595 0.8966054975 -0.4749078790
C 2.9601024995 -0.5430248114 0.1410496733
C -1.3285933693 -0.6410895869 -0.0712604222
H 1.8194085679 2.0702167267 -0.5988277282
H 1.0710671602 0.6065579215 -1.2146344993
H -0.3076575365 2.7461839050 -0.1612866699
H -0.2378627475 1.6551531995 1.2275331138
H 2.4468113264 1.0960018700 1.4370167059
H 1.2297127637 -0.1683684162 1.3468389979
H -1.2704988015 0.9491259299 -1.5640455987
H 3.7310656000 -0.8072491298 0.8640696447
H 3.4371256948 -0.0632845904 -0.7133472266
H -2.5432643621 2.1520849835 0.6930123940
H -3.2834926874 1.6599628672 -0.7262042831
H 2.7178100064 -2.1982719631 -1.1361624210
H 2.2482990815 -2.4828183187 0.4368503414
H 1.2567750610 -1.5723871751 -0.5319861416
H -2.9757946227 0.4721100866 0.5082732629
25
-34.15545846
O -2.3777023451 -0.9177733836 0.6489622803
O -0.4009151018 -1.4460423055 -0.2363506290
N -2.5928675119 1.3702602408 -0.1958788709
N 2.2705863538 -1.5552728060 -0.6959671399
C 1.2626408325 1.2650767695 -0.1999508619
C -0.1676900608 1.6997857440 0.1433859727
C 1.8576282855 0.3222355626 0.8537184709
C -1.2484356318 0.8255651567 -0.5054674146
C 2.9084111559 -0.6168744122 0.2764230940
C -1.3258577703 -0.6732783629 0.0271901706
H 1.8935551201 2.1501893106 -0.2772016053
H 1.2458750287 0.7908735642 -1.1798122422
H -0.3171466901 2.7186794233 -0.2201486160
H -0.2967658760 1.7007241110 1.2272150872
H 2.3089550370 0.9081814099 1.6535209776
H 1.0686425282 -0.2818531316 1.3016188748
H -1.0876822336 0.7817119785 -1.5847014431
H 3.3524799930 -1.1924945383 1.0873745985
H 3.6963304583 -0.0519426265 -0.2212122945
H -2.9913275573 0.5162412164 0.3615908966
H -2.5589278358 2.2025641690 0.3999047288
H 2.6187881911 -2.5117770389 -0.5815509942
H 1.2217357961 -1.5619680721 -0.5321789754
H 2.4345460024 -1.2660811775 -1.6650051770
H -3.1696789754 1.5558016135 -1.0193997348
25
-34.15372845
O -0.8759513206 -1.0507714945 0.7395130487
O -1.0471783147 -1.2210634731 -1.4857197061
N -2.0100713682 1.1638582833 0.9733772433
N 1.7020599065 -1.7447966827 0.8365171560
C 0.9224701988 1.3977552196 0.1299066020
C -0.3034955554 1.7474745141 -0.7290850754
C 1.8313865205 0.3371413309 -0.5152183962
C -1.5397536849 0.8954837973 -0.4124258872
C 2.5286316072 -0.5462459399 0.5204181162
C -1.1337894558 -0.6241211220 -0.4224393440
H 0.5999134506 1.0447379113 1.1088504940
H 1.5075448530 2.3032957331 0.2891927248
H -0.0603736240 1.6138572758 -1.7832639404
H -0.5689030950 2.7975685720 -0.5933476217
H 1.2520636483 -0.2949117808 -1.1892484363
H 2.5821478388 0.8362393886 -1.1261103610
H -2.3194229349 1.0797505602 -1.1497937252
H 2.6951475316 0.0179430508 1.4383444706
H 3.4942793529 -0.8824584649 0.1428332329
H -1.6366862386 2.0406897131 1.3566364027
H -3.0319635647 1.1643591957 1.0502533535
H 2.0123953855 -2.1956055335 1.7033301803
H 0.6617699349 -1.4988916853 0.9180060433
H 1.7620357216 -2.4244612968 0.0700935319
H -1.6155014744 0.3110635636 1.4848864268
25
-34.15296366
O -0.9806508876 -0.9847570656 0.9016095316
O -1.0342961735 -1.4975369515 -1.2790158618
N -2.0600699488 1.3003330160 0.6598555012
N 1.6483681534 -1.4770759046 1.0193012553
C 0.8930197778 1.4409262402 0.3162871289
C -0.1018287155 1.4892580056 -0.8622608316
C 2.2127299908 0.7309357778 -0.0043863659
C -1.4370888955 0.7837090011 -0.5889439421
C 2.1074033787 -0.7768274559 -0.2134156979
C -1.1447588094 -0.7374998177 -0.3265126418
H 0.4424872625 0.9591479756 1.1822965353
H 1.1476108940 2.4604379775 0.6056686014
H 0.3344753521 1.0203527521 -1.7446629435
H -0.3104542151 2.5259312846 -1.1308040482
H 2.6400205950 1.1617275571 -0.9101356051
H 2.9157320587 0.9267783500 0.8074716570
H -2.0965188879 0.8923814409 -1.4486238017
H 3.0903725751 -1.1592546293 -0.4905246456
H 1.4081882505 -1.0149227036 -1.0156071158
H -3.0831759004 1.3363322990 0.6000761811
H -1.7780292676 0.5566802205 1.3629478428
H 0.5935972201 -1.3552723672 1.1310943683
H 1.8230458958 -2.4842696303 0.9418471118
H 2.1352867113 -1.1124232113 1.8467861942
H -1.6994055067 2.2241075607 0.9291898687
25
-34.15141381
O -1.3943785432 -1.0307567325 0.9025777408
O -0.4225106153 -1.2766835982 -1.0942963862
N -2.1382332497 1.3072100696 0.6820859684
N 1.8303685962 -1.9710495392 0.0707587666
C 0.9607624662 1.6168605590 0.1707894371
C -0.1959862014 1.7114505222 -0.8393306352
C 1.9799050629 0.5013843169 -0.1348429740
C -1.4247901716 0.8500833461 -0.5368218294
C 1.9371661639 -0.6960029627 0.8273160062
C -1.0467465663 -0.6453837331 -0.2300826075
H 0.5821582777 1.4936196592 1.1855414824
H 1.4853467761 2.5717069600 0.1437592453
H 0.1684852019 1.4328271659 -1.8279890567
H -0.5295558410 2.7487716363 -0.9068101296
H 1.7990108513 0.1423863706 -1.1479579769
H 2.9805578856 0.9303497506 -0.1217137710
H -2.0885910031 0.8601550715 -1.4024758281
H 1.0694995988 -0.6377417602 1.4853144468
H 2.8334350246 -0.7245407799 1.4470296049
H -1.6564311817 2.0816802979 1.1506799283
H -3.1156981590 1.5603768219 0.5197984898
H 2.6238854975 -2.0888513426 -0.5678832194
H 1.7803639034 -2.7755595157 0.7040003308
H 0.9218328585 -1.9204346441 -0.4990801476
H -2.0745975566 0.3957317695 1.2728068192
25
-34.15122546
O -1.4776298354 -0.9779162840 0.9689911790
O -0.4925234496 -1.3628551244 -1.0036507954
N -2.1639329473 1.3683833598 0.5828976748
N 1.8796097534 -1.9018199765 -0.0324662357
C 0.9212842989 1.5787487906 0.2022882964
C -0.1575522111 1.5949891498 -0.8983466380
C 2.0807266384 0.5903317386 -0.0312863938
C -1.4336452672 0.8124987799 -0.5833640723
C 2.0372506913 -0.6914975753 0.8153963432
C -1.1077721831 -0.6712596326 -0.1784565029
H 0.4772061571 1.3762504059 1.1769048440
H 1.3491352329 2.5796760629 0.2516191005
H 0.2519128515 1.1866703660 -1.8221067218
H -0.4443786804 2.6264492546 -1.1098247151
H 2.1069094344 0.3214704229 -1.0870338715
H 3.0133169149 1.1098047495 0.1839278312
H -2.0690635873 0.7917157945 -1.4696202320
H 1.1994207751 -0.6693189818 1.5132782369
H 2.9569334194 -0.7921592267 1.3922309558
H -1.6800036187 2.1679532791 1.0053931771
H -3.1334683512 1.6233161070 0.3791665015
H 2.6232513605 -1.9579134554 -0.7362257317
H 1.8912989959 -2.7525757838 0.5392212719
H 0.9273933706 -1.8444066710 -0.5257136187
H -2.1333692549 0.5050567581 1.2388793063
25
-34.15102491
O -2.7007099152 -1.6852260154 -0.4665848832
O -3.5578446821 0.0350414384 0.6950996250
N -1.7249631955 1.6316108345 0.4298941943
N 4.8503432460 -0.2765578553 0.0049566510
C 1.1047501969 0.4501828237 -0.3379055216
C -0.1445140391 -0.2886936630 0.1459845288
C 2.3741940866 -0.2828979224 0.0965187068
C -1.4339016427 0.3793165613 -0.3028003047
C 3.6160850376 0.4539254738 -0.3951671902
C -2.6934727541 -0.5488189533 -0.0039847520
H 1.1202182595 1.4641232269 0.0631636060
H 1.0744507790 0.5160807326 -1.4261342120
H -0.1466884742 -0.3764861277 1.2347858649
H -0.1614173698 -1.3033571375 -0.2567289346
H 2.3826839006 -0.3550960319 1.1849100754
H 2.3458274825 -1.2937888279 -0.3120785959
H -1.4193517541 0.5529473168 -1.3796978179
H 3.6476836700 1.4575725004 0.0303995802
H 3.5908364898 0.5356651831 -1.4823627548
H -1.7888195298 2.4639789876 -0.1581990321
H -2.7206446817 1.3385249836 0.8116492773
H 5.6915565940 0.2174183391 -0.3259194480
H 4.8962264202 -0.3605243899 1.0314916029
H 4.8443782262 -1.2261178281 -0.3963638456
H -1.0847775954 1.7904079765 1.2125175138
25
-34.15096274
O -3.0664396656 -0.6811514259 0.4732552569
O -1.1830540829 -1.6357953168 -0.2743493731
N -2.5677177347 1.6455787239 -0.0357930901
N 3.4161836207 -1.4947231742 0.0043983586
C 1.0526738400 0.4198619079 -0.3271341828
C -0.1392962419 1.1756334823 0.2601668795
C 2.3301787116 0.6870764296 0.4694516416
C -1.4374925629 0.7826136622 -0.4432938226
C 3.5433351448 -0.0155439223 -0.1263696890
C -1.9232732343 -0.6849210780 -0.0466890927
H 1.1914397265 0.7321338619 -1.3635085567
H 0.7858921189 -0.6373779302 -0.3204311762
H 0.0218209785 2.2505182304 0.1468292069
H -0.2266241551 0.9455046686 1.3247226359
H 2.5278796928 1.7590277621 0.4727119199
H 2.1882724928 0.3782271446 1.5058219746
H -1.3025118280 0.8032883597 -1.5259809861
H 4.4479425383 0.3027435209 0.3921583620
H 3.6346463266 0.2349775607 -1.1836334244
H -3.0211573542 2.1410074345 -0.8045922540
H -3.2218629847 0.8371482423 0.3587042034
H 4.2253281871 -1.9726822174 -0.4155872948
H 3.3489651634 -1.7581308116 0.9983748528
H 2.5570336567 -1.8144640232 -0.4679619280
H -2.3203269015 2.3008181314 0.7098687755
25
-34.15085064
O -3.5353488917 0.0322108098 0.4084906022
O -2.4254579214 -1.9084093375 0.6284254091
N -1.7744403653 1.3652775888 -0.6438663864
N 4.1843308757 -0.1227935311 -1.0149931633
C 1.2054477125 0.3338345853 -0.2573250215
C -0.0723683835 -0.0922253390 0.4701395235
C 2.4067626747 0.2476116899 0.6871054133
C -1.3095998224 -0.0201188789 -0.4106994862
C 3.6989393857 0.7735207317 0.0732627710
C -2.5481677099 -0.7347193516 0.2919952398
H 1.1039179513 1.3593179418 -0.6150772671
H 1.3305507433 -0.3269971917 -1.1156830038
H -0.2323641779 0.5179203592 1.3619095074
H 0.0148258366 -1.1272987239 0.8068454593
H 2.1939049448 0.8432977651 1.5748900270
H 2.5356727482 -0.7824695024 1.0211816116
H -1.1430786561 -0.5391980852 -1.3555500184
H 4.4681954117 0.8303339300 0.8434799998
H 3.5410323285 1.7707403176 -0.3387001734
H -2.7959421798 1.2370421526 -0.2418926774
H -1.2663143192 2.0543977636 -0.0827359543
H 4.3101055621 -1.0795234870 -0.6518667702
H 3.4997443822 -0.1538144186 -1.7841531054
H 5.0842467659 0.2120203831 -1.3873716549
H -1.8037773550 1.6476905491 -1.6249441216
25
-34.15048917
O -1.9774584219 -0.8822136242 -1.3447844876
O -0.7353222350 -1.3040874260 0.4712779300
N -1.2018711229 0.8204001109 1.6759883927
N 1.9464051556 -1.6923491374 0.3496868798
C 0.8058411616 1.0462579377 -0.9468506072
C -0.4754178942 1.7785998054 -0.5275051890
C 1.8226970853 0.7789686777 0.1752315710
C -1.5019161161 0.9155320919 0.2220611780
C 2.6576744598 -0.4659344426 -0.1177787426
C -1.4255171488 -0.5796126398 -0.2987743324
H 1.2927227489 1.6467822241 -1.7161034319
H 0.5115436171 0.1136959367 -1.4299162738
H -0.9626535770 2.1244807715 -1.4395462399
H -0.2313854006 2.6631384020 0.0636669169
H 2.4836824282 1.6401220787 0.2631946203
H 1.3379895832 0.6415415675 1.1385387246
H -2.5044279955 1.3042479639 0.0511624497
H 3.6220420553 -0.4105315490 0.3870533340
H 2.8346807725 -0.5412209492 -1.1903941633
H -0.5537202797 1.5469435382 2.0029686770
H -2.0510266978 0.8448716653 2.2497725116
H 0.8939724574 -1.6086997720 0.2334441992
H 2.2630899289 -2.5235198565 -0.1606712473
H 2.1155048181 -1.8450780227 1.3513380594
H -0.7798223195 -0.1709536166 1.7101046448
25
-34.15043307
O -3.3078583007 -0.3518099727 0.5480566241
O -1.8441149941 -2.0073969491 0.1397595703
N -1.9672328473 1.5403727742 -0.2372000800
N 3.6044925126 -0.8358440764 -0.6659647320
C 1.1684565146 1.0446332415 -0.4943940738
C 0.1082467096 0.2177955398 0.2395309885
C 2.4785715426 1.1657908669 0.2896724500
C -1.2458520269 0.2676289576 -0.4513144135
C 3.1696429470 -0.1592315605 0.5890169973
C -2.2263428078 -0.8414226742 0.1422529421
H 0.7932826990 2.0569151989 -0.6482792542
H 1.3354698371 0.6191303209 -1.4843783659
H -0.0031461344 0.5661490514 1.2686433794
H 0.3791631733 -0.8378892566 0.2841197320
H 3.1579098277 1.8188328559 -0.2597110183
H 2.2713923792 1.6478155935 1.2450536588
H -1.1432114953 0.0569418065 -1.5170262136
H 2.5004236326 -0.8240708177 1.1341470919
H 4.0493855213 0.0280822082 1.2049984895
H -1.5070190112 2.1488324010 0.4454597884
H -2.1837208522 2.0584218679 -1.0901715209
H 4.2094106392 -0.2068158824 -1.2153183037
H 4.1256865653 -1.6973771969 -0.4514692989
H 2.7834959501 -1.0854874671 -1.2358687369
H -2.8845837684 1.1083956166 0.2016053653
Calculate free energies on populated, DFT optimized conformers
Note
Using sorting parts: part0 and part1 in order to reduce the number of computational costly DFT geometry optimizations
For the cheap-prescreening in part 0, the default level of theory is B97-D/def2-SV(P). Using a higher level of theory, e.g., a hybrid DFA as B3LYP-D3 with a large basis set leads to unnecessarily high computational costs in this part.
example settings are read from .censorc file
$ censo -inp ensemble.xyz -inprc /path/to/.censorc > censo.out &
______________________________________________________________
| |
| |
| CENSO - Commandline ENSO |
| v 1.0.3 |
| energetic sorting of CREST Conformer Rotamer Ensembles |
| University of Bonn, MCTC |
| Feb 2021 |
| based on ENSO version 2.0.1 |
| F. Bohle and S. Grimme |
| |
|______________________________________________________________|
This program is distributed in the hope that it will be useful,
but WITHOUT ANY WARRANTY; without even the implied warranty of
MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE.
----------------------------------------------------------------------------------------------------
PARAMETERS
----------------------------------------------------------------------------------------------------
The configuration file .censorc is read from /home/bohle/1projects/from_tmp1/CENSO/documentation-calcs/3-part0-2/.censorc.
Reading conformer rotamer ensemble from: /home/bohle/1projects/from_tmp1/CENSO/documentation-calcs/3-part0-2/ensemble.xyz.
Reading file: censo_solvents.json
--------------------------------------------------
CRE SORTING SETTINGS
--------------------------------------------------
number of atoms in system: 25
number of considered conformers: 22
number of all conformers from input: 22
charge: 1
unpaired: 0
solvent: h2o
temperature: 298.15
evaluate at different temperatures: on
temperature range: 273.15, 278.15, 283.15, 288.15, ...
calculate mRRHO contribution: on
consider symmetry for mRRHO contribution: off
cautious checking for error and failed calculations: on
checking the DFT-ensemble using CREST: off
maxthreads: 4
omp: 2
--------------------------------------------------
CRE CHEAP-PRESCREENING - PART0
--------------------------------------------------
part0: on
starting number of considered conformers: 22
program for part0: tm
functional for fast single-point: b97-d
basis set for fast single-point: def2-SV(P)
threshold g_thr(0) for sorting in part0: 4.0
Solvent model used with xTB: alpb
short-notation:
b97-d-D3/def2-SV(P) // GFNn-xTB (Input geometry)
--------------------------------------------------
CRE PRESCREENING - PART1
--------------------------------------------------
part1: on
starting number of considered conformers: 22
program for part1: tm
functional for initial evaluation: r2scan-3c
basis set for initial evaluation: def2-mTZVPP
calculate mRRHO contribution: on
program for mRRHO contribution: xtb
GFN version for mRRHO and/or GBSA_Gsolv: gfn2
Apply constraint to input geometry during mRRHO calculation: on
solvent model applied with xTB: alpb
evaluate at different temperatures: off
threshold g_thr(1) and G_thr(1) for sorting in part1: 3.5
solvent model for Gsolv contribution of part1: cosmors
short-notation:
r2scan-3c + COSMORS[h2o] + GmRRHO(GFN2[alpb]-bhess) // GFNn-xTB (Input geometry)
--------------------------------------------------
CRE OPTIMIZATION - PART2
--------------------------------------------------
part2: on
program: tm
functional for part2: r2scan-3c
basis set for part2: def2-mTZVPP
using xTB-optimizer for optimization: on
using the new ensemble optimizer: on
optimize all conformers below this G_thr(opt,2) threshold: 2.5
spearmanthr: 0.935
optimization level in part2: lax
solvent model applied in the optimization: dcosmors
solvent model for Gsolv contribution: cosmors
evaluate at different temperatures: on
Boltzmann sum threshold G_thr(2) for sorting in part2: 99.0
calculate mRRHO contribution: on
program for mRRHO contribution: xtb
GFN version for mRRHO and/or GBSA_Gsolv: gfn2
Apply constraint to input geometry during mRRHO calculation: on
solvent model applied with xTB: alpb
short-notation:
r2scan-3c + COSMORS[h2o] + GmRRHO(GFN2[alpb]-bhess) // r2scan-3c[DCOSMORS]
END of parameters
------------------------------------------------------------
PATHS of external QM programs
------------------------------------------------------------
The following program paths are used:
xTB: /home/abt-grimme/AK-bin/xtb
TURBOMOLE: /home/abt-grimme/TURBOMOLE.7.5.1/bin/em64t-unknown-linux-gnu_smp
Setup of COSMO-RS:
ctd = BP_TZVPD_FINE_C30_1601.ctd
cdir = "/home/bohle/COSMOlogic/COSMOthermX19/COSMOtherm/CTDATA-FILES"
ldir = "/home/bohle/COSMOlogic/COSMOthermX19/COSMOtherm/CTDATA-FILES"
Using /home/bohle/COSMOlogic/COSMOthermX19/COSMOtherm/DATABASE-COSMO/BP-TZVP-COSMO
as path to the COSMO-RS NORMAL DATABASE.
Using cefine from /home/bohle/bin/cefine
PARNODES for TM or COSMO-RS calculation was set to 2
Using COSMOtherm from /home/bohle/COSMOlogic/COSMOthermX19/COSMOtherm/BIN-LINUX/cosmotherm
----------------------------------------------------------------------------------------------------
Processing data from previous run (enso.json)
----------------------------------------------------------------------------------------------------
INFORMATION: No restart information exists and is created during this run!
----------------------------------------------------------------------------------------------------
CRE CHEAP-PRESCREENING - PART0
----------------------------------------------------------------------------------------------------
program: tm
functional for part0: b97-d
basis set for part0: def2-SV(P)
threshold g_thr(0): 4.0
starting number of considered conformers: 22
Calculating efficient gas-phase single-point energies:
The efficient gas-phase single-point is calculated for:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF10, CONF11
CONF12, CONF13, CONF14, CONF15, CONF16, CONF17, CONF18, CONF19, CONF20, CONF21, CONF22
Constructed folders!
Starting 22 ALPB-Gsolv calculations
Running single-point in CONF1/part0_sp
Running single-point in CONF2/part0_sp
Running single-point in CONF3/part0_sp
Running single-point in CONF4/part0_sp
Running ALPB_GSOLV calculation in 3-part0-2/CONF1/part0_sp
Running ALPB_GSOLV calculation in 3-part0-2/CONF3/part0_sp
Running ALPB_GSOLV calculation in 3-part0-2/CONF2/part0_sp
Running ALPB_GSOLV calculation in 3-part0-2/CONF4/part0_sp
Running single-point in CONF5/part0_sp
Running single-point in CONF6/part0_sp
Running single-point in CONF7/part0_sp
Running single-point in CONF8/part0_sp
Running ALPB_GSOLV calculation in 3-part0-2/CONF5/part0_sp
Running ALPB_GSOLV calculation in 3-part0-2/CONF6/part0_sp
Running ALPB_GSOLV calculation in 3-part0-2/CONF7/part0_sp
Running ALPB_GSOLV calculation in 3-part0-2/CONF8/part0_sp
Running single-point in CONF9/part0_sp
Running single-point in CONF11/part0_sp
Running single-point in CONF10/part0_sp
Running single-point in CONF12/part0_sp
Running ALPB_GSOLV calculation in 3-part0-2/CONF9/part0_sp
Running ALPB_GSOLV calculation in 3-part0-2/CONF10/part0_sp
Running ALPB_GSOLV calculation in 3-part0-2/CONF11/part0_sp
Running ALPB_GSOLV calculation in 3-part0-2/CONF12/part0_sp
Running single-point in CONF13/part0_sp
Running single-point in CONF15/part0_sp
Running single-point in CONF14/part0_sp
Running single-point in CONF16/part0_sp
Running ALPB_GSOLV calculation in 3-part0-2/CONF13/part0_sp
Running ALPB_GSOLV calculation in 3-part0-2/CONF15/part0_sp
Running ALPB_GSOLV calculation in 3-part0-2/CONF14/part0_sp
Running single-point in CONF17/part0_sp
Running ALPB_GSOLV calculation in 3-part0-2/CONF16/part0_sp
Running single-point in CONF18/part0_sp
Running single-point in CONF19/part0_sp
Running ALPB_GSOLV calculation in 3-part0-2/CONF17/part0_sp
Running single-point in CONF20/part0_sp
Running ALPB_GSOLV calculation in 3-part0-2/CONF18/part0_sp
Running ALPB_GSOLV calculation in 3-part0-2/CONF19/part0_sp
Running single-point in CONF21/part0_sp
Running single-point in CONF22/part0_sp
Running ALPB_GSOLV calculation in 3-part0-2/CONF20/part0_sp
Running ALPB_GSOLV calculation in 3-part0-2/CONF22/part0_sp
Running ALPB_GSOLV calculation in 3-part0-2/CONF21/part0_sp
Tasks completed!
The efficient gas-phase single-point was successful for CONF1/part0_sp: E(DFT) = -496.55110270 Gsolv = -0.12279245
The efficient gas-phase single-point was successful for CONF2/part0_sp: E(DFT) = -496.55473079 Gsolv = -0.12241044
The efficient gas-phase single-point was successful for CONF3/part0_sp: E(DFT) = -496.55551554 Gsolv = -0.12099969
The efficient gas-phase single-point was successful for CONF4/part0_sp: E(DFT) = -496.55384665 Gsolv = -0.12156811
The efficient gas-phase single-point was successful for CONF5/part0_sp: E(DFT) = -496.55341065 Gsolv = -0.12251050
The efficient gas-phase single-point was successful for CONF6/part0_sp: E(DFT) = -496.55368828 Gsolv = -0.12175678
The efficient gas-phase single-point was successful for CONF7/part0_sp: E(DFT) = -496.54759208 Gsolv = -0.12365913
The efficient gas-phase single-point was successful for CONF8/part0_sp: E(DFT) = -496.55118920 Gsolv = -0.12200372
The efficient gas-phase single-point was successful for CONF9/part0_sp: E(DFT) = -496.54970422 Gsolv = -0.12167549
The efficient gas-phase single-point was successful for CONF10/part0_sp: E(DFT) = -496.55157272 Gsolv = -0.12001541
The efficient gas-phase single-point was successful for CONF11/part0_sp: E(DFT) = -496.54991969 Gsolv = -0.12296934
The efficient gas-phase single-point was successful for CONF12/part0_sp: E(DFT) = -496.55212770 Gsolv = -0.11751864
The efficient gas-phase single-point was successful for CONF13/part0_sp: E(DFT) = -496.55077718 Gsolv = -0.11854291
The efficient gas-phase single-point was successful for CONF14/part0_sp: E(DFT) = -496.55028792 Gsolv = -0.12090071
The efficient gas-phase single-point was successful for CONF15/part0_sp: E(DFT) = -496.54987374 Gsolv = -0.12311773
The efficient gas-phase single-point was successful for CONF16/part0_sp: E(DFT) = -496.55360018 Gsolv = -0.11274054
The efficient gas-phase single-point was successful for CONF17/part0_sp: E(DFT) = -496.55289531 Gsolv = -0.11378052
The efficient gas-phase single-point was successful for CONF18/part0_sp: E(DFT) = -496.50423172 Gsolv = -0.16468285
The efficient gas-phase single-point was successful for CONF19/part0_sp: E(DFT) = -496.52384365 Gsolv = -0.14390846
The efficient gas-phase single-point was successful for CONF20/part0_sp: E(DFT) = -496.50481476 Gsolv = -0.16519450
The efficient gas-phase single-point was successful for CONF21/part0_sp: E(DFT) = -496.53860751 Gsolv = -0.13212346
The efficient gas-phase single-point was successful for CONF22/part0_sp: E(DFT) = -496.50938650 Gsolv = -0.15826546
----------------------------------------------------------------------------------------------------
Removing high lying conformers by improved energy description
----------------------------------------------------------------------------------------------------
CONF# G [Eh] ΔG [kcal/mol] E [Eh] Gsolv [Eh] Gtot ΔE(DFT) ΔGsolv ΔGtot
GFN2-xTB GFN2-xTB b97-d-D3(0)/def2-SV(P) alpb [Eh] [kcal/mol] [kcal/mol] [kcal/mol]
[ALPB] [ALPB] [gfn2]
CONF1 -34.1746819 0.00 -496.5511027 -0.1227924 -496.6738951 2.28 -0.24 2.04
CONF2 -34.1743917 0.18 -496.5547308 -0.1224104 -496.6771412 0.00 0.00 0.00 <------
CONF3 -34.1742792 0.25 -496.5555155 -0.1209997 -496.6765152 -0.49 0.89 0.39
CONF4 -34.1739821 0.44 -496.5538467 -0.1215681 -496.6754148 0.55 0.53 1.08
CONF5 -34.1740224 0.41 -496.5534106 -0.1225105 -496.6759212 0.83 -0.06 0.77
CONF6 -34.1736309 0.66 -496.5536883 -0.1217568 -496.6754451 0.65 0.41 1.06
CONF7 -34.1725555 1.33 -496.5475921 -0.1236591 -496.6712512 4.48 -0.78 3.70
CONF8 -34.1721876 1.57 -496.5511892 -0.1220037 -496.6731929 2.22 0.26 2.48
CONF9 -34.1719439 1.72 -496.5497042 -0.1216755 -496.6713797 3.15 0.46 3.62
CONF10 -34.1714765 2.01 -496.5515727 -0.1200154 -496.6715881 1.98 1.50 3.48
CONF11 -34.1712638 2.14 -496.5499197 -0.1229693 -496.6728890 3.02 -0.35 2.67
CONF12 -34.1704841 2.63 -496.5521277 -0.1175186 -496.6696463 1.63 3.07 4.70
CONF13 -34.1703687 2.71 -496.5507772 -0.1185429 -496.6693201 2.48 2.43 4.91
CONF14 -34.1694191 3.30 -496.5502879 -0.1209007 -496.6711886 2.79 0.95 3.74
CONF15 -34.1691431 3.48 -496.5498737 -0.1231177 -496.6729915 3.05 -0.44 2.60
CONF16 -34.1664418 5.17 -496.5536002 -0.1127405 -496.6663407 0.71 6.07 6.78
CONF17 -34.1661652 5.34 -496.5528953 -0.1137805 -496.6666758 1.15 5.42 6.57
CONF18 -34.1660003 5.45 -496.5042317 -0.1646828 -496.6689146 31.69 -26.53 5.16
CONF19 -34.1664801 5.15 -496.5238437 -0.1439085 -496.6677521 19.38 -13.49 5.89
CONF20 -34.1658387 5.55 -496.5048148 -0.1651945 -496.6700093 31.32 -26.85 4.48
CONF21 -34.1670164 4.81 -496.5386075 -0.1321235 -496.6707310 10.12 -6.10 4.02
CONF22 -34.1656180 5.69 -496.5093865 -0.1582655 -496.6676520 28.45 -22.50 5.95
----------------------------------------------------------------------------------------------------
--------------------------------------------------
Conformers considered further
--------------------------------------------------
These conformers are below the 4.000 kcal/mol g_thr(0) threshold.
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF10, CONF11
CONF14, CONF15
Calculating Boltzmann averaged (free) energy of ensemble on input geometries (not DFT optimized)!
temperature /K: avE(T) /a.u. avG(T) /a.u.
----------------------------------------------------------------------------------------------------
298.15 -496.5544580 -496.6764445 <<==part0==
----------------------------------------------------------------------------------------------------
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>END of Part0<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
Ran part0 in 25.3156 seconds
----------------------------------------------------------------------------------------------------
CRE PRESCREENING - PART1
----------------------------------------------------------------------------------------------------
program: tm
functional for part1 and 2: r2scan-3c
basis set for part1 and 2: def2-mTZVPP
Solvent: h2o
solvent model for Gsolv contribution: cosmors
threshold g_thr(1) and G_thr(1): 3.5
starting number of considered conformers: 13
calculate mRRHO contribution: on
program for mRRHO contribution: xtb
GFN version for mRRHO and/or GBSA_Gsolv: gfn2
Apply constraint to input geometry during mRRHO calculation: on
temperature: 298.15
Calculating single-point energies and solvation contribution (G_solv):
The prescreening COSMO-RS is calculated for:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF10, CONF11
CONF14, CONF15
Constructed folders!
Constructed folders!
Starting 13 COSMO-RS-Gsolv calculations.
Running COSMO-RS calculation in CONF1/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF2/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF3/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF4/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF5/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF6/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF7/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF8/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF9/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF10/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF11/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF14/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF15/r2scan-3c/COSMO
Tasks completed!
prescreening COSMO-RS calculation was successful for CONF1/r2scan-3c/COSMO: -0.10999518
prescreening COSMO-RS calculation was successful for CONF2/r2scan-3c/COSMO: -0.10920229
prescreening COSMO-RS calculation was successful for CONF3/r2scan-3c/COSMO: -0.10753681
prescreening COSMO-RS calculation was successful for CONF4/r2scan-3c/COSMO: -0.10829026
prescreening COSMO-RS calculation was successful for CONF5/r2scan-3c/COSMO: -0.10936797
prescreening COSMO-RS calculation was successful for CONF6/r2scan-3c/COSMO: -0.10838241
prescreening COSMO-RS calculation was successful for CONF7/r2scan-3c/COSMO: -0.11058677
prescreening COSMO-RS calculation was successful for CONF8/r2scan-3c/COSMO: -0.10882956
prescreening COSMO-RS calculation was successful for CONF9/r2scan-3c/COSMO: -0.10889910
prescreening COSMO-RS calculation was successful for CONF10/r2scan-3c/COSMO: -0.10665800
prescreening COSMO-RS calculation was successful for CONF11/r2scan-3c/COSMO: -0.10960927
prescreening COSMO-RS calculation was successful for CONF14/r2scan-3c/COSMO: -0.10711489
prescreening COSMO-RS calculation was successful for CONF15/r2scan-3c/COSMO: -0.10994652
--------------------------------------------------
Removing high lying conformers
--------------------------------------------------
CONF# E(GFNn-xTB) ΔE(GFNn-xTB) E [Eh] Gsolv [Eh] Gtot ΔGtot
[a.u.] [kcal/mol] r2scan-3c COSMO-RS-normal [Eh] [kcal/mol]
[r2scan-3c]
CONF1 -34.1599548 0.00 -497.3021467 -0.1099952 -497.4121419 1.82
CONF2 -34.1594484 0.32 -497.3049956 -0.1092023 -497.4141979 0.53
CONF3 -34.1592530 0.44 -497.3064930 -0.1075368 -497.4140298 0.64
CONF4 -34.1590954 0.54 -497.3049058 -0.1082903 -497.4131961 1.16
CONF5 -34.1589250 0.65 -497.3056747 -0.1093680 -497.4150427 0.00 <------
CONF6 -34.1587465 0.76 -497.3049670 -0.1083824 -497.4133494 1.06
CONF7 -34.1578065 1.35 -497.2983644 -0.1105868 -497.4089512 3.82
CONF8 -34.1571241 1.78 -497.3029570 -0.1088296 -497.4117866 2.04
CONF9 -34.1570372 1.83 -497.3008248 -0.1088991 -497.4097239 3.34
CONF10 -34.1563217 2.28 -497.3038986 -0.1066580 -497.4105566 2.82
CONF11 -34.1559251 2.53 -497.3008250 -0.1096093 -497.4104343 2.89
CONF14 -34.1537285 3.91 -497.3022176 -0.1071149 -497.4093325 3.58
CONF15 -34.1529637 4.39 -497.3009381 -0.1099465 -497.4108846 2.61
--------------------------------------------------
Conformers considered further
--------------------------------------------------
Below the g_thr(1) threshold of 3.5 kcal/mol.
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF8, CONF9, CONF10, CONF11, CONF15
--------------------------------------------------
Calculating prescreening G_mRRHO with implicit solvation!
The prescreening G_mRRHO calculation is now performed for:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF8, CONF9, CONF10, CONF11, CONF15
Constructed folders!
Starting 11 G_RRHO calculations.
Running GFN2-xTB mRRHO in CONF1/rrho_part1
Running GFN2-xTB mRRHO in CONF2/rrho_part1
Running GFN2-xTB mRRHO in CONF3/rrho_part1
Running GFN2-xTB mRRHO in CONF4/rrho_part1
Running GFN2-xTB mRRHO in CONF5/rrho_part1
Running GFN2-xTB mRRHO in CONF6/rrho_part1
Running GFN2-xTB mRRHO in CONF8/rrho_part1
Running GFN2-xTB mRRHO in CONF9/rrho_part1
Running GFN2-xTB mRRHO in CONF10/rrho_part1
Running GFN2-xTB mRRHO in CONF11/rrho_part1
Running GFN2-xTB mRRHO in CONF15/rrho_part1
Tasks completed!
The prescreening G_mRRHO calculation @ c1 was successful for CONF1/rrho_part1: 0.18287159
The prescreening G_mRRHO calculation @ c1 was successful for CONF2/rrho_part1: 0.18311981
The prescreening G_mRRHO calculation @ c1 was successful for CONF3/rrho_part1: 0.18287068
The prescreening G_mRRHO calculation @ c1 was successful for CONF4/rrho_part1: 0.18270975
The prescreening G_mRRHO calculation @ c1 was successful for CONF5/rrho_part1: 0.18272101
The prescreening G_mRRHO calculation @ c1 was successful for CONF6/rrho_part1: 0.18277331
The prescreening G_mRRHO calculation @ c1 was successful for CONF8/rrho_part1: 0.18239977
The prescreening G_mRRHO calculation @ c1 was successful for CONF9/rrho_part1: 0.18270475
The prescreening G_mRRHO calculation @ c1 was successful for CONF10/rrho_part1: 0.18264350
The prescreening G_mRRHO calculation @ c1 was successful for CONF11/rrho_part1: 0.18284194
The prescreening G_mRRHO calculation @ c1 was successful for CONF15/rrho_part1: 0.18323466
--------------------------------------------------
* Gibbs free energies of part1 *
--------------------------------------------------
CONF# G(GFNn-xTB) ΔG(GFNn-xTB) E [Eh] Gsolv [Eh] GmRRHO [Eh] Gtot ΔGtot
[a.u.] [kcal/mol] r2scan-3c COSMO-RS-normal GFN2 [Eh] [kcal/mol]
[r2scan-3c] [alpb]-bhess
CONF1 -33.9770832 0.00 -497.3021467 -0.1099952 0.1828716 -497.2292703 1.91
CONF2 -33.9763286 0.47 -497.3049956 -0.1092023 0.1831198 -497.2310781 0.78
CONF3 -33.9763824 0.44 -497.3064930 -0.1075368 0.1828707 -497.2311591 0.73
CONF4 -33.9763856 0.44 -497.3049058 -0.1082903 0.1827098 -497.2304863 1.15
CONF5 -33.9762040 0.55 -497.3056747 -0.1093680 0.1827210 -497.2323217 0.00 <------
CONF6 -33.9759732 0.70 -497.3049670 -0.1083824 0.1827733 -497.2305761 1.10
CONF8 -33.9747243 1.48 -497.3029570 -0.1088296 0.1823998 -497.2293868 1.84
CONF9 -33.9743324 1.73 -497.3008248 -0.1088991 0.1827047 -497.2270192 3.33
CONF10 -33.9736782 2.14 -497.3038986 -0.1066580 0.1826435 -497.2279131 2.77
CONF11 -33.9730831 2.51 -497.3008250 -0.1096093 0.1828419 -497.2275923 2.97
CONF15 -33.9697290 4.61 -497.3009381 -0.1099465 0.1832347 -497.2276500 2.93
Additional global 'fuzzy-threshold' based on the standard deviation of (G_mRRHO):
Std_dev(G_mRRHO) = 0.142 kcal/mol
Fuzzythreshold = 0.096 kcal/mol
Final sorting threshold G_thr(1) = 3.500 + 0.096 = 3.596 kcal/mol
Spearman correlation coefficient between (E + Solv) and (E + Solv + mRRHO) = 0.973
All relative (free) energies are below the initial G_thr(1) threshold of 3.5 kcal/mol.
All conformers are considered further.
Calculating Boltzmann averaged free energy of ensemble on input geometries (not DFT optimized)!
temperature /K: avE(T) /a.u. avGmRRHO(T) /a.u. avGsolv(T) /a.u. avG(T) /a.u.
----------------------------------------------------------------------------------------------------
298.15 -497.3054081 0.1827983 -0.1089068 -497.2315166 <<==part1==
----------------------------------------------------------------------------------------------------
Calculating unbiased GFNn-xTB energy
Constructed folders!
Starting 11 xTB - single-point calculations.
gfn2-xTB energy for CONF1/GFN_unbiased = -34.1764957
gfn2-xTB energy for CONF2/GFN_unbiased = -34.1762055
gfn2-xTB energy for CONF3/GFN_unbiased = -34.1760929
gfn2-xTB energy for CONF4/GFN_unbiased = -34.1757959
gfn2-xTB energy for CONF5/GFN_unbiased = -34.1758362
gfn2-xTB energy for CONF6/GFN_unbiased = -34.1754446
gfn2-xTB energy for CONF8/GFN_unbiased = -34.1740014
gfn2-xTB energy for CONF9/GFN_unbiased = -34.1737577
gfn2-xTB energy for CONF10/GFN_unbiased = -34.1732903
gfn2-xTB energy for CONF11/GFN_unbiased = -34.1730775
gfn2-xTB energy for CONF15/GFN_unbiased = -34.1709569
Tasks completed!
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>END of Part1<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
Ran part1 in 166.4199 seconds
----------------------------------------------------------------------------------------------------
CRE OPTIMIZATION - PART2
----------------------------------------------------------------------------------------------------
program: tm
functional for part2: r2scan-3c
basis set for part2: def2-mTZVPP
using the xTB-optimizer for optimization: on
using the new ensemble optimizer: on
optimize all conformers below this G_thr(opt,2) threshold: 2.5
Spearman threshold: 0.935
number of optimization iterations: 8
radsize: 10
optimization level in part2: lax
solvent: h2o
solvent model applied in the optimization: dcosmors
solvent model for Gsolv contribution: cosmors
temperature: 298.15
evalulate at different temperatures: on
temperature range: 273.15, 278.15, 283.15, 288.15, ...
Boltzmann sum threshold G_thr(2) for sorting in part2: 99.0
calculate mRRHO contribution: on
program for mRRHO contribution: xtb
GFN version for mRRHO and/or GBSA_Gsolv: gfn2
Apply constraint to input geometry during mRRHO calculation: on
Optimizing geometries at DFT level with implicit solvation!
The optimization is calculated for:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF8, CONF9, CONF10, CONF11, CONF15
Constructed folders!
Preparing 11 calculations.
Tasks completed!
************************Starting optimizations************************
Starting threshold is set to 2.5 + 60.0 % = 4.0 kcal/mol
Lower limit is set to G_thr(opt,2) = 2.5 kcal/mol
*******************************CYCLE 1********************************
Starting 11 optimizations.
Running optimization in CONF1/r2scan-3c
Running optimization in CONF2/r2scan-3c
Running optimization in CONF3/r2scan-3c
Running optimization in CONF4/r2scan-3c
Running optimization in CONF5/r2scan-3c
Running optimization in CONF6/r2scan-3c
Running optimization in CONF8/r2scan-3c
Running optimization in CONF9/r2scan-3c
Running optimization in CONF10/r2scan-3c
Running optimization in CONF11/r2scan-3c
Running optimization in CONF15/r2scan-3c
Tasks completed!
Constructed folders!
Starting 11 G_RRHO calculations.
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Tasks completed!
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF1/r2scan-3c/rrho_crude: 0.18253283
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF2/r2scan-3c/rrho_crude: 0.18217211
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF3/r2scan-3c/rrho_crude: 0.18223212
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF4/r2scan-3c/rrho_crude: 0.18258415
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF5/r2scan-3c/rrho_crude: 0.18245612
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF6/r2scan-3c/rrho_crude: 0.18170042
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF8/r2scan-3c/rrho_crude: 0.18260313
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF9/r2scan-3c/rrho_crude: 0.18272834
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF10/r2scan-3c/rrho_crude: 0.18229001
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF11/r2scan-3c/rrho_crude: 0.18150537
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF15/r2scan-3c/rrho_crude: 0.18100199
Max number of performed iterations: 8
Spearman rank evaluation is performed in the next cycle.
CONF1 initial ΔG = 2.13 kcal/mol and current ΔG = 2.59 kcal/mol. (not_converged)
CONF2 initial ΔG = 0.86 kcal/mol and current ΔG = 0.46 kcal/mol. (not_converged)
CONF3 initial ΔG = 0.97 kcal/mol and current ΔG = 1.47 kcal/mol. (not_converged)
CONF4 initial ΔG = 1.62 kcal/mol and current ΔG = 1.40 kcal/mol. (not_converged)
CONF5 initial ΔG = 0.00 kcal/mol and current ΔG = 0.42 kcal/mol. (not_converged)
CONF6 initial ΔG = 0.60 kcal/mol and current ΔG = 1.15 kcal/mol. (not_converged)
CONF8 initial ΔG = 2.26 kcal/mol and current ΔG = 2.31 kcal/mol. (not_converged)
CONF9 initial ΔG = 3.90 kcal/mol and current ΔG = 4.42 kcal/mol. (not_converged)
CONF10 initial ΔG = 2.86 kcal/mol and current ΔG = 2.92 kcal/mol. (not_converged)
CONF11 initial ΔG = 2.26 kcal/mol and current ΔG = 3.43 kcal/mol. (not_converged)
CONF15 initial ΔG = 0.23 kcal/mol and current ΔG = 0.00 kcal/mol. (not_converged)
CONF9 is above 4.0 kcal/mol and gradient norm (0.004056375026) is below 0.01.
CONF9 is above threshold, dont optimize further and remove conformer.
CYCLE 1 performed in 1310.1062 seconds
*******************************CYCLE 2********************************
Starting 10 optimizations.
Running optimization in CONF1/r2scan-3c
Running optimization in CONF2/r2scan-3c
Running optimization in CONF3/r2scan-3c
Running optimization in CONF4/r2scan-3c
Running optimization in CONF5/r2scan-3c
Running optimization in CONF6/r2scan-3c
Running optimization in CONF8/r2scan-3c
Running optimization in CONF10/r2scan-3c
Running optimization in CONF11/r2scan-3c
Running optimization in CONF15/r2scan-3c
Tasks completed!
Geometry optimization converged for: CONF3 within 12 cycles
Max number of performed iterations: 16
CONF1 previous ΔG = 2.59 kcal/mol and current ΔG = 2.54 kcal/mol. (not_converged)
CONF2 previous ΔG = 0.46 kcal/mol and current ΔG = 0.00 kcal/mol. (not_converged)
CONF3 previous ΔG = 1.47 kcal/mol and current ΔG = 1.92 kcal/mol. (converged)
CONF4 previous ΔG = 1.40 kcal/mol and current ΔG = 10.87 kcal/mol. (not_converged)
CONF5 previous ΔG = 0.42 kcal/mol and current ΔG = 0.99 kcal/mol. (not_converged)
CONF6 previous ΔG = 1.15 kcal/mol and current ΔG = 1.57 kcal/mol. (not_converged)
CONF8 previous ΔG = 2.31 kcal/mol and current ΔG = 1.85 kcal/mol. (not_converged)
CONF10 previous ΔG = 2.92 kcal/mol and current ΔG = 3.68 kcal/mol. (not_converged)
CONF11 previous ΔG = 3.43 kcal/mol and current ΔG = 3.29 kcal/mol. (not_converged)
CONF15 previous ΔG = 0.00 kcal/mol and current ΔG = 0.03 kcal/mol. (not_converged)
Spearman coeff. from 8 --> 11 = 0.9758
Spearman coeff. from 9 --> 12 = 0.9879
Spearman coeff. from 10 --> 13 = 0.7455
Spearman coeff. from 11 --> 14 = 1.0000
Evaluating Spearman coeff. from 12 --> 15 = 0.9515
Evaluating Spearman coeff. from 13 --> 16 = 0.8667
Final averaged Spearman correlation coefficient: 0.9091
CONF4 is above 4.0 kcal/mol but gradient norm (0.108293192291) is above 0.01 --> not sorted out!
CYCLE 2 performed in 980.4814 seconds
*******************************CYCLE 3********************************
Starting 9 optimizations.
Running optimization in CONF1/r2scan-3c
Running optimization in CONF2/r2scan-3c
Running optimization in CONF4/r2scan-3c
Running optimization in CONF5/r2scan-3c
Running optimization in CONF6/r2scan-3c
Running optimization in CONF8/r2scan-3c
Running optimization in CONF10/r2scan-3c
Running optimization in CONF11/r2scan-3c
Running optimization in CONF15/r2scan-3c
Tasks completed!
Geometry optimization converged for: CONF5 within 23 cycles
Geometry optimization converged for: CONF11 within 21 cycles
Max number of performed iterations: 24
CONF1 previous ΔG = 2.54 kcal/mol and current ΔG = 2.38 kcal/mol. (not_converged)
CONF2 previous ΔG = 0.00 kcal/mol and current ΔG = 0.00 kcal/mol. (not_converged)
CONF3 previous ΔG = 1.92 kcal/mol and current ΔG = 2.09 kcal/mol. (converged)
CONF4 previous ΔG = 10.87 kcal/mol and current ΔG = 0.07 kcal/mol. (not_converged)
CONF5 previous ΔG = 0.99 kcal/mol and current ΔG = 1.25 kcal/mol. (converged)
CONF6 previous ΔG = 1.57 kcal/mol and current ΔG = 1.51 kcal/mol. (not_converged)
CONF8 previous ΔG = 1.85 kcal/mol and current ΔG = 1.89 kcal/mol. (not_converged)
CONF10 previous ΔG = 3.68 kcal/mol and current ΔG = 3.36 kcal/mol. (not_converged)
CONF11 previous ΔG = 3.29 kcal/mol and current ΔG = 3.30 kcal/mol. (converged)
CONF15 previous ΔG = 0.03 kcal/mol and current ΔG = 0.22 kcal/mol. (not_converged)
Spearman coeff. from 16 --> 19 = 0.6606
Spearman coeff. from 17 --> 20 = 0.6000
Spearman coeff. from 18 --> 21 = 0.7455
Spearman coeff. from 19 --> 22 = 0.8182
Evaluating Spearman coeff. from 20 --> 23 = 0.7091
Evaluating Spearman coeff. from 21 --> 24 = 1.0000
Final averaged Spearman correlation coefficient: 0.8545
CYCLE 3 performed in 830.9565 seconds
*******************************CYCLE 4********************************
Starting 7 optimizations.
Running optimization in CONF1/r2scan-3c
Running optimization in CONF2/r2scan-3c
Running optimization in CONF4/r2scan-3c
Running optimization in CONF6/r2scan-3c
Running optimization in CONF8/r2scan-3c
Running optimization in CONF10/r2scan-3c
Running optimization in CONF15/r2scan-3c
Tasks completed!
Geometry optimization converged for: CONF8 within 26 cycles
Geometry optimization converged for: CONF15 within 25 cycles
Max number of performed iterations: 32
CONF1 previous ΔG = 2.38 kcal/mol and current ΔG = 1.84 kcal/mol. (not_converged)
CONF2 previous ΔG = 0.00 kcal/mol and current ΔG = 0.00 kcal/mol. (not_converged)
CONF3 previous ΔG = 2.09 kcal/mol and current ΔG = 2.11 kcal/mol. (converged)
CONF4 previous ΔG = 0.07 kcal/mol and current ΔG = 0.01 kcal/mol. (not_converged)
CONF5 previous ΔG = 1.25 kcal/mol and current ΔG = 1.26 kcal/mol. (converged)
CONF6 previous ΔG = 1.51 kcal/mol and current ΔG = 1.57 kcal/mol. (not_converged)
CONF8 previous ΔG = 1.89 kcal/mol and current ΔG = 1.91 kcal/mol. (converged)
CONF10 previous ΔG = 3.36 kcal/mol and current ΔG = 6.58 kcal/mol. (not_converged)
CONF11 previous ΔG = 3.30 kcal/mol and current ΔG = 3.32 kcal/mol. (converged)
CONF15 previous ΔG = 0.22 kcal/mol and current ΔG = 0.23 kcal/mol. (converged)
Spearman coeff. from 24 --> 27 = 0.9636
Spearman coeff. from 25 --> 28 = 0.9879
Spearman coeff. from 26 --> 29 = 0.9758
Spearman coeff. from 27 --> 30 = 0.8424
Evaluating Spearman coeff. from 28 --> 31 = 0.9515
Evaluating Spearman coeff. from 29 --> 32 = 0.9636
Final averaged Spearman correlation coefficient: 0.9576
PES is assumed to be parallel
Updated optimization threshold to: 3.50 kcal/mol
CONF10 is above 3.5 kcal/mol but gradient norm (0.09473608827) is above 0.01 --> not sorted out!
CYCLE 4 performed in 613.0740 seconds
*******************************CYCLE 5********************************
Starting 5 optimizations.
Running optimization in CONF1/r2scan-3c
Running optimization in CONF2/r2scan-3c
Running optimization in CONF4/r2scan-3c
Running optimization in CONF6/r2scan-3c
Running optimization in CONF10/r2scan-3c
Tasks completed!
Geometry optimization converged for: CONF2 within 34 cycles
Geometry optimization converged for: CONF6 within 39 cycles
Max number of performed iterations: 40
CONF1 previous ΔG = 1.84 kcal/mol and current ΔG = 7.79 kcal/mol. (not_converged)
CONF2 previous ΔG = 0.00 kcal/mol and current ΔG = 0.38 kcal/mol. (converged)
CONF3 previous ΔG = 2.11 kcal/mol and current ΔG = 2.53 kcal/mol. (converged)
CONF4 previous ΔG = 0.01 kcal/mol and current ΔG = 0.35 kcal/mol. (not_converged)
CONF5 previous ΔG = 1.26 kcal/mol and current ΔG = 1.68 kcal/mol. (converged)
CONF6 previous ΔG = 1.57 kcal/mol and current ΔG = 1.94 kcal/mol. (converged)
CONF8 previous ΔG = 1.91 kcal/mol and current ΔG = 2.33 kcal/mol. (converged)
CONF10 previous ΔG = 6.58 kcal/mol and current ΔG = 0.00 kcal/mol. (not_converged)
CONF11 previous ΔG = 3.32 kcal/mol and current ΔG = 3.74 kcal/mol. (converged)
CONF15 previous ΔG = 0.23 kcal/mol and current ΔG = 0.65 kcal/mol. (converged)
Spearman coeff. from 32 --> 35 = 0.7333
Spearman coeff. from 33 --> 36 = 0.9273
Spearman coeff. from 34 --> 37 = 0.8182
Spearman coeff. from 35 --> 38 = 1.0000
Evaluating Spearman coeff. from 36 --> 39 = 0.9879
Evaluating Spearman coeff. from 37 --> 40 = 0.8545
Final averaged Spearman correlation coefficient: 0.9212
CONF1 is above 3.5 kcal/mol but gradient norm (0.083474094636) is above 0.01 --> not sorted out!
CYCLE 5 performed in 403.3400 seconds
*******************************CYCLE 6********************************
Starting 3 optimizations.
Running optimization in CONF1/r2scan-3c
Running optimization in CONF4/r2scan-3c
Running optimization in CONF10/r2scan-3c
Tasks completed!
Geometry optimization converged for: CONF4 within 44 cycles
Max number of performed iterations: 48
CONF1 previous ΔG = 7.79 kcal/mol and current ΔG = 2.37 kcal/mol. (not_converged)
CONF2 previous ΔG = 0.38 kcal/mol and current ΔG = 1.25 kcal/mol. (converged)
CONF3 previous ΔG = 2.53 kcal/mol and current ΔG = 3.40 kcal/mol. (converged)
CONF4 previous ΔG = 0.35 kcal/mol and current ΔG = 1.19 kcal/mol. (converged)
CONF5 previous ΔG = 1.68 kcal/mol and current ΔG = 2.55 kcal/mol. (converged)
CONF6 previous ΔG = 1.94 kcal/mol and current ΔG = 2.80 kcal/mol. (converged)
CONF8 previous ΔG = 2.33 kcal/mol and current ΔG = 3.20 kcal/mol. (converged)
CONF10 previous ΔG = 0.00 kcal/mol and current ΔG = 0.00 kcal/mol. (not_converged)
CONF11 previous ΔG = 3.74 kcal/mol and current ΔG = 4.61 kcal/mol. (converged)
CONF15 previous ΔG = 0.65 kcal/mol and current ΔG = 1.52 kcal/mol. (converged)
Spearman coeff. from 40 --> 43 = 0.8788
Spearman coeff. from 41 --> 44 = 0.6242
Spearman coeff. from 42 --> 45 = 0.9273
Spearman coeff. from 43 --> 46 = 1.0000
Evaluating Spearman coeff. from 44 --> 47 = 0.6485
Evaluating Spearman coeff. from 45 --> 48 = 0.9879
Final averaged Spearman correlation coefficient: 0.8182
CYCLE 6 performed in 296.2324 seconds
*******************************CYCLE 7********************************
Starting 2 optimizations.
Running optimization in CONF1/r2scan-3c
Running optimization in CONF10/r2scan-3c
Tasks completed!
Max number of performed iterations: 56
CONF1 previous ΔG = 2.37 kcal/mol and current ΔG = 2.60 kcal/mol. (not_converged)
CONF2 previous ΔG = 1.25 kcal/mol and current ΔG = 1.66 kcal/mol. (converged)
CONF3 previous ΔG = 3.40 kcal/mol and current ΔG = 3.81 kcal/mol. (converged)
CONF4 previous ΔG = 1.19 kcal/mol and current ΔG = 1.61 kcal/mol. (converged)
CONF5 previous ΔG = 2.55 kcal/mol and current ΔG = 2.97 kcal/mol. (converged)
CONF6 previous ΔG = 2.80 kcal/mol and current ΔG = 3.22 kcal/mol. (converged)
CONF8 previous ΔG = 3.20 kcal/mol and current ΔG = 3.61 kcal/mol. (converged)
CONF10 previous ΔG = 0.00 kcal/mol and current ΔG = 0.00 kcal/mol. (not_converged)
CONF11 previous ΔG = 4.61 kcal/mol and current ΔG = 5.02 kcal/mol. (converged)
CONF15 previous ΔG = 1.52 kcal/mol and current ΔG = 1.94 kcal/mol. (converged)
Spearman coeff. from 48 --> 51 = 1.0000
Spearman coeff. from 49 --> 52 = 1.0000
Spearman coeff. from 50 --> 53 = 1.0000
Spearman coeff. from 51 --> 54 = 1.0000
Evaluating Spearman coeff. from 52 --> 55 = 1.0000
Evaluating Spearman coeff. from 53 --> 56 = 1.0000
Final averaged Spearman correlation coefficient: 1.0000
PES is assumed to be parallel
Updated optimization threshold to: 3.00 kcal/mol
CYCLE 7 performed in 252.6446 seconds
*******************************CYCLE 8********************************
Starting 2 optimizations.
Running optimization in CONF1/r2scan-3c
Running optimization in CONF10/r2scan-3c
Tasks completed!
Max number of performed iterations: 64
CONF1 previous ΔG = 2.60 kcal/mol and current ΔG = 2.71 kcal/mol. (not_converged)
CONF2 previous ΔG = 1.66 kcal/mol and current ΔG = 1.87 kcal/mol. (converged)
CONF3 previous ΔG = 3.81 kcal/mol and current ΔG = 4.02 kcal/mol. (converged)
CONF4 previous ΔG = 1.61 kcal/mol and current ΔG = 1.81 kcal/mol. (converged)
CONF5 previous ΔG = 2.97 kcal/mol and current ΔG = 3.17 kcal/mol. (converged)
CONF6 previous ΔG = 3.22 kcal/mol and current ΔG = 3.42 kcal/mol. (converged)
CONF8 previous ΔG = 3.61 kcal/mol and current ΔG = 3.82 kcal/mol. (converged)
CONF10 previous ΔG = 0.00 kcal/mol and current ΔG = 0.00 kcal/mol. (not_converged)
CONF11 previous ΔG = 5.02 kcal/mol and current ΔG = 5.23 kcal/mol. (converged)
CONF15 previous ΔG = 1.94 kcal/mol and current ΔG = 2.14 kcal/mol. (converged)
Spearman coeff. from 56 --> 59 = 1.0000
Spearman coeff. from 57 --> 60 = 1.0000
Spearman coeff. from 58 --> 61 = 1.0000
Spearman coeff. from 59 --> 62 = 1.0000
Evaluating Spearman coeff. from 60 --> 63 = 1.0000
Evaluating Spearman coeff. from 61 --> 64 = 1.0000
Final averaged Spearman correlation coefficient: 1.0000
PES is assumed to be parallel
Updated optimization threshold to: 2.50 kcal/mol
CONF1 is above 2.5 kcal/mol and gradient norm (0.001262363132) is below 0.01.
CONF1 is removed because of the lowered threshold!
CONF1 is above threshold, dont optimize further and remove conformer.
CYCLE 8 performed in 251.6678 seconds
*******************************CYCLE 9********************************
Starting 1 optimizations.
Running optimization in CONF10/r2scan-3c
Tasks completed!
Max number of performed iterations: 72
CONF2 previous ΔG = 1.87 kcal/mol and current ΔG = 1.84 kcal/mol. (converged)
CONF3 previous ΔG = 4.02 kcal/mol and current ΔG = 3.99 kcal/mol. (converged)
CONF4 previous ΔG = 1.81 kcal/mol and current ΔG = 1.78 kcal/mol. (converged)
CONF5 previous ΔG = 3.17 kcal/mol and current ΔG = 3.14 kcal/mol. (converged)
CONF6 previous ΔG = 3.42 kcal/mol and current ΔG = 3.39 kcal/mol. (converged)
CONF8 previous ΔG = 3.82 kcal/mol and current ΔG = 3.79 kcal/mol. (converged)
CONF10 previous ΔG = 0.00 kcal/mol and current ΔG = 0.00 kcal/mol. (not_converged)
CONF11 previous ΔG = 5.23 kcal/mol and current ΔG = 5.20 kcal/mol. (converged)
CONF15 previous ΔG = 2.14 kcal/mol and current ΔG = 2.11 kcal/mol. (converged)
Spearman coeff. from 64 --> 67 = 1.0000
Spearman coeff. from 65 --> 68 = 1.0000
Spearman coeff. from 66 --> 69 = 1.0000
Spearman coeff. from 67 --> 70 = 1.0000
Evaluating Spearman coeff. from 68 --> 71 = 1.0000
Evaluating Spearman coeff. from 69 --> 72 = 1.0000
Final averaged Spearman correlation coefficient: 1.0000
PES is assumed to be parallel
Current optimization threshold: 2.50 kcal/mol
CYCLE 9 performed in 202.2149 seconds
*******************************CYCLE 10*******************************
Starting 1 optimizations.
Running optimization in CONF10/r2scan-3c
Tasks completed!
Geometry optimization converged for: CONF10 within 80 cycles
***********************Finished optimizations!************************
Timings:
Cycle: [s]
1 1310.11
2 980.48
3 830.96
4 613.07
5 403.34
6 296.23
7 252.64
8 251.67
9 202.21
10 200.34
sum: 5341.06
CONVERGED optimizations for the following remaining conformers:
Converged optimization for CONF2 after 34 cycles: -497.4508410
Converged optimization for CONF3 after 12 cycles: -497.4474711
Converged optimization for CONF4 after 44 cycles: -497.4513419
Converged optimization for CONF5 after 23 cycles: -497.4490457
Converged optimization for CONF6 after 39 cycles: -497.4478893
Converged optimization for CONF8 after 26 cycles: -497.4481610
Converged optimization for CONF10 after 80 cycles: -497.4538409
Converged optimization for CONF11 after 21 cycles: -497.4448166
Converged optimization for CONF15 after 25 cycles: -497.4492319
Calculating single-point energies and solvation contribution (G_solv)!
The low level gsolv calculation is now calculated for:
Constructed folders!
CONF2, CONF3, CONF4, CONF5, CONF6, CONF8, CONF10, CONF11, CONF15
Starting 9 COSMO-RS-Gsolv calculations.
Running COSMO-RS calculation in CONF2/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF3/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF4/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF5/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF6/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF8/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF10/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF11/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF15/r2scan-3c/COSMO
Tasks completed!
lowlevel COSMO-RS calculation was successful for CONF2/r2scan-3c/COSMO: -0.13589836
lowlevel COSMO-RS calculation was successful for CONF3/r2scan-3c/COSMO: -0.12865185
lowlevel COSMO-RS calculation was successful for CONF4/r2scan-3c/COSMO: -0.13533997
lowlevel COSMO-RS calculation was successful for CONF5/r2scan-3c/COSMO: -0.12940253
lowlevel COSMO-RS calculation was successful for CONF6/r2scan-3c/COSMO: -0.12980575
lowlevel COSMO-RS calculation was successful for CONF8/r2scan-3c/COSMO: -0.12959608
lowlevel COSMO-RS calculation was successful for CONF10/r2scan-3c/COSMO: -0.15938290
lowlevel COSMO-RS calculation was successful for CONF11/r2scan-3c/COSMO: -0.13131124
lowlevel COSMO-RS calculation was successful for CONF15/r2scan-3c/COSMO: -0.13806979
Calculating lowlevel G_mRRHO with implicit solvation on DFT geometry!
The lowlevel G_mRRHO calculation is now performed for:
CONF2, CONF3, CONF4, CONF5, CONF6, CONF8, CONF10, CONF11, CONF15
Constructed folders!
Starting 9 G_RRHO calculations.
Running GFN2-xTB mRRHO in CONF2/rrho_part2
Running GFN2-xTB mRRHO in CONF3/rrho_part2
Running GFN2-xTB mRRHO in CONF4/rrho_part2
Running GFN2-xTB mRRHO in CONF5/rrho_part2
Running GFN2-xTB mRRHO in CONF6/rrho_part2
Running GFN2-xTB mRRHO in CONF8/rrho_part2
Running GFN2-xTB mRRHO in CONF10/rrho_part2
Running GFN2-xTB mRRHO in CONF11/rrho_part2
Running GFN2-xTB mRRHO in CONF15/rrho_part2
Tasks completed!
The lowlevel G_mRRHO calculation @ c1 was successful for CONF2/rrho_part2: 0.18028469
The lowlevel G_mRRHO calculation @ c1 was successful for CONF3/rrho_part2: 0.18207391
The lowlevel G_mRRHO calculation @ c1 was successful for CONF4/rrho_part2: 0.18117449
The lowlevel G_mRRHO calculation @ c1 was successful for CONF5/rrho_part2: 0.18231888
The lowlevel G_mRRHO calculation @ c1 was successful for CONF6/rrho_part2: 0.18172189
The lowlevel G_mRRHO calculation @ c1 was successful for CONF8/rrho_part2: 0.18218769
The lowlevel G_mRRHO calculation @ c1 was successful for CONF10/rrho_part2: 0.17889859
The lowlevel G_mRRHO calculation @ c1 was successful for CONF11/rrho_part2: 0.18153296
The lowlevel G_mRRHO calculation @ c1 was successful for CONF15/rrho_part2: 0.18070768
--------------------------------------------------
* Gibbs free energies of part2 *
--------------------------------------------------
CONF# E(GFNn-xTB) ΔE(GFNn-xTB) E [Eh] Gsolv [Eh] GmRRHO [Eh] Gtot ΔGtot Boltzmannweight
[a.u.] [kcal/mol] r2scan-3c COSMO-RS-normal GFN2 [Eh] [kcal/mol] % at 298.15 K
[r2scan-3c] [alpb]-bhess
CONF2 -34.1594484 0.32 -497.2837810 -0.1358984 0.1802847 -497.2393947 0.17 21.10
CONF3 -34.1592530 0.44 -497.2915335 -0.1286518 0.1820739 -497.2381114 0.98 5.42
CONF4 -34.1590954 0.54 -497.2853431 -0.1353400 0.1811745 -497.2395086 0.10 23.80
CONF5 -34.1589250 0.65 -497.2917403 -0.1294025 0.1823189 -497.2388239 0.53 11.53
CONF6 -34.1587465 0.76 -497.2896963 -0.1298058 0.1817219 -497.2377802 1.19 3.82
CONF8 -34.1571241 1.78 -497.2905928 -0.1295961 0.1821877 -497.2380012 1.05 4.82
CONF10 -34.1563217 2.28 -497.2591855 -0.1593829 0.1788986 -497.2396698 0.00 28.23 <------
CONF11 -34.1559251 2.53 -497.2848139 -0.1313112 0.1815330 -497.2345922 3.19 0.13
CONF15 -34.1529637 4.39 -497.2792896 -0.1380698 0.1807077 -497.2366518 1.89 1.15
Calculating ALPB_Gsolv values for evaluation of the std. dev. of Gsolv.
Constructed folders!
Starting 9 ALPB-Gsolv calculations
Running ALPB_GSOLV calculation in 3-part0-2/CONF2/alpb_gsolv
Running ALPB_GSOLV calculation in 3-part0-2/CONF3/alpb_gsolv
Running ALPB_GSOLV calculation in 3-part0-2/CONF4/alpb_gsolv
Running ALPB_GSOLV calculation in 3-part0-2/CONF5/alpb_gsolv
Running ALPB_GSOLV calculation in 3-part0-2/CONF6/alpb_gsolv
Running ALPB_GSOLV calculation in 3-part0-2/CONF8/alpb_gsolv
Running ALPB_GSOLV calculation in 3-part0-2/CONF10/alpb_gsolv
Running ALPB_GSOLV calculation in 3-part0-2/CONF11/alpb_gsolv
Running ALPB_GSOLV calculation in 3-part0-2/CONF15/alpb_gsolv
Tasks completed!
ALPB_GSOLV calculation was successful for CONF2/alpb_gsolv: -0.14572423
ALPB_GSOLV calculation was successful for CONF3/alpb_gsolv: -0.14136373
ALPB_GSOLV calculation was successful for CONF4/alpb_gsolv: -0.14534319
ALPB_GSOLV calculation was successful for CONF5/alpb_gsolv: -0.14108828
ALPB_GSOLV calculation was successful for CONF6/alpb_gsolv: -0.14215086
ALPB_GSOLV calculation was successful for CONF8/alpb_gsolv: -0.14150861
ALPB_GSOLV calculation was successful for CONF10/alpb_gsolv: -0.16262359
ALPB_GSOLV calculation was successful for CONF11/alpb_gsolv: -0.14230853
ALPB_GSOLV calculation was successful for CONF15/alpb_gsolv: -0.14881714
SD of solvation models (all units in kcal/mol):
CONFX ΔG(COSMO-RS) ΔG(DCOSMO-RS_gsolv) ΔG(ALPB_gsolv) SD(COSMO-RS 40%, DCOSMO-RS_gsolv 40%, ALPB_gsolv 20%)
----------------------------------------------------------------------------------------------------
CONF2 14.74 17.32 10.60 3.01
CONF3 19.28 24.30 13.34 4.97
CONF4 15.09 17.98 10.84 3.21
CONF5 18.81 23.44 13.51 4.51
CONF6 18.56 22.88 12.85 4.53
CONF8 18.69 23.27 13.25 4.54
CONF10 0.00 0.00 0.00 0.00
CONF11 17.62 21.74 12.75 4.08
CONF15 13.37 15.51 8.66 3.06
----------------------------------------------------------------------------------------------------
Calculating Boltzmann averaged free energy of ensemble!
temperature /K: avE(T) /a.u. avGmRRHO(T) /a.u. avGsolv(T) /a.u. avG(T) /a.u.
----------------------------------------------------------------------------------------------------
273.15 -497.2736863 0.1842434 -0.1489372 -497.2383801
278.15 -497.2749007 0.1835285 -0.1471130 -497.2384852
283.15 -497.2760521 0.1828067 -0.1453716 -497.2386171
288.15 -497.2771316 0.1820765 -0.1437255 -497.2387805
293.15 -497.2781302 0.1813377 -0.1421818 -497.2389743
298.15 -497.2790497 0.1805897 -0.1407374 -497.2391974 <<==part2==
303.15 -497.2798874 0.1798315 -0.1393937 -497.2394496
308.15 -497.2806529 0.1790640 -0.1381406 -497.2397295
313.15 -497.2813425 0.1782861 -0.1369800 -497.2400363
318.15 -497.2819636 0.1774979 -0.1359037 -497.2403694
323.15 -497.2825199 0.1767001 -0.1349073 -497.2407270
328.15 -497.2830226 0.1758934 -0.1339792 -497.2411084
333.15 -497.2834731 0.1750769 -0.1331172 -497.2415134
338.15 -497.2838754 0.1742509 -0.1323164 -497.2419409
343.15 -497.2842354 0.1734163 -0.1315708 -497.2423900
348.15 -497.2845604 0.1725735 -0.1308732 -497.2428601
353.15 -497.2848497 0.1717219 -0.1302228 -497.2433506
358.15 -497.2851101 0.1708621 -0.1296129 -497.2438610
363.15 -497.2853448 0.1699946 -0.1290401 -497.2443904
368.15 -497.2855549 0.1691186 -0.1285027 -497.2449389
373.15 -497.2857417 0.1682355 -0.1279985 -497.2455047
----------------------------------------------------------------------------------------------------
--------------------------------------------------
Conformers considered further
--------------------------------------------------
Conformers that are below the Boltzmann threshold G_thr(2) of 99.0%:
CONF10, CONF4, CONF2, CONF5, CONF3, CONF8, CONF6, CONF15
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>END of Part2<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
Ran part2 in 5486.1036 seconds
Part : #conf time
--------------------------------------------------
Input : 22 -
Part0_all : 13 25.32s
Part1_initial_sort : 11 -
Part1_all : 11 166.42s
Part2_opt : 9 -
Part2_all : 8 5486.10s
--------------------------------------------------
All parts : 5677.84s
CENSO all done!
$CENSO global configuration file: .censorc
$VERSION:1.0.3
ORCA: /home/$USER/orca_4_2_1_linux_x86-64_openmpi216
ORCA version: 4.2.1
GFN-xTB: /home/$USER/bin/xtb
CREST: /home/$USER/bin/crest
mpshift: mpshift
escf: escf
#COSMO-RS
ctd = BP_TZVPD_FINE_C30_1601.ctd cdir = "/home/$USER/COSMOlogic/COSMOthermX19/COSMOtherm/CTDATA-FILES" ldir = "/home/$USER/COSMOlogic/COSMOthermX19/COSMOtherm/CTDATA-FILES"
cosmothermversion: 19
$ENDPROGRAMS
$CRE SORTING SETTINGS:
$GENERAL SETTINGS:
nconf: all # ['all', 'number e.g. 10 up to all conformers']
charge: 1 # ['number e.g. 0']
unpaired: 0 # ['number e.g. 0']
solvent: h2o # ['gas', 'acetone', 'chcl3', 'acetonitrile', 'ch2cl2', 'dmso', 'h2o', 'methanol', 'thf', '...']
prog_rrho: xtb # ['xtb', 'prog']
temperature: 298.15 # ['temperature in K e.g. 298.15']
trange: [273.15, 378.15, 5] # ['temperature range [start, end, step]']
multitemp: on # ['on', 'off']
evaluate_rrho: on # ['on', 'off']
consider_sym: off # ['on', 'off']
bhess: on # ['on', 'off']
imagthr: automatic # ['automatic or e.g., -100 # in cm-1']
sthr: automatic # ['automatic or e.g., 50 # in cm-1']
scale: automatic # ['automatic or e.g., 1.0 ']
rmsdbias: off # ['on', 'off']
sm_rrho: alpb # ['alpb', 'gbsa']
check: on # ['on', 'off']
prog: tm # ['tm', 'orca']
func: r2scan-3c # ['pbe', 'b97-d', 'pbeh-3c', 'tpss', 'b97-d3', 'r2scan-3c', 'b97-3c']
basis: automatic # ['automatic', 'def2-mSVP', 'def2-mTZVP', 'def2-mTZVP', 'def2-TZVP', '...']
maxthreads: 4 # ['number of threads e.g. 2']
omp: 2 # ['number cores per thread e.g. 4']
cosmorsparam: automatic # ['automatic', '12-fine', '12-normal', '13-fine', '13-normal', '14-fine', '...']
$PART0 - CHEAP-PRESCREENING - SETTINGS:
part0: on # ['on', 'off']
func0: b97-d # ['pbeh-3c', 'b97-3c', 'b97-d3', 'pbe', 'r2scan-3c', 'tpss', 'b97-d']
basis0: def2-SV(P) # ['automatic', 'def2-mSVP', 'def2-mTZVP', 'def2-mTZVP', 'def2-TZVP', '...']
part0_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
part0_threshold: 4.0 # ['number e.g. 4.0']
$PART1 - PRESCREENING - SETTINGS:
# func and basis is set under GENERAL SETTINGS
part1: on # ['on', 'off']
smgsolv1: cosmors # ['alpb_gsolv', 'dcosmors', 'cosmo', 'smd', 'cosmors-fine', 'cpcm', 'gbsa_gsolv', '...']
part1_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
part1_threshold: 3.5 # ['number e.g. 5.0']
$PART2 - OPTIMIZATION - SETTINGS:
# func and basis is set under GENERAL SETTINGS
part2: on # ['on', 'off']
opt_limit: 2.5 # ['number e.g. 4.0']
sm2: dcosmors # ['cosmo', 'cpcm', 'default', 'smd', 'dcosmors']
smgsolv2: cosmors # ['cosmors-fine', 'gbsa_gsolv', 'cosmo', 'cpcm', 'smd_gsolv', 'alpb_gsolv', 'smd', '...']
part2_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
ancopt: on # ['on']
hlow: 0.01 # ['lowest force constant in ANC generation, e.g. 0.01']
opt_spearman: on # ['on', 'off']
part2_threshold: 99 # ['Boltzmann sum threshold in %. e.g. 95 (between 1 and 100)']
optlevel2: automatic # ['crude', 'sloppy', 'loose', 'lax', 'normal', 'tight', 'vtight', 'extreme', '...']
optcycles: 8 # ['number e.g. 5 or 10']
spearmanthr: -4.0 # ['value between -1 and 1, if outside set automatically']
radsize: 10 # ['number e.g. 8 or 10']
crestcheck: off # ['on', 'off']
$PART3 - REFINEMENT - SETTINGS:
part3: off # ['on', 'off']
prog3: prog # ['tm', 'orca', 'prog']
func3: pw6b95 # ['dsd-blyp', 'pw6b95', 'b97-d3', 'r2scan-3c', 'pbe0', 'wb97x']
basis3: def2-TZVPD # ['SVP', 'SV(P)', 'TZVP', 'TZVPP', 'QZVP', 'QZVPP', 'def2-SV(P)', 'def2-mSVP', '...']
smgsolv3: cosmors # ['alpb_gsolv', 'dcosmors', 'cosmo', 'smd', 'cosmors-fine', 'cpcm', 'gbsa_gsolv', '...']
part3_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
part3_threshold: 99 # ['Boltzmann sum threshold in %. e.g. 95 (between 1 and 100)']
$NMR PROPERTY SETTINGS:
$PART4 SETTINGS:
part4: off # ['on', 'off']
couplings: on # ['on', 'off']
progJ: prog # ['tm', 'orca', 'adf', 'prog']
funcJ: pbe0 # ['tpss', 'pbe0', 'pbeh-3c']
basisJ: def2-TZVP # ['SVP', 'SV(P)', 'TZVP', 'TZVPP', 'QZVP', 'QZVPP', 'def2-SV(P)', 'def2-mSVP', '...']
sm4J: default # ['dcosmors', 'smd', 'cosmo', 'cpcm']
shieldings: on # ['on', 'off']
progS: prog # ['tm', 'orca', 'adf', 'prog']
funcS: pbe0 # ['dsd-blyp', 'pbeh-3c', 'tpss', 'pbe0', 'kt2']
basisS: def2-TZVP # ['SVP', 'SV(P)', 'TZVP', 'TZVPP', 'QZVP', 'QZVPP', 'def2-SV(P)', 'def2-mSVP', '...']
sm4S: default # ['dcosmors', 'smd', 'cosmo', 'cpcm']
reference_1H: TMS # ['TMS']
reference_13C: TMS # ['TMS']
reference_19F: CFCl3 # ['CFCl3']
reference_29Si: TMS # ['TMS']
reference_31P: TMP # ['TMP', 'PH3']
1H_active: on # ['on', 'off']
13C_active: on # ['on', 'off']
19F_active: off # ['on', 'off']
29Si_active: off # ['on', 'off']
31P_active: off # ['on', 'off']
resonance_frequency: 300.0 # ['MHz number of your experimental spectrometer setup']
$OPTICAL ROTATION PROPERTY SETTINGS:
$PART5 SETTINGS:
optical_rotation: off # ['on', 'off']
funcOR: pbe # ['functional for opt_rot e.g. pbe']
funcOR_SCF: r2scan-3c # ['functional for SCF in opt_rot e.g. r2scan-3c']
basisOR: def2-SVPD # ['basis set for opt_rot e.g. def2-SVPD']
frequency_optical_rot: [589.0] # ['list of freq in nm to evaluate opt rot at e.g. [589, 700]']
$END CENSORC
25
-34.15995484
O -2.7404108553 -0.9210263756 0.0505546155
O -0.6532148483 -1.7303331963 0.1381884568
N -2.1811530948 1.4662136641 -0.3280604715
N 1.9999042791 -1.6115147219 -0.1737452561
C 1.2832974514 1.7494376339 0.2478643443
C 0.0382242755 1.0547713107 0.8044626435
C 2.1972182535 0.8461961224 -0.5852172042
C -0.9624505804 0.6276788621 -0.2713633850
C 2.7304045776 -0.3664138808 0.1827513585
C -1.5037598584 -0.8390454546 0.0048901917
H 1.8561477225 2.1402514306 1.0892838382
H 0.9852377848 2.6009114549 -0.3642611073
H -0.4518270956 1.7282098745 1.5089809935
H 0.3322545004 0.1775252924 1.3825123935
H 1.6820235078 0.5059218506 -1.4832357051
H 3.0380251642 1.4546200515 -0.9165284484
H -0.4868586610 0.6089411942 -1.2530945560
H 2.6388482788 -0.2021893439 1.2562535001
H 3.7853986884 -0.5188774904 -0.0473087348
H -2.2380899515 2.1426216511 0.4407329422
H -2.3218532286 1.9466721688 -1.2205963738
H 2.3394829642 -2.3992859458 0.3882351555
H 0.9350292280 -1.5547588441 -0.0165809300
H 2.1475890354 -1.8368760327 -1.1636223077
H -2.9328940343 0.6971914121 -0.1921053926
25
-34.15944839
O -2.5910945174 -0.7526707388 0.4737473250
O -0.5258609320 -1.6055473558 0.4259946366
N -2.1883047577 1.2429283468 -0.9276187078
N 2.0221224751 -1.5482527990 -0.3635876013
C 1.0959612422 0.9269941773 1.0928268792
C -0.1118183900 1.5287127427 0.3646185610
C 2.3883955950 0.8232150730 0.2771806877
C -0.9137052232 0.5859776609 -0.5447048684
C 2.3984857923 -0.1999516224 -0.8542391926
C -1.3796771114 -0.7342464594 0.2052799197
H 0.8202702309 -0.0346561349 1.5230052762
H 1.3150554091 1.5881707649 1.9329336393
H 0.2252898792 2.3837647705 -0.2251981715
H -0.7970460540 1.8894152433 1.1356879771
H 2.6174364896 1.7989347868 -0.1517789136
H 3.1942064704 0.5825472961 0.9724285437
H -0.3499369029 0.3059081761 -1.4335569624
H 3.4048136146 -0.2402683769 -1.2749539435
H 1.7122153210 0.0852818535 -1.6492945642
H -2.2438155060 2.2226460139 -0.6308218074
H -2.4094139539 1.1700691932 -1.9248068681
H 0.9927667520 -1.5675551657 -0.0428313977
H 2.1377830502 -2.2542373686 -1.0973459319
H 2.6054467969 -1.8083924261 0.4393088723
H -2.8796747565 0.6310313638 -0.3580585669
25
-34.15925304
O -2.6152400314 -1.0465052003 0.2378304729
O -0.5362082855 -1.4663149505 -0.4593671015
N -2.4557715395 1.4039918868 0.4362491058
N 2.1447268012 -1.5911295291 -0.0908488427
C 1.1849792140 1.1291603967 -0.7123125066
C 0.0549239500 1.3190958463 0.3109988160
C 2.5516538580 0.8614163999 -0.0796419105
C -1.2770304521 0.8481909298 -0.2699298296
C 2.6278492228 -0.4408272994 0.7143702153
C -1.4798499370 -0.7275247362 -0.1594413730
H 1.2638776944 2.0288103522 -1.3231621409
H 0.9216680773 0.3153070715 -1.3864974308
H -0.0132519646 2.3752873127 0.5749134480
H 0.2604586805 0.7576942571 1.2223482952
H 3.2969117377 0.8423750312 -0.8755852689
H 2.8115908685 1.6810094994 0.5898815386
H -1.3353089850 1.1014685130 -1.3310936133
H 2.0240292821 -0.3842292575 1.6193674755
H 3.6638250279 -0.6160286136 1.0070988283
H -2.2160156909 1.7709567167 1.3631094761
H -2.9723151833 2.1102297821 -0.0929227069
H 2.6253531272 -1.6216026098 -0.9974699479
H 2.3064470721 -2.4761725884 0.4009650983
H 1.0998172314 -1.5111491960 -0.2664030290
H -3.0342449423 0.4895941432 0.5473101358
25
-34.15909539
O -2.5859531813 -0.8098009525 0.4685129327
O -0.4816245052 -1.5436026390 0.3102868940
N -2.3339104715 1.3007756781 -0.7836622820
N 2.0492564794 -1.5871472251 -0.5900906398
C 1.1052196056 0.9870949301 0.9271737021
C -0.2140772115 1.5925038037 0.4451868362
C 2.1667497593 0.8464686753 -0.1771749747
C -1.0154302176 0.6861215046 -0.4984780600
C 2.9073785151 -0.4823039670 -0.0859386105
C -1.3840762559 -0.7076318190 0.1707141950
H 0.8920163207 0.0201435075 1.3815317896
H 1.4930774855 1.6249540613 1.7213050551
H -0.0067258320 2.5368714102 -0.0630175124
H -0.8282372023 1.7972548662 1.3249938991
H 1.7135886317 0.9350726379 -1.1636100949
H 2.8915967677 1.6540129472 -0.0882838850
H -0.4677225963 0.5007855037 -1.4227063242
H 3.1688824777 -0.6847294678 0.9524589517
H 3.8237876908 -0.4492687854 -0.6762385697
H -2.5870605138 1.3036833133 -1.7755142934
H -2.9689761577 0.6014426661 -0.2449843511
H 1.9723283573 -1.5417442554 -1.6118847971
H 2.4360322309 -2.5007606995 -0.3327999527
H 1.0548817542 -1.5294780724 -0.1881461707
H -2.4296444696 2.2452869585 -0.3976639893
25
-34.15892503
O -2.4565204045 -0.8423454324 0.7070928712
O -0.5980888346 -1.6084799899 -0.2741310513
N -2.2081439307 1.5499049911 0.0797136762
N 2.0665657616 -1.5738488309 0.1665385721
C 0.8489992162 1.1021214800 0.5754910780
C 0.1319038004 1.1643519245 -0.7806607655
C 2.3611287197 0.8895018711 0.4504901307
C -1.3084663819 0.6750096988 -0.7114456668
C 2.7727042231 -0.3622083435 -0.3226992137
C -1.4583757981 -0.7537800183 -0.0274928276
H 0.4155006230 0.3036523983 1.1797441237
H 0.7020515096 2.0340597672 1.1228295970
H 0.6304477123 0.5210588124 -1.5026223849
H 0.1602968624 2.1778026423 -1.1819014190
H 2.8058653217 1.7473421346 -0.0537220547
H 2.7810957945 0.8489865537 1.4562868383
H -1.7106345304 0.5638398621 -1.7216356404
H 3.8478441803 -0.5048945341 -0.2050768697
H 2.5630334022 -0.2516058916 -1.3860041489
H -2.8269415029 2.1396743466 -0.4823076953
H -2.7642911303 0.7978509993 0.6164252281
H 2.4375926491 -2.4160896797 -0.2859521821
H 2.1793786728 -1.6693140839 1.1823811296
H 1.0245787819 -1.5392035408 -0.0507409777
H -1.6858363886 2.1297513428 0.7452520673
25
-34.15874648
O -2.3605981473 -0.8116042758 0.8659409274
O -0.5843274484 -1.6230194998 -0.2244865945
N -2.2109877661 1.5364887021 0.0742080589
N 2.0757147084 -1.4672125485 -0.4830865528
C 0.8597549995 1.2262238823 0.4517476395
C 0.0874032105 1.1513231513 -0.8771132949
C 2.3512317789 0.8985689618 0.2982329522
C -1.3361657088 0.6347191585 -0.7132778341
C 2.6824356905 -0.5712975375 0.5355813092
C -1.4234260659 -0.7566236862 0.0528412632
H 0.4221004807 0.5356789162 1.1753502177
H 0.7733707657 2.2309740830 0.8650565898
H 0.5827203837 0.4802075195 -1.5756071724
H 0.0631682297 2.1335262804 -1.3514203271
H 2.7003072358 1.2129278451 -0.6855204577
H 2.9172351560 1.4667500260 1.0360745366
H -1.7833398578 0.4616073570 -1.6955611361
H 2.3142140534 -0.8770131145 1.5154931358
H 3.7656889000 -0.7005750162 0.5151167425
H -1.6671444199 2.1735125324 0.6661186569
H -2.8827343086 2.0695764162 -0.4835506438
H 2.3643871815 -1.1879653946 -1.4273853498
H 2.3725094498 -2.4360036549 -0.3237631449
H 1.0071911744 -1.4659435853 -0.4227446705
H -2.7090441201 0.8070062032 0.6949537917
25
-34.15780648
O -0.6998712600 -1.7595821198 0.3315266042
O -2.7564980797 -0.8922468771 0.1127087169
N -2.0912345729 1.3897515697 -0.6207661035
N 1.9218668343 -1.5538051846 -0.0911139226
C 1.2724611881 1.7665596691 0.2614262584
C -0.0126245071 1.0963347719 0.7686474620
C 2.5091659413 0.8642624497 0.2746224048
C -0.9098130242 0.5407794733 -0.3415020783
C 2.4929985644 -0.3196472927 -0.6935656161
C -1.5185682699 -0.8616584773 0.0830057999
H 1.4961397665 2.6156227165 0.9073501428
H 1.1219744652 2.1603919352 -0.7434483203
H -0.5894557289 1.8123867185 1.3552290309
H 0.2456564204 0.2915541326 1.4587568868
H 3.3629575544 1.4905499194 0.0148013745
H 2.6790096911 0.5030846620 1.2896168914
H -0.3423105707 0.3842564461 -1.2596402044
H 3.5203242755 -0.5424656780 -0.9864282797
H 1.9321769951 -0.0751719518 -1.5947809502
H -2.8797605313 0.6907420298 -0.3760235139
H -2.1503924665 2.2120232522 -0.0104200923
H 2.3803263882 -1.7503210865 0.8055549093
H 0.8561607870 -1.5268128372 0.0892049642
H 2.0800598725 -2.3543999415 -0.7123089418
H -2.1774862389 1.6805852373 -1.5986092278
25
-34.15712409
O -0.5169415866 -1.4677953372 -0.4155166026
O -2.6718105717 -1.1029442546 0.0532698902
N -2.5041882719 1.3222450456 0.5356135683
N 2.0547500815 -1.4660199090 0.3952674841
C 1.1460672076 1.2350620947 -0.5905524477
C 0.0199991306 1.2471137062 0.4571994904
C 2.5324119546 0.9449216441 0.0010319703
C -1.3028765754 0.8219664596 -0.1753086903
C 2.9759584940 -0.4928036507 -0.2449354813
C -1.5027299259 -0.7570468322 -0.1889883845
H 1.1567284127 2.2014747982 -1.0932584224
H 0.9206483903 0.4869950226 -1.3509226784
H -0.0730555237 2.2557886152 0.8619704582
H 0.2503077610 0.5750867956 1.2847618985
H 3.2724883861 1.5930094341 -0.4660504111
H 2.5411017268 1.1667575642 1.0680525607
H -1.3439398378 1.1490834108 -1.2170635960
H 3.9809942022 -0.6416689206 0.1522940223
H 2.9906906878 -0.6917352586 -1.3167852841
H -3.1147358709 0.4302326474 0.4978236161
H -2.3055244586 1.5552316017 1.5147727778
H 1.0589542040 -1.3722826387 0.0237179616
H 2.3537713222 -2.4274392985 0.2004063016
H 2.0384435533 -1.3281897829 1.4119124615
H -2.9671283162 2.1123945703 0.0792841186
25
-34.15703719
O -0.4840204654 -1.5908390855 -0.1977054306
O -2.6424172363 -1.0172969494 -0.2313294170
N -2.3824391815 1.4118161237 0.1129756179
N 2.0591606020 -1.4080559447 0.6444993748
C 1.3255949204 1.5932768744 0.1316443482
C 0.0292286137 1.1405631347 0.8180158808
C 1.9611296764 0.5914767293 -0.8382657525
C -1.0774245534 0.7702791240 -0.1744053472
C 2.8498636045 -0.4645206094 -0.1922799795
C -1.4218007565 -0.7850501416 -0.2021503198
H 2.0511714191 1.8626404605 0.8996124027
H 1.1017826165 2.5033885185 -0.4265728164
H -0.3083479528 1.9763847128 1.4332069953
H 0.2033356421 0.3027453082 1.4918974312
H 1.1939462785 0.0852292002 -1.4240400771
H 2.5871303840 1.1474547523 -1.5362113415
H -0.7775416723 1.0312499156 -1.1919311777
H 3.6205966803 0.0055444316 0.4194317100
H 3.3361661233 -1.0318279746 -0.9856062704
H -2.6284628403 2.1753990573 -0.5216811303
H -3.0325627338 0.5540981187 -0.0284710758
H 2.4910941867 -2.3367192092 0.6680136974
H 1.9781398838 -1.0663385144 1.6071566991
H 1.0702227114 -1.5059322479 0.2455456256
H -2.4592842029 1.7302943098 1.0848423546
25
-34.15632171
O -0.6377330316 -1.5602972801 -0.5925407457
O -1.9438226066 -0.8962122015 1.0968963302
N -2.0691310397 1.5115608178 0.5520085904
N 2.0161394685 -1.6864701960 -0.6808935380
C 1.0245208700 1.5678424724 0.3114580637
C 0.0268814843 1.3822638313 -0.8376423667
C 1.7125418505 0.2918072015 0.8062089597
C -1.3255436976 0.7602094261 -0.4861917907
C 2.5798867778 -0.3735189736 -0.2699515146
C -1.2821941821 -0.7269243894 0.0579815663
H 0.5483025277 2.0449086772 1.1692286093
H 1.7966540418 2.2561425086 -0.0359536522
H 0.4649670843 0.7632830938 -1.6196837604
H -0.1661732017 2.3619672870 -1.2796881790
H 2.3322565126 0.5621906956 1.6605054657
H 0.9655697921 -0.4140620317 1.1690705974
H -1.9201614836 0.7061105980 -1.4020292431
H 3.5936748701 -0.5312821922 0.0979019662
H 2.6369135707 0.2686143194 -1.1490685384
H -2.9086496435 1.9840429736 0.2094100180
H -2.3313717130 0.6888337551 1.2084854943
H 2.1820919942 -2.3913736082 0.0460188498
H 0.9581287874 -1.6201065231 -0.7926007692
H 2.4297604389 -2.0163864988 -1.5591512145
H -1.4715716162 2.1835379113 1.0453989384
25
-34.15592507
O -2.1033969149 -0.8344078612 0.9487088877
O -0.5699612374 -1.6680274471 -0.4517081377
N -1.9569275917 1.5465619838 0.2627686808
N 2.0837337252 -1.3633283870 -0.5773762363
C 1.1075416509 1.5419562581 -0.0103918263
C 0.1209945172 1.1686244794 -1.1251953409
C 1.4352918321 0.4570859531 1.0214669735
C -1.2503428799 0.6557309102 -0.6867527508
C 2.5057372966 -0.5407868185 0.5902823997
C -1.2948162683 -0.7745608298 0.0075959523
H 0.7274518351 2.4141375453 0.5251162579
H 2.0330446007 1.8691474761 -0.4864359338
H 0.5312965142 0.4086384701 -1.7869917742
H -0.0324587227 2.0621064981 -1.7347668750
H 1.8169519144 0.9463147999 1.9183948838
H 0.5367641743 -0.0848292604 1.3149274400
H -1.8634330878 0.5270669166 -1.5839659846
H 2.7015999487 -1.2099014501 1.4282318085
H 3.4296803314 -0.0167769980 0.3414844052
H -1.2992225514 2.1272618717 0.7958603647
H -2.6755557698 2.1370116562 -0.1630903688
H 2.2813676133 -0.8782194673 -1.4586367594
H 2.5735263109 -2.2635096262 -0.5875520766
H 1.0325521121 -1.5556317222 -0.5353874345
H -2.3935764356 0.8063761382 0.9145561672
25
-34.15547499
O -2.3288681540 -0.9689367039 0.5974852723
O -0.4008216845 -1.3559960891 -0.4559450313
N -2.6333104330 1.3901145964 0.0148791472
N 2.2688734235 -1.7836122490 -0.3135245436
C 1.2045970266 1.2147830910 -0.3207104722
C -0.1766849998 1.7049883679 0.1388725454
C 1.9421007389 0.4104322018 0.7578355455
C -1.3241367595 0.8966054975 -0.4749078790
C 2.9601024995 -0.5430248114 0.1410496733
C -1.3285933693 -0.6410895869 -0.0712604222
H 1.8194085679 2.0702167267 -0.5988277282
H 1.0710671602 0.6065579215 -1.2146344993
H -0.3076575365 2.7461839050 -0.1612866699
H -0.2378627475 1.6551531995 1.2275331138
H 2.4468113264 1.0960018700 1.4370167059
H 1.2297127637 -0.1683684162 1.3468389979
H -1.2704988015 0.9491259299 -1.5640455987
H 3.7310656000 -0.8072491298 0.8640696447
H 3.4371256948 -0.0632845904 -0.7133472266
H -2.5432643621 2.1520849835 0.6930123940
H -3.2834926874 1.6599628672 -0.7262042831
H 2.7178100064 -2.1982719631 -1.1361624210
H 2.2482990815 -2.4828183187 0.4368503414
H 1.2567750610 -1.5723871751 -0.5319861416
H -2.9757946227 0.4721100866 0.5082732629
25
-34.15545846
O -2.3777023451 -0.9177733836 0.6489622803
O -0.4009151018 -1.4460423055 -0.2363506290
N -2.5928675119 1.3702602408 -0.1958788709
N 2.2705863538 -1.5552728060 -0.6959671399
C 1.2626408325 1.2650767695 -0.1999508619
C -0.1676900608 1.6997857440 0.1433859727
C 1.8576282855 0.3222355626 0.8537184709
C -1.2484356318 0.8255651567 -0.5054674146
C 2.9084111559 -0.6168744122 0.2764230940
C -1.3258577703 -0.6732783629 0.0271901706
H 1.8935551201 2.1501893106 -0.2772016053
H 1.2458750287 0.7908735642 -1.1798122422
H -0.3171466901 2.7186794233 -0.2201486160
H -0.2967658760 1.7007241110 1.2272150872
H 2.3089550370 0.9081814099 1.6535209776
H 1.0686425282 -0.2818531316 1.3016188748
H -1.0876822336 0.7817119785 -1.5847014431
H 3.3524799930 -1.1924945383 1.0873745985
H 3.6963304583 -0.0519426265 -0.2212122945
H -2.9913275573 0.5162412164 0.3615908966
H -2.5589278358 2.2025641690 0.3999047288
H 2.6187881911 -2.5117770389 -0.5815509942
H 1.2217357961 -1.5619680721 -0.5321789754
H 2.4345460024 -1.2660811775 -1.6650051770
H -3.1696789754 1.5558016135 -1.0193997348
25
-34.15372845
O -0.8759513206 -1.0507714945 0.7395130487
O -1.0471783147 -1.2210634731 -1.4857197061
N -2.0100713682 1.1638582833 0.9733772433
N 1.7020599065 -1.7447966827 0.8365171560
C 0.9224701988 1.3977552196 0.1299066020
C -0.3034955554 1.7474745141 -0.7290850754
C 1.8313865205 0.3371413309 -0.5152183962
C -1.5397536849 0.8954837973 -0.4124258872
C 2.5286316072 -0.5462459399 0.5204181162
C -1.1337894558 -0.6241211220 -0.4224393440
H 0.5999134506 1.0447379113 1.1088504940
H 1.5075448530 2.3032957331 0.2891927248
H -0.0603736240 1.6138572758 -1.7832639404
H -0.5689030950 2.7975685720 -0.5933476217
H 1.2520636483 -0.2949117808 -1.1892484363
H 2.5821478388 0.8362393886 -1.1261103610
H -2.3194229349 1.0797505602 -1.1497937252
H 2.6951475316 0.0179430508 1.4383444706
H 3.4942793529 -0.8824584649 0.1428332329
H -1.6366862386 2.0406897131 1.3566364027
H -3.0319635647 1.1643591957 1.0502533535
H 2.0123953855 -2.1956055335 1.7033301803
H 0.6617699349 -1.4988916853 0.9180060433
H 1.7620357216 -2.4244612968 0.0700935319
H -1.6155014744 0.3110635636 1.4848864268
25
-34.15296366
O -0.9806508876 -0.9847570656 0.9016095316
O -1.0342961735 -1.4975369515 -1.2790158618
N -2.0600699488 1.3003330160 0.6598555012
N 1.6483681534 -1.4770759046 1.0193012553
C 0.8930197778 1.4409262402 0.3162871289
C -0.1018287155 1.4892580056 -0.8622608316
C 2.2127299908 0.7309357778 -0.0043863659
C -1.4370888955 0.7837090011 -0.5889439421
C 2.1074033787 -0.7768274559 -0.2134156979
C -1.1447588094 -0.7374998177 -0.3265126418
H 0.4424872625 0.9591479756 1.1822965353
H 1.1476108940 2.4604379775 0.6056686014
H 0.3344753521 1.0203527521 -1.7446629435
H -0.3104542151 2.5259312846 -1.1308040482
H 2.6400205950 1.1617275571 -0.9101356051
H 2.9157320587 0.9267783500 0.8074716570
H -2.0965188879 0.8923814409 -1.4486238017
H 3.0903725751 -1.1592546293 -0.4905246456
H 1.4081882505 -1.0149227036 -1.0156071158
H -3.0831759004 1.3363322990 0.6000761811
H -1.7780292676 0.5566802205 1.3629478428
H 0.5935972201 -1.3552723672 1.1310943683
H 1.8230458958 -2.4842696303 0.9418471118
H 2.1352867113 -1.1124232113 1.8467861942
H -1.6994055067 2.2241075607 0.9291898687
25
-34.15141381
O -1.3943785432 -1.0307567325 0.9025777408
O -0.4225106153 -1.2766835982 -1.0942963862
N -2.1382332497 1.3072100696 0.6820859684
N 1.8303685962 -1.9710495392 0.0707587666
C 0.9607624662 1.6168605590 0.1707894371
C -0.1959862014 1.7114505222 -0.8393306352
C 1.9799050629 0.5013843169 -0.1348429740
C -1.4247901716 0.8500833461 -0.5368218294
C 1.9371661639 -0.6960029627 0.8273160062
C -1.0467465663 -0.6453837331 -0.2300826075
H 0.5821582777 1.4936196592 1.1855414824
H 1.4853467761 2.5717069600 0.1437592453
H 0.1684852019 1.4328271659 -1.8279890567
H -0.5295558410 2.7487716363 -0.9068101296
H 1.7990108513 0.1423863706 -1.1479579769
H 2.9805578856 0.9303497506 -0.1217137710
H -2.0885910031 0.8601550715 -1.4024758281
H 1.0694995988 -0.6377417602 1.4853144468
H 2.8334350246 -0.7245407799 1.4470296049
H -1.6564311817 2.0816802979 1.1506799283
H -3.1156981590 1.5603768219 0.5197984898
H 2.6238854975 -2.0888513426 -0.5678832194
H 1.7803639034 -2.7755595157 0.7040003308
H 0.9218328585 -1.9204346441 -0.4990801476
H -2.0745975566 0.3957317695 1.2728068192
25
-34.15122546
O -1.4776298354 -0.9779162840 0.9689911790
O -0.4925234496 -1.3628551244 -1.0036507954
N -2.1639329473 1.3683833598 0.5828976748
N 1.8796097534 -1.9018199765 -0.0324662357
C 0.9212842989 1.5787487906 0.2022882964
C -0.1575522111 1.5949891498 -0.8983466380
C 2.0807266384 0.5903317386 -0.0312863938
C -1.4336452672 0.8124987799 -0.5833640723
C 2.0372506913 -0.6914975753 0.8153963432
C -1.1077721831 -0.6712596326 -0.1784565029
H 0.4772061571 1.3762504059 1.1769048440
H 1.3491352329 2.5796760629 0.2516191005
H 0.2519128515 1.1866703660 -1.8221067218
H -0.4443786804 2.6264492546 -1.1098247151
H 2.1069094344 0.3214704229 -1.0870338715
H 3.0133169149 1.1098047495 0.1839278312
H -2.0690635873 0.7917157945 -1.4696202320
H 1.1994207751 -0.6693189818 1.5132782369
H 2.9569334194 -0.7921592267 1.3922309558
H -1.6800036187 2.1679532791 1.0053931771
H -3.1334683512 1.6233161070 0.3791665015
H 2.6232513605 -1.9579134554 -0.7362257317
H 1.8912989959 -2.7525757838 0.5392212719
H 0.9273933706 -1.8444066710 -0.5257136187
H -2.1333692549 0.5050567581 1.2388793063
25
-34.15102491
O -2.7007099152 -1.6852260154 -0.4665848832
O -3.5578446821 0.0350414384 0.6950996250
N -1.7249631955 1.6316108345 0.4298941943
N 4.8503432460 -0.2765578553 0.0049566510
C 1.1047501969 0.4501828237 -0.3379055216
C -0.1445140391 -0.2886936630 0.1459845288
C 2.3741940866 -0.2828979224 0.0965187068
C -1.4339016427 0.3793165613 -0.3028003047
C 3.6160850376 0.4539254738 -0.3951671902
C -2.6934727541 -0.5488189533 -0.0039847520
H 1.1202182595 1.4641232269 0.0631636060
H 1.0744507790 0.5160807326 -1.4261342120
H -0.1466884742 -0.3764861277 1.2347858649
H -0.1614173698 -1.3033571375 -0.2567289346
H 2.3826839006 -0.3550960319 1.1849100754
H 2.3458274825 -1.2937888279 -0.3120785959
H -1.4193517541 0.5529473168 -1.3796978179
H 3.6476836700 1.4575725004 0.0303995802
H 3.5908364898 0.5356651831 -1.4823627548
H -1.7888195298 2.4639789876 -0.1581990321
H -2.7206446817 1.3385249836 0.8116492773
H 5.6915565940 0.2174183391 -0.3259194480
H 4.8962264202 -0.3605243899 1.0314916029
H 4.8443782262 -1.2261178281 -0.3963638456
H -1.0847775954 1.7904079765 1.2125175138
25
-34.15096274
O -3.0664396656 -0.6811514259 0.4732552569
O -1.1830540829 -1.6357953168 -0.2743493731
N -2.5677177347 1.6455787239 -0.0357930901
N 3.4161836207 -1.4947231742 0.0043983586
C 1.0526738400 0.4198619079 -0.3271341828
C -0.1392962419 1.1756334823 0.2601668795
C 2.3301787116 0.6870764296 0.4694516416
C -1.4374925629 0.7826136622 -0.4432938226
C 3.5433351448 -0.0155439223 -0.1263696890
C -1.9232732343 -0.6849210780 -0.0466890927
H 1.1914397265 0.7321338619 -1.3635085567
H 0.7858921189 -0.6373779302 -0.3204311762
H 0.0218209785 2.2505182304 0.1468292069
H -0.2266241551 0.9455046686 1.3247226359
H 2.5278796928 1.7590277621 0.4727119199
H 2.1882724928 0.3782271446 1.5058219746
H -1.3025118280 0.8032883597 -1.5259809861
H 4.4479425383 0.3027435209 0.3921583620
H 3.6346463266 0.2349775607 -1.1836334244
H -3.0211573542 2.1410074345 -0.8045922540
H -3.2218629847 0.8371482423 0.3587042034
H 4.2253281871 -1.9726822174 -0.4155872948
H 3.3489651634 -1.7581308116 0.9983748528
H 2.5570336567 -1.8144640232 -0.4679619280
H -2.3203269015 2.3008181314 0.7098687755
25
-34.15085064
O -3.5353488917 0.0322108098 0.4084906022
O -2.4254579214 -1.9084093375 0.6284254091
N -1.7744403653 1.3652775888 -0.6438663864
N 4.1843308757 -0.1227935311 -1.0149931633
C 1.2054477125 0.3338345853 -0.2573250215
C -0.0723683835 -0.0922253390 0.4701395235
C 2.4067626747 0.2476116899 0.6871054133
C -1.3095998224 -0.0201188789 -0.4106994862
C 3.6989393857 0.7735207317 0.0732627710
C -2.5481677099 -0.7347193516 0.2919952398
H 1.1039179513 1.3593179418 -0.6150772671
H 1.3305507433 -0.3269971917 -1.1156830038
H -0.2323641779 0.5179203592 1.3619095074
H 0.0148258366 -1.1272987239 0.8068454593
H 2.1939049448 0.8432977651 1.5748900270
H 2.5356727482 -0.7824695024 1.0211816116
H -1.1430786561 -0.5391980852 -1.3555500184
H 4.4681954117 0.8303339300 0.8434799998
H 3.5410323285 1.7707403176 -0.3387001734
H -2.7959421798 1.2370421526 -0.2418926774
H -1.2663143192 2.0543977636 -0.0827359543
H 4.3101055621 -1.0795234870 -0.6518667702
H 3.4997443822 -0.1538144186 -1.7841531054
H 5.0842467659 0.2120203831 -1.3873716549
H -1.8037773550 1.6476905491 -1.6249441216
25
-34.15048917
O -1.9774584219 -0.8822136242 -1.3447844876
O -0.7353222350 -1.3040874260 0.4712779300
N -1.2018711229 0.8204001109 1.6759883927
N 1.9464051556 -1.6923491374 0.3496868798
C 0.8058411616 1.0462579377 -0.9468506072
C -0.4754178942 1.7785998054 -0.5275051890
C 1.8226970853 0.7789686777 0.1752315710
C -1.5019161161 0.9155320919 0.2220611780
C 2.6576744598 -0.4659344426 -0.1177787426
C -1.4255171488 -0.5796126398 -0.2987743324
H 1.2927227489 1.6467822241 -1.7161034319
H 0.5115436171 0.1136959367 -1.4299162738
H -0.9626535770 2.1244807715 -1.4395462399
H -0.2313854006 2.6631384020 0.0636669169
H 2.4836824282 1.6401220787 0.2631946203
H 1.3379895832 0.6415415675 1.1385387246
H -2.5044279955 1.3042479639 0.0511624497
H 3.6220420553 -0.4105315490 0.3870533340
H 2.8346807725 -0.5412209492 -1.1903941633
H -0.5537202797 1.5469435382 2.0029686770
H -2.0510266978 0.8448716653 2.2497725116
H 0.8939724574 -1.6086997720 0.2334441992
H 2.2630899289 -2.5235198565 -0.1606712473
H 2.1155048181 -1.8450780227 1.3513380594
H -0.7798223195 -0.1709536166 1.7101046448
25
-34.15043307
O -3.3078583007 -0.3518099727 0.5480566241
O -1.8441149941 -2.0073969491 0.1397595703
N -1.9672328473 1.5403727742 -0.2372000800
N 3.6044925126 -0.8358440764 -0.6659647320
C 1.1684565146 1.0446332415 -0.4943940738
C 0.1082467096 0.2177955398 0.2395309885
C 2.4785715426 1.1657908669 0.2896724500
C -1.2458520269 0.2676289576 -0.4513144135
C 3.1696429470 -0.1592315605 0.5890169973
C -2.2263428078 -0.8414226742 0.1422529421
H 0.7932826990 2.0569151989 -0.6482792542
H 1.3354698371 0.6191303209 -1.4843783659
H -0.0031461344 0.5661490514 1.2686433794
H 0.3791631733 -0.8378892566 0.2841197320
H 3.1579098277 1.8188328559 -0.2597110183
H 2.2713923792 1.6478155935 1.2450536588
H -1.1432114953 0.0569418065 -1.5170262136
H 2.5004236326 -0.8240708177 1.1341470919
H 4.0493855213 0.0280822082 1.2049984895
H -1.5070190112 2.1488324010 0.4454597884
H -2.1837208522 2.0584218679 -1.0901715209
H 4.2094106392 -0.2068158824 -1.2153183037
H 4.1256865653 -1.6973771969 -0.4514692989
H 2.7834959501 -1.0854874671 -1.2358687369
H -2.8845837684 1.1083956166 0.2016053653
Calculation of NMR spectra
cis-4-hexen-1-ol
Example of calculating the 1H-NMR spectrum of cis-4-hexen-1-ol in CHCl3 at 400 MHz using TURBOMOLE
$ cat coord
$coord
-4.5787202885 -0.8553740252 1.3686940377 C
-3.4023148672 -0.1930996279 -1.1104654057 C
-1.3502307141 1.1896725766 -1.5219716436 C
0.3006494795 2.4620980359 0.3955352829 C
0.2362191223 4.5001215909 0.0503890823 H
3.0642161355 1.6025835718 0.2096773622 C
3.4178595291 -1.1774351958 0.9222138508 C
2.5415191057 -2.8708599609 -0.9480135970 O
0.8099880164 -2.4078504756 -1.3346662145 H
5.4277591776 -1.6058314070 1.1410607567 H
2.4658368243 -1.5681688895 2.7281288136 H
4.2123967195 2.7454371194 1.4856132673 H
3.7614131313 1.8836292778 -1.7127605110 H
-0.3830548458 2.1258051759 2.3091187341 H
-0.7518930894 1.4941446048 -3.4579714157 H
-4.3910561682 -0.9675515200 -2.7291675973 H
-3.5318795176 -0.1003881407 2.9630921609 H
-6.5027613245 -0.1227486813 1.4413316443 H
-4.6902514935 -2.9039535570 1.5605291206 H
$end
Input structure.
Start with your (at best already optimized) input structure and create the conformers
and rotamers for the CENSO and ANMR calculation.
$ crest coord -gfn2 -g chcl3 -T 4 -nmr > crest.out
In our case CREST found 86 conformers within an energy window of 6 kcal/mol.
We then create a new folder for the CENSO reranking and copy the necessary files:
$ mkdir censo
$ cp crest_conformers.xyz coord anmr_nucinfo anmr_rotamer censo/
$ cd censo/
CENSO requires only the file crest_conformers.xyz, but ANMR needs the last
three files. Here we want to calculate everything with TURBOMOLE using r2SCAN-3c and
DCOSMO-RS for geometry optimization and PBE0/def2-TZVP for the NMR part using the command line.
To save computational costs, a threshold of 95 percent of Boltzmann_weights is used in part 2
which is in most cases sufficient to reproduce the experimental spectrum. In our case,
this reduces the number of conformers in part 4 from 74 to 58.
Note
settings are read from global .censorc file (default settings in censo)
$ censo --input crest_conformers.xyz -solvent chcl3 --prog tm --thrpart2 95 -part4 on
-funcJ pbe0 -funcS pbe0 -basisJ def2-TZVP -basisS def2-TZVP -cactive off > censo.out
______________________________________________________________
| |
| |
| CENSO - Commandline ENSO |
| v 1.1.2 |
| energetic sorting of CREST Conformer Rotamer Ensembles |
| University of Bonn, MCTC |
| Feb 2021 |
| based on ENSO version 2.0.1 |
| F. Bohle and S. Grimme |
| |
|______________________________________________________________|
Please cite:
S. Grimme, F. Bohle, A. Hansen, P. Pracht, S. Spicher, and M. Stahn
J. Phys. Chem. A 2021, 125, 19, 4039-4054.
DOI: https://doi.org/10.1021/acs.jpca.1c00971
This program is distributed in the hope that it will be useful,
but WITHOUT ANY WARRANTY; without even the implied warranty of
MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE.
----------------------------------------------------------------------------------------------------
PARAMETERS
----------------------------------------------------------------------------------------------------
program call: censo --input crest_conformers.xyz -solvent chcl3 --prog tm -thrpart2 95 -part4 on -funcJ pbe0 -funcS pbe0 -basisJ def2-TZVP -basisS def2-TZVP -cactive off
The configuration file .censorc is read from /home/gorges/.censorc.
Reading conformer rotamer ensemble from: /tmp1/gorges/3870734.majestix.thch.uni-bonn.de/crest_conformers.xyz.
Reading file: censo_solvents.json
--------------------------------------------------
CRE SORTING SETTINGS
--------------------------------------------------
number of atoms in system: 19
number of considered conformers: 86
number of all conformers from input: 86
charge: 0
unpaired: 0
solvent: chcl3
temperature: 298.15
evaluate at different temperatures: on
temperature range: 273.15, 278.15, 283.15, 288.15, ...
calculate mRRHO contribution: on
consider symmetry for mRRHO contribution: on
cautious checking for error and failed calculations: on
checking the DFT-ensemble using CREST: off
maxthreads: 7
omp: 4
automatically balance maxthreads and omp: off
--------------------------------------------------
CRE CHEAP-PRESCREENING - PART0
--------------------------------------------------
part0: on
starting number of considered conformers: 86
program for part0: tm
functional for fast single-point: b97-d3(0)
basis set for fast single-point: def2-SV(P)
threshold g_thr(0) for sorting in part0: 4.0
Solvent model used with xTB: alpb
short-notation:
b97-d3(0)/def2-SV(P) // GFNn-xTB (Input geometry)
--------------------------------------------------
CRE PRESCREENING - PART1
--------------------------------------------------
part1: on
program for part1: tm
functional for initial evaluation: r2scan-3c
basis set for initial evaluation: def2-mTZVPP
calculate mRRHO contribution: on
program for mRRHO contribution: xtb
GFN version for mRRHO and/or GBSA_Gsolv: gfn2
Apply constraint to input geometry during mRRHO calculation: on
solvent model applied with xTB: alpb
evaluate at different temperatures: off
threshold g_thr(1) and G_thr(1) for sorting in part1: 3.5
solvent model for Gsolv contribution of part1: cosmors
short-notation:
r2scan-3c + COSMORS[chcl3] + GmRRHO(GFN2[alpb]-bhess) // GFNn-xTB (Input geometry)
--------------------------------------------------
CRE OPTIMIZATION - PART2
--------------------------------------------------
part2: on
program: tm
functional for part2: r2scan-3c
basis set for part2: def2-mTZVPP
using xTB-optimizer for optimization: on
using the new ensemble optimizer: on
optimize all conformers below this G_thr(opt,2) threshold: 2.5
spearmanthr: 0.939
optimization level in part2: lax
solvent model applied in the optimization: dcosmors
solvent model for Gsolv contribution: cosmors
evaluate at different temperatures: on
Boltzmann sum threshold G_thr(2) for sorting in part2: 95.0
calculate mRRHO contribution: on
program for mRRHO contribution: xtb
GFN version for mRRHO and/or GBSA_Gsolv: gfn2
Apply constraint to input geometry during mRRHO calculation: on
solvent model applied with xTB: alpb
short-notation:
r2scan-3c + COSMORS[chcl3] + GmRRHO(GFN2[alpb]-bhess) // r2scan-3c[DCOSMORS]
--------------------------------------------------
NMR MODE SETTINGS
--------------------------------------------------
part4: on
calculate couplings (J): on
program for coupling calculations: tm
solvation model for coupling calculations: dcosmors
functional for coupling calculation: pbe0
basis set for coupling calculation: def2-TZVP
calculate shieldings (S): on
program for shielding calculations: tm
solvation model for shielding calculations: dcosmors
functional for shielding calculation: pbe0
basis set for shielding calculation: def2-TZVP
Calculating proton spectrum: on
reference for 1H: TMS
resonance frequency: 300.0
END of parameters
------------------------------------------------------------
PATHS of external QM programs
------------------------------------------------------------
The following program paths are used:
xTB: /home/abt-grimme/AK-bin/xtb
TURBOMOLE: /home/abt-grimme/TURBOMOLE.7.5//bin/em64t-unknown-linux-gnu_smp
escf: /home/abt-grimme/TURBOMOLE.7.5/bin/em64t-unknown-linux-gnu/escf
mpshift: /home/abt-grimme/TURBOMOLE.7.5/bin/em64t-unknown-linux-gnu/mpshift
Setup of COSMO-RS:
ctd = BP_TZVP_C30_1601.ctd
cdir = "/software/cluster/COSMOthermX16/COSMOtherm/CTDATA-FILES"
ldir = "/software/cluster/COSMOthermX16/COSMOtherm/CTDATA-FILES"
Using /software/cluster/COSMOthermX16/COSMOtherm/DATABASE-COSMO/BP-TZVP-COSMO
as path to the COSMO-RS NORMAL DATABASE.
Using cefine from /tmp/_MEIfYCvi7/cefine
PARNODES for TM or COSMO-RS calculation was set to 4
Using COSMOtherm from /software/cluster/COSMOthermX16/COSMOtherm/BIN-LINUX/cosmotherm
----------------------------------------------------------------------------------------------------
Processing data from previous run (enso.json)
----------------------------------------------------------------------------------------------------
Reading file: enso.json
Backing up enso.json to enso.json.1.
----------------------------------------------------------------------------------------------------
CRE CHEAP-PRESCREENING - PART0
----------------------------------------------------------------------------------------------------
program: tm
functional for part0: b97-d3(0)
basis set for part0: def2-SV(P)
threshold g_thr(0): 4.0
starting number of considered conformers: 86
temperature: 298.15
Calculating efficient gas-phase single-point energies:
The efficient gas-phase single-point was calculated before for:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF10, CONF11
CONF12, CONF13, CONF14, CONF15, CONF16, CONF17, CONF18, CONF19, CONF20, CONF21, CONF22
CONF23, CONF24, CONF25, CONF26, CONF27, CONF28, CONF29, CONF30, CONF31, CONF32, CONF33
CONF34, CONF35, CONF36, CONF37, CONF38, CONF39, CONF40, CONF41, CONF42, CONF43, CONF44
CONF45, CONF46, CONF47, CONF48, CONF49, CONF50, CONF51, CONF52, CONF53, CONF54, CONF55
CONF56, CONF57, CONF58, CONF59, CONF60, CONF61, CONF62, CONF63, CONF64, CONF65, CONF66
CONF67, CONF68, CONF69, CONF70, CONF71, CONF72, CONF73, CONF74, CONF75, CONF76, CONF77
CONF78, CONF79, CONF80, CONF81, CONF82, CONF83, CONF84, CONF85, CONF86
No conformers are considered additionally.
The efficient gas-phase single-point was successful for CONF1/r2scan-3c: E(DFT) = -310.50644556 Gsolv = -0.01259316
The efficient gas-phase single-point was successful for CONF2/r2scan-3c: E(DFT) = -310.50549173 Gsolv = -0.01254507
The efficient gas-phase single-point was successful for CONF3/r2scan-3c: E(DFT) = -310.50556800 Gsolv = -0.01252321
The efficient gas-phase single-point was successful for CONF4/r2scan-3c: E(DFT) = -310.50452092 Gsolv = -0.01239310
The efficient gas-phase single-point was successful for CONF5/r2scan-3c: E(DFT) = -310.50324027 Gsolv = -0.01290135
The efficient gas-phase single-point was successful for CONF6/r2scan-3c: E(DFT) = -310.50430238 Gsolv = -0.01302933
The efficient gas-phase single-point was successful for CONF7/r2scan-3c: E(DFT) = -310.50362157 Gsolv = -0.01294663
The efficient gas-phase single-point was successful for CONF8/r2scan-3c: E(DFT) = -310.50501659 Gsolv = -0.01244904
The efficient gas-phase single-point was successful for CONF9/r2scan-3c: E(DFT) = -310.50323217 Gsolv = -0.01304501
The efficient gas-phase single-point was successful for CONF10/r2scan-3c: E(DFT) = -310.50316255 Gsolv = -0.01250823
The efficient gas-phase single-point was successful for CONF11/r2scan-3c: E(DFT) = -310.50365564 Gsolv = -0.01301507
The efficient gas-phase single-point was successful for CONF12/r2scan-3c: E(DFT) = -310.50301128 Gsolv = -0.01313443
The efficient gas-phase single-point was successful for CONF13/r2scan-3c: E(DFT) = -310.50235295 Gsolv = -0.01312964
The efficient gas-phase single-point was successful for CONF14/r2scan-3c: E(DFT) = -310.50229552 Gsolv = -0.01303556
The efficient gas-phase single-point was successful for CONF15/r2scan-3c: E(DFT) = -310.50299453 Gsolv = -0.01304084
The efficient gas-phase single-point was successful for CONF16/r2scan-3c: E(DFT) = -310.50201155 Gsolv = -0.01303909
The efficient gas-phase single-point was successful for CONF17/r2scan-3c: E(DFT) = -310.50257043 Gsolv = -0.01304199
The efficient gas-phase single-point was successful for CONF18/r2scan-3c: E(DFT) = -310.50262684 Gsolv = -0.01298126
The efficient gas-phase single-point was successful for CONF19/r2scan-3c: E(DFT) = -310.50238589 Gsolv = -0.01299909
The efficient gas-phase single-point was successful for CONF20/r2scan-3c: E(DFT) = -310.50303283 Gsolv = -0.01271583
The efficient gas-phase single-point was successful for CONF21/r2scan-3c: E(DFT) = -310.50298056 Gsolv = -0.01300713
The efficient gas-phase single-point was successful for CONF22/r2scan-3c: E(DFT) = -310.50141939 Gsolv = -0.01316957
The efficient gas-phase single-point was successful for CONF23/r2scan-3c: E(DFT) = -310.50224518 Gsolv = -0.01302580
The efficient gas-phase single-point was successful for CONF24/r2scan-3c: E(DFT) = -310.50126153 Gsolv = -0.01317505
The efficient gas-phase single-point was successful for CONF25/r2scan-3c: E(DFT) = -310.50149071 Gsolv = -0.01298526
The efficient gas-phase single-point was successful for CONF26/r2scan-3c: E(DFT) = -310.50172224 Gsolv = -0.01284566
The efficient gas-phase single-point was successful for CONF27/r2scan-3c: E(DFT) = -310.50103871 Gsolv = -0.01317593
The efficient gas-phase single-point was successful for CONF28/r2scan-3c: E(DFT) = -310.50188946 Gsolv = -0.01286462
The efficient gas-phase single-point was successful for CONF29/r2scan-3c: E(DFT) = -310.50193595 Gsolv = -0.01285422
The efficient gas-phase single-point was successful for CONF30/r2scan-3c: E(DFT) = -310.50300675 Gsolv = -0.01304005
The efficient gas-phase single-point was successful for CONF31/r2scan-3c: E(DFT) = -310.50296021 Gsolv = -0.01303605
The efficient gas-phase single-point was successful for CONF32/r2scan-3c: E(DFT) = -310.50273943 Gsolv = -0.01296876
The efficient gas-phase single-point was successful for CONF33/r2scan-3c: E(DFT) = -310.50055463 Gsolv = -0.01314045
The efficient gas-phase single-point was successful for CONF34/r2scan-3c: E(DFT) = -310.50017301 Gsolv = -0.01313233
The efficient gas-phase single-point was successful for CONF35/r2scan-3c: E(DFT) = -310.50221173 Gsolv = -0.01269156
The efficient gas-phase single-point was successful for CONF36/r2scan-3c: E(DFT) = -310.50351951 Gsolv = -0.01280423
The efficient gas-phase single-point was successful for CONF37/r2scan-3c: E(DFT) = -310.50129783 Gsolv = -0.01294235
The efficient gas-phase single-point was successful for CONF38/r2scan-3c: E(DFT) = -310.50175782 Gsolv = -0.01310259
The efficient gas-phase single-point was successful for CONF39/r2scan-3c: E(DFT) = -310.50215763 Gsolv = -0.01310764
The efficient gas-phase single-point was successful for CONF40/r2scan-3c: E(DFT) = -310.50165835 Gsolv = -0.01310779
The efficient gas-phase single-point was successful for CONF41/r2scan-3c: E(DFT) = -310.50163520 Gsolv = -0.01303483
The efficient gas-phase single-point was successful for CONF42/r2scan-3c: E(DFT) = -310.50109848 Gsolv = -0.01306423
The efficient gas-phase single-point was successful for CONF43/r2scan-3c: E(DFT) = -310.50319141 Gsolv = -0.01269743
The efficient gas-phase single-point was successful for CONF44/r2scan-3c: E(DFT) = -310.50104233 Gsolv = -0.01307209
The efficient gas-phase single-point was successful for CONF45/r2scan-3c: E(DFT) = -310.50069824 Gsolv = -0.01308548
The efficient gas-phase single-point was successful for CONF46/r2scan-3c: E(DFT) = -310.50002287 Gsolv = -0.01306407
The efficient gas-phase single-point was successful for CONF47/r2scan-3c: E(DFT) = -310.50020045 Gsolv = -0.01306941
The efficient gas-phase single-point was successful for CONF48/r2scan-3c: E(DFT) = -310.50229899 Gsolv = -0.01266393
The efficient gas-phase single-point was successful for CONF49/r2scan-3c: E(DFT) = -310.50216429 Gsolv = -0.01293336
The efficient gas-phase single-point was successful for CONF50/r2scan-3c: E(DFT) = -310.50195524 Gsolv = -0.01293931
The efficient gas-phase single-point was successful for CONF51/r2scan-3c: E(DFT) = -310.50068600 Gsolv = -0.01277049
The efficient gas-phase single-point was successful for CONF52/r2scan-3c: E(DFT) = -310.50083692 Gsolv = -0.01280485
The efficient gas-phase single-point was successful for CONF53/r2scan-3c: E(DFT) = -310.50057125 Gsolv = -0.01285096
The efficient gas-phase single-point was successful for CONF54/r2scan-3c: E(DFT) = -310.50114145 Gsolv = -0.01304802
The efficient gas-phase single-point was successful for CONF55/r2scan-3c: E(DFT) = -310.50096503 Gsolv = -0.01304828
The efficient gas-phase single-point was successful for CONF56/r2scan-3c: E(DFT) = -310.50043866 Gsolv = -0.01305560
The efficient gas-phase single-point was successful for CONF57/r2scan-3c: E(DFT) = -310.50252236 Gsolv = -0.01297975
The efficient gas-phase single-point was successful for CONF58/r2scan-3c: E(DFT) = -310.50518631 Gsolv = -0.01258490
The efficient gas-phase single-point was successful for CONF59/r2scan-3c: E(DFT) = -310.50435689 Gsolv = -0.01261304
The efficient gas-phase single-point was successful for CONF60/r2scan-3c: E(DFT) = -310.50490609 Gsolv = -0.01266506
The efficient gas-phase single-point was successful for CONF61/r2scan-3c: E(DFT) = -310.50435133 Gsolv = -0.01274205
The efficient gas-phase single-point was successful for CONF62/r2scan-3c: E(DFT) = -310.50314978 Gsolv = -0.01279024
The efficient gas-phase single-point was successful for CONF63/r2scan-3c: E(DFT) = -310.50186647 Gsolv = -0.01292045
The efficient gas-phase single-point was successful for CONF64/r2scan-3c: E(DFT) = -310.50307118 Gsolv = -0.01301270
The efficient gas-phase single-point was successful for CONF65/r2scan-3c: E(DFT) = -310.50225800 Gsolv = -0.01319383
The efficient gas-phase single-point was successful for CONF66/r2scan-3c: E(DFT) = -310.50193130 Gsolv = -0.01319355
The efficient gas-phase single-point was successful for CONF67/r2scan-3c: E(DFT) = -310.50228846 Gsolv = -0.01319400
The efficient gas-phase single-point was successful for CONF68/r2scan-3c: E(DFT) = -310.50220287 Gsolv = -0.01320023
The efficient gas-phase single-point was successful for CONF69/r2scan-3c: E(DFT) = -310.50276338 Gsolv = -0.01308969
The efficient gas-phase single-point was successful for CONF70/r2scan-3c: E(DFT) = -310.50136803 Gsolv = -0.01286081
The efficient gas-phase single-point was successful for CONF71/r2scan-3c: E(DFT) = -310.50131530 Gsolv = -0.01287980
The efficient gas-phase single-point was successful for CONF72/r2scan-3c: E(DFT) = -310.50142923 Gsolv = -0.01284848
The efficient gas-phase single-point was successful for CONF73/r2scan-3c: E(DFT) = -310.50036983 Gsolv = -0.01321449
The efficient gas-phase single-point was successful for CONF74/r2scan-3c: E(DFT) = -310.50166270 Gsolv = -0.01308829
The efficient gas-phase single-point was successful for CONF75/r2scan-3c: E(DFT) = -310.50118989 Gsolv = -0.01319950
The efficient gas-phase single-point was successful for CONF76/r2scan-3c: E(DFT) = -310.49985984 Gsolv = -0.01304093
The efficient gas-phase single-point was successful for CONF77/r2scan-3c: E(DFT) = -310.50238832 Gsolv = -0.01278828
The efficient gas-phase single-point was successful for CONF78/r2scan-3c: E(DFT) = -310.50035337 Gsolv = -0.01310207
The efficient gas-phase single-point was successful for CONF79/r2scan-3c: E(DFT) = -310.50039686 Gsolv = -0.01312173
The efficient gas-phase single-point was successful for CONF80/r2scan-3c: E(DFT) = -310.50094664 Gsolv = -0.01294095
The efficient gas-phase single-point was successful for CONF81/r2scan-3c: E(DFT) = -310.49909407 Gsolv = -0.01298452
The efficient gas-phase single-point was successful for CONF82/r2scan-3c: E(DFT) = -310.49903148 Gsolv = -0.01298146
The efficient gas-phase single-point was successful for CONF83/r2scan-3c: E(DFT) = -310.50011988 Gsolv = -0.01306104
The efficient gas-phase single-point was successful for CONF84/r2scan-3c: E(DFT) = -310.50030930 Gsolv = -0.01307692
The efficient gas-phase single-point was successful for CONF85/r2scan-3c: E(DFT) = -310.50026527 Gsolv = -0.01313995
The efficient gas-phase single-point was successful for CONF86/r2scan-3c: E(DFT) = -310.49752615 Gsolv = -0.01292175
----------------------------------------------------------------------------------------------------
Removing high lying conformers by improved energy description
----------------------------------------------------------------------------------------------------
CONF# E [Eh] ΔE [kcal/mol] E [Eh] Gsolv [Eh] gtot ΔE(DFT) ΔGsolv Δgtot
GFN2-xTB GFN2-xTB b97-d3(0)/def2-SV(P) alpb [Eh] [kcal/mol] [kcal/mol] [kcal/mol]
[alpb] [alpb] [gfn2]
CONF1 -23.0107276 0.00 -310.5064456 -0.0125932 -310.5190387 0.00 0.00 0.00 <------
CONF2 -23.0101222 0.38 -310.5054917 -0.0125451 -310.5180368 0.60 0.03 0.63
CONF3 -23.0101206 0.38 -310.5055680 -0.0125232 -310.5180912 0.55 0.04 0.59
CONF4 -23.0098044 0.58 -310.5045209 -0.0123931 -310.5169140 1.21 0.13 1.33
CONF5 -23.0097569 0.61 -310.5032403 -0.0129013 -310.5161416 2.01 -0.19 1.82
CONF6 -23.0096876 0.65 -310.5043024 -0.0130293 -310.5173317 1.34 -0.27 1.07
CONF7 -23.0096343 0.69 -310.5036216 -0.0129466 -310.5165682 1.77 -0.22 1.55
CONF8 -23.0096174 0.70 -310.5050166 -0.0124490 -310.5174656 0.90 0.09 0.99
CONF9 -23.0095115 0.76 -310.5032322 -0.0130450 -310.5162772 2.02 -0.28 1.73
CONF10 -23.0094779 0.78 -310.5031625 -0.0125082 -310.5156708 2.06 0.05 2.11
CONF11 -23.0094724 0.79 -310.5036556 -0.0130151 -310.5166707 1.75 -0.26 1.49
CONF12 -23.0094402 0.81 -310.5030113 -0.0131344 -310.5161457 2.16 -0.34 1.82
CONF13 -23.0094039 0.83 -310.5023530 -0.0131296 -310.5154826 2.57 -0.34 2.23
CONF14 -23.0092672 0.92 -310.5022955 -0.0130356 -310.5153311 2.60 -0.28 2.33
CONF15 -23.0092160 0.95 -310.5029945 -0.0130408 -310.5160354 2.17 -0.28 1.88
CONF16 -23.0092080 0.95 -310.5020116 -0.0130391 -310.5150506 2.78 -0.28 2.50
CONF17 -23.0091923 0.96 -310.5025704 -0.0130420 -310.5156124 2.43 -0.28 2.15
CONF18 -23.0091398 1.00 -310.5026268 -0.0129813 -310.5156081 2.40 -0.24 2.15
CONF19 -23.0091297 1.00 -310.5023859 -0.0129991 -310.5153850 2.55 -0.25 2.29
CONF20 -23.0090825 1.03 -310.5030328 -0.0127158 -310.5157487 2.14 -0.08 2.06
CONF21 -23.0090389 1.06 -310.5029806 -0.0130071 -310.5159877 2.17 -0.26 1.91
CONF22 -23.0090378 1.06 -310.5014194 -0.0131696 -310.5145890 3.15 -0.36 2.79
CONF23 -23.0090092 1.08 -310.5022452 -0.0130258 -310.5152710 2.64 -0.27 2.36
CONF24 -23.0089916 1.09 -310.5012615 -0.0131750 -310.5144366 3.25 -0.37 2.89
CONF25 -23.0089768 1.10 -310.5014907 -0.0129853 -310.5144760 3.11 -0.25 2.86
CONF26 -23.0090337 1.06 -310.5017222 -0.0128457 -310.5145679 2.96 -0.16 2.81
CONF27 -23.0089880 1.09 -310.5010387 -0.0131759 -310.5142146 3.39 -0.37 3.03
CONF28 -23.0088839 1.16 -310.5018895 -0.0128646 -310.5147541 2.86 -0.17 2.69
CONF29 -23.0088827 1.16 -310.5019360 -0.0128542 -310.5147902 2.83 -0.16 2.67
CONF30 -23.0088560 1.17 -310.5030068 -0.0130400 -310.5160468 2.16 -0.28 1.88
CONF31 -23.0088493 1.18 -310.5029602 -0.0130361 -310.5159963 2.19 -0.28 1.91
CONF32 -23.0088201 1.20 -310.5027394 -0.0129688 -310.5157082 2.33 -0.24 2.09
CONF33 -23.0087944 1.21 -310.5005546 -0.0131404 -310.5136951 3.70 -0.34 3.35
CONF34 -23.0087841 1.22 -310.5001730 -0.0131323 -310.5133053 3.94 -0.34 3.60
CONF35 -23.0087018 1.27 -310.5022117 -0.0126916 -310.5149033 2.66 -0.06 2.60
CONF36 -23.0085315 1.38 -310.5035195 -0.0128042 -310.5163237 1.84 -0.13 1.70
CONF37 -23.0084271 1.44 -310.5012978 -0.0129424 -310.5142402 3.23 -0.22 3.01
CONF38 -23.0083912 1.47 -310.5017578 -0.0131026 -310.5148604 2.94 -0.32 2.62
CONF39 -23.0083734 1.48 -310.5021576 -0.0131076 -310.5152653 2.69 -0.32 2.37
CONF40 -23.0083684 1.48 -310.5016583 -0.0131078 -310.5147661 3.00 -0.32 2.68
CONF41 -23.0083492 1.49 -310.5016352 -0.0130348 -310.5146700 3.02 -0.28 2.74
CONF42 -23.0083209 1.51 -310.5010985 -0.0130642 -310.5141627 3.36 -0.30 3.06
CONF43 -23.0082969 1.53 -310.5031914 -0.0126974 -310.5158888 2.04 -0.07 1.98
CONF44 -23.0082292 1.57 -310.5010423 -0.0130721 -310.5141144 3.39 -0.30 3.09
CONF45 -23.0082256 1.57 -310.5006982 -0.0130855 -310.5137837 3.61 -0.31 3.30
CONF46 -23.0082146 1.58 -310.5000229 -0.0130641 -310.5130869 4.03 -0.30 3.73
CONF47 -23.0082062 1.58 -310.5002005 -0.0130694 -310.5132699 3.92 -0.30 3.62
CONF48 -23.0081931 1.59 -310.5022990 -0.0126639 -310.5149629 2.60 -0.04 2.56
CONF49 -23.0081343 1.63 -310.5021643 -0.0129334 -310.5150976 2.69 -0.21 2.47
CONF50 -23.0080982 1.65 -310.5019552 -0.0129393 -310.5148946 2.82 -0.22 2.60
CONF51 -23.0080894 1.66 -310.5006860 -0.0127705 -310.5134565 3.61 -0.11 3.50
CONF52 -23.0080887 1.66 -310.5008369 -0.0128049 -310.5136418 3.52 -0.13 3.39
CONF53 -23.0080770 1.66 -310.5005713 -0.0128510 -310.5134222 3.69 -0.16 3.52
CONF54 -23.0080314 1.69 -310.5011414 -0.0130480 -310.5141895 3.33 -0.29 3.04
CONF55 -23.0080039 1.71 -310.5009650 -0.0130483 -310.5140133 3.44 -0.29 3.15
CONF56 -23.0077011 1.90 -310.5004387 -0.0130556 -310.5134943 3.77 -0.29 3.48
CONF57 -23.0076712 1.92 -310.5025224 -0.0129797 -310.5155021 2.46 -0.24 2.22
CONF58 -23.0075551 1.99 -310.5051863 -0.0125849 -310.5177712 0.79 0.01 0.80
CONF59 -23.0075531 1.99 -310.5043569 -0.0126130 -310.5169699 1.31 -0.01 1.30
CONF60 -23.0075409 2.00 -310.5049061 -0.0126651 -310.5175712 0.97 -0.05 0.92
CONF61 -23.0075072 2.02 -310.5043513 -0.0127421 -310.5170934 1.31 -0.09 1.22
CONF62 -23.0074774 2.04 -310.5031498 -0.0127902 -310.5159400 2.07 -0.12 1.94
CONF63 -23.0073284 2.13 -310.5018665 -0.0129205 -310.5147869 2.87 -0.21 2.67
CONF64 -23.0071119 2.27 -310.5030712 -0.0130127 -310.5160839 2.12 -0.26 1.85
CONF65 -23.0071098 2.27 -310.5022580 -0.0131938 -310.5154518 2.63 -0.38 2.25
CONF66 -23.0071072 2.27 -310.5019313 -0.0131935 -310.5151248 2.83 -0.38 2.46
CONF67 -23.0071096 2.27 -310.5022885 -0.0131940 -310.5154825 2.61 -0.38 2.23
CONF68 -23.0071027 2.27 -310.5022029 -0.0132002 -310.5154031 2.66 -0.38 2.28
CONF69 -23.0069880 2.35 -310.5027634 -0.0130897 -310.5158531 2.31 -0.31 2.00
CONF70 -23.0068383 2.44 -310.5013680 -0.0128608 -310.5142288 3.19 -0.17 3.02
CONF71 -23.0068349 2.44 -310.5013153 -0.0128798 -310.5141951 3.22 -0.18 3.04
CONF72 -23.0068337 2.44 -310.5014292 -0.0128485 -310.5142777 3.15 -0.16 2.99
CONF73 -23.0067855 2.47 -310.5003698 -0.0132145 -310.5135843 3.81 -0.39 3.42
CONF74 -23.0066627 2.55 -310.5016627 -0.0130883 -310.5147510 3.00 -0.31 2.69
CONF75 -23.0066210 2.58 -310.5011899 -0.0131995 -310.5143894 3.30 -0.38 2.92
CONF76 -23.0063840 2.73 -310.4998598 -0.0130409 -310.5129008 4.13 -0.28 3.85
CONF77 -23.0061505 2.87 -310.5023883 -0.0127883 -310.5151766 2.55 -0.12 2.42
CONF78 -23.0060644 2.93 -310.5003534 -0.0131021 -310.5134554 3.82 -0.32 3.50
CONF79 -23.0060556 2.93 -310.5003969 -0.0131217 -310.5135186 3.80 -0.33 3.46
CONF80 -23.0056253 3.20 -310.5009466 -0.0129409 -310.5138876 3.45 -0.22 3.23
CONF81 -23.0053013 3.41 -310.4990941 -0.0129845 -310.5120786 4.61 -0.25 4.37
CONF82 -23.0052859 3.41 -310.4990315 -0.0129815 -310.5120129 4.65 -0.24 4.41
CONF83 -23.0051513 3.50 -310.5001199 -0.0130610 -310.5131809 3.97 -0.29 3.68
CONF84 -23.0051460 3.50 -310.5003093 -0.0130769 -310.5133862 3.85 -0.30 3.55
CONF85 -23.0048789 3.67 -310.5002653 -0.0131399 -310.5134052 3.88 -0.34 3.54
CONF86 -23.0029504 4.88 -310.4975262 -0.0129217 -310.5104479 5.60 -0.21 5.39
----------------------------------------------------------------------------------------------------
Number of conformers observed within the following Δg windows:
Δg [kcal/mol] #CONF sum(Boltzmann_weights)
---------------------------------------------
0 - 0.5 1 0.24
0 - 1.0 6 0.56
0 - 1.5 11 0.70
0 - 2.0 24 0.85
0 - 2.5 41 0.94
0 - 3.0 58 0.98
0 - 3.5 73 0.99
0 - 4.0 83 1.00
0 - 4.5 85 1.00
0 - 5.0 85 1.00
0 - 5.5 86 1.00
---------------------------------------------
--------------------------------------------------
Conformers considered further
--------------------------------------------------
These conformers are below the 4.000 kcal/mol g_thr(0) threshold.
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF10, CONF11
CONF12, CONF13, CONF14, CONF15, CONF16, CONF17, CONF18, CONF19, CONF20, CONF21, CONF22
CONF23, CONF24, CONF25, CONF26, CONF27, CONF28, CONF29, CONF30, CONF31, CONF32, CONF33
CONF34, CONF35, CONF36, CONF37, CONF38, CONF39, CONF40, CONF41, CONF42, CONF43, CONF44
CONF45, CONF46, CONF47, CONF48, CONF49, CONF50, CONF51, CONF52, CONF53, CONF54, CONF55
CONF56, CONF57, CONF58, CONF59, CONF60, CONF61, CONF62, CONF63, CONF64, CONF65, CONF66
CONF67, CONF68, CONF69, CONF70, CONF71, CONF72, CONF73, CONF74, CONF75, CONF76, CONF77
CONF78, CONF79, CONF80, CONF83, CONF84, CONF85
Calculating Boltzmann averaged (free) energy of ensemble on input geometries (not DFT optimized)!
temperature /K: avE(T) /a.u. avG(T) /a.u.
----------------------------------------------------------------------------------------------------
298.15 -310.5046287 -310.5173461 <<==part0==
----------------------------------------------------------------------------------------------------
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>END of Part0<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
Ran part0 in 0.1865 seconds
----------------------------------------------------------------------------------------------------
CRE PRESCREENING - PART1
----------------------------------------------------------------------------------------------------
program: tm
functional for part1 and 2: r2scan-3c
basis set for part1 and 2: def2-mTZVPP
Solvent: chcl3
solvent model for Gsolv contribution: cosmors
threshold g_thr(1) and G_thr(1): 3.5
starting number of considered conformers: 83
calculate mRRHO contribution: on
program for mRRHO contribution: xtb
GFN version for mRRHO and/or GBSA_Gsolv: gfn2
Apply constraint to input geometry during mRRHO calculation: on
temperature: 298.15
Calculating single-point energies and solvation contribution (G_solv):
The prescreening_single-point was calculated before for:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF10, CONF11
CONF12, CONF13, CONF14, CONF15, CONF16, CONF17, CONF18, CONF19, CONF20, CONF21, CONF22
CONF23, CONF24, CONF25, CONF26, CONF27, CONF28, CONF29, CONF30, CONF31, CONF32, CONF33
CONF34, CONF35, CONF36, CONF37, CONF38, CONF39, CONF40, CONF41, CONF42, CONF43, CONF44
CONF45, CONF46, CONF47, CONF48, CONF49, CONF50, CONF51, CONF52, CONF53, CONF54, CONF55
CONF56, CONF57, CONF58, CONF59, CONF60, CONF61, CONF62, CONF63, CONF64, CONF65, CONF66
CONF67, CONF68, CONF69, CONF70, CONF71, CONF72, CONF73, CONF74, CONF75, CONF76, CONF77
CONF78, CONF79, CONF80, CONF83, CONF84, CONF85
No conformers are considered additionally.
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF1/r2scan-3c: -0.00749496
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF2/r2scan-3c: -0.00744587
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF3/r2scan-3c: -0.00741805
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF4/r2scan-3c: -0.00644510
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF5/r2scan-3c: -0.00793157
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF6/r2scan-3c: -0.00732222
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF7/r2scan-3c: -0.00798114
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF8/r2scan-3c: -0.00701204
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF9/r2scan-3c: -0.00773387
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF10/r2scan-3c: -0.00745434
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF11/r2scan-3c: -0.00769430
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF12/r2scan-3c: -0.00806872
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF13/r2scan-3c: -0.00813639
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF14/r2scan-3c: -0.00767170
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF15/r2scan-3c: -0.00771300
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF16/r2scan-3c: -0.00770135
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF17/r2scan-3c: -0.00770460
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF18/r2scan-3c: -0.00797719
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF19/r2scan-3c: -0.00797990
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF20/r2scan-3c: -0.00694890
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF21/r2scan-3c: -0.00730959
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF22/r2scan-3c: -0.00811097
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF23/r2scan-3c: -0.00773655
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF24/r2scan-3c: -0.00816990
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF25/r2scan-3c: -0.00766066
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF26/r2scan-3c: -0.00787301
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF27/r2scan-3c: -0.00815090
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF28/r2scan-3c: -0.00803486
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF29/r2scan-3c: -0.00802964
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF30/r2scan-3c: -0.00785297
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF31/r2scan-3c: -0.00768147
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF32/r2scan-3c: -0.00770491
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF33/r2scan-3c: -0.00809670
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF34/r2scan-3c: -0.00812084
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF35/r2scan-3c: -0.00744830
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF36/r2scan-3c: -0.00729869
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF37/r2scan-3c: -0.00732253
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF38/r2scan-3c: -0.00796110
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF39/r2scan-3c: -0.00800842
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF40/r2scan-3c: -0.00800695
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF41/r2scan-3c: -0.00785621
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF42/r2scan-3c: -0.00777196
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF43/r2scan-3c: -0.00686110
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF44/r2scan-3c: -0.00796971
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF45/r2scan-3c: -0.00800838
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF46/r2scan-3c: -0.00791748
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF47/r2scan-3c: -0.00793228
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF48/r2scan-3c: -0.00754897
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF49/r2scan-3c: -0.00759862
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF50/r2scan-3c: -0.00771436
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF51/r2scan-3c: -0.00720396
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF52/r2scan-3c: -0.00724495
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF53/r2scan-3c: -0.00740309
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF54/r2scan-3c: -0.00813890
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF55/r2scan-3c: -0.00803376
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF56/r2scan-3c: -0.00802618
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF57/r2scan-3c: -0.00787894
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF58/r2scan-3c: -0.00656993
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF59/r2scan-3c: -0.00660332
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF60/r2scan-3c: -0.00665323
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF61/r2scan-3c: -0.00672072
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF62/r2scan-3c: -0.00679624
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF63/r2scan-3c: -0.00774301
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF64/r2scan-3c: -0.00727539
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF65/r2scan-3c: -0.00823067
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF66/r2scan-3c: -0.00822543
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF67/r2scan-3c: -0.00821952
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF68/r2scan-3c: -0.00824090
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF69/r2scan-3c: -0.00760791
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF70/r2scan-3c: -0.00792993
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF71/r2scan-3c: -0.00797498
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF72/r2scan-3c: -0.00791216
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF73/r2scan-3c: -0.00820864
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF74/r2scan-3c: -0.00772638
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF75/r2scan-3c: -0.00820207
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF76/r2scan-3c: -0.00765267
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF77/r2scan-3c: -0.00716368
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF78/r2scan-3c: -0.00700951
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF79/r2scan-3c: -0.00707422
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF80/r2scan-3c: -0.00754296
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF83/r2scan-3c: -0.00750673
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF84/r2scan-3c: -0.00754007
prescreening COSMO-RS calculation was successful for 3870734.majestix.thch.uni-bonn.de/CONF85/r2scan-3c: -0.00759296
--------------------------------------------------
Removing high lying conformers
--------------------------------------------------
CONF# E(GFNn-xTB) ΔE(GFNn-xTB) E [Eh] Gsolv [Eh] gtot Δgtot
[a.u.] [kcal/mol] r2scan-3c COSMO-RS-normal [Eh] [kcal/mol]
[r2scan-3c]
CONF1 -23.0093455 0.00 -310.9770910 -0.0074950 -310.9845859 0.00 <------
CONF2 -23.0087399 0.38 -310.9757758 -0.0074459 -310.9832217 0.86
CONF3 -23.0087382 0.38 -310.9758415 -0.0074181 -310.9832596 0.83
CONF4 -23.0084220 0.58 -310.9758145 -0.0064451 -310.9822596 1.46
CONF5 -23.0083752 0.61 -310.9743875 -0.0079316 -310.9823191 1.42
CONF6 -23.0083052 0.65 -310.9745964 -0.0073222 -310.9819186 1.67
CONF7 -23.0082519 0.69 -310.9744677 -0.0079811 -310.9824488 1.34
CONF8 -23.0082350 0.70 -310.9757295 -0.0070120 -310.9827415 1.16
CONF9 -23.0081291 0.76 -310.9745570 -0.0077339 -310.9822909 1.44
CONF10 -23.0080955 0.78 -310.9743226 -0.0074543 -310.9817770 1.76
CONF11 -23.0080901 0.79 -310.9744861 -0.0076943 -310.9821804 1.51
CONF12 -23.0080578 0.81 -310.9741780 -0.0080687 -310.9822467 1.47
CONF13 -23.0080214 0.83 -310.9741538 -0.0081364 -310.9822902 1.44
CONF14 -23.0078848 0.92 -310.9733007 -0.0076717 -310.9809724 2.27
CONF15 -23.0078336 0.95 -310.9735503 -0.0077130 -310.9812633 2.09
CONF16 -23.0078261 0.95 -310.9731490 -0.0077013 -310.9808504 2.34
CONF17 -23.0078099 0.96 -310.9733525 -0.0077046 -310.9810571 2.21
CONF18 -23.0077574 1.00 -310.9730612 -0.0079772 -310.9810384 2.23
CONF19 -23.0077473 1.00 -310.9730253 -0.0079799 -310.9810052 2.25
CONF20 -23.0077001 1.03 -310.9743520 -0.0069489 -310.9813009 2.06
CONF21 -23.0076566 1.06 -310.9732578 -0.0073096 -310.9805674 2.52
CONF22 -23.0076554 1.06 -310.9729525 -0.0081110 -310.9810635 2.21
CONF23 -23.0076269 1.08 -310.9732776 -0.0077366 -310.9810142 2.24
CONF24 -23.0076092 1.09 -310.9728437 -0.0081699 -310.9810136 2.24
CONF25 -23.0075945 1.10 -310.9722243 -0.0076607 -310.9798849 2.95
CONF26 -23.0076514 1.06 -310.9734086 -0.0078730 -310.9812816 2.07
CONF27 -23.0076056 1.09 -310.9727742 -0.0081509 -310.9809251 2.30
CONF28 -23.0075015 1.16 -310.9733748 -0.0080349 -310.9814096 1.99
CONF29 -23.0075003 1.16 -310.9733671 -0.0080296 -310.9813968 2.00
CONF30 -23.0074736 1.17 -310.9740731 -0.0078530 -310.9819261 1.67
CONF31 -23.0074670 1.18 -310.9741187 -0.0076815 -310.9818002 1.75
CONF32 -23.0074378 1.20 -310.9741085 -0.0077049 -310.9818134 1.74
CONF33 -23.0074120 1.21 -310.9721512 -0.0080967 -310.9802479 2.72
CONF34 -23.0074018 1.22 -310.9718597 -0.0081208 -310.9799806 2.89
CONF35 -23.0073194 1.27 -310.9733092 -0.0074483 -310.9807575 2.40
CONF36 -23.0071492 1.38 -310.9738901 -0.0072987 -310.9811887 2.13
CONF37 -23.0070448 1.44 -310.9711644 -0.0073225 -310.9784869 3.83
CONF38 -23.0070094 1.47 -310.9726276 -0.0079611 -310.9805887 2.51
CONF39 -23.0069911 1.48 -310.9726003 -0.0080084 -310.9806087 2.50
CONF40 -23.0069860 1.48 -310.9726416 -0.0080069 -310.9806486 2.47
CONF41 -23.0069668 1.49 -310.9727955 -0.0078562 -310.9806517 2.47
CONF42 -23.0069385 1.51 -310.9728220 -0.0077720 -310.9805939 2.51
CONF43 -23.0069145 1.53 -310.9728106 -0.0068611 -310.9796717 3.08
CONF44 -23.0068468 1.57 -310.9717566 -0.0079697 -310.9797263 3.05
CONF45 -23.0068432 1.57 -310.9718972 -0.0080084 -310.9799056 2.94
CONF46 -23.0068324 1.58 -310.9717215 -0.0079175 -310.9796390 3.10
CONF47 -23.0068240 1.58 -310.9718636 -0.0079323 -310.9797959 3.01
CONF48 -23.0068107 1.59 -310.9734980 -0.0075490 -310.9810470 2.22
CONF49 -23.0067519 1.63 -310.9729840 -0.0075986 -310.9805826 2.51
CONF50 -23.0067158 1.65 -310.9727818 -0.0077144 -310.9804962 2.57
CONF51 -23.0067070 1.66 -310.9705106 -0.0072040 -310.9777146 4.31
CONF52 -23.0067063 1.66 -310.9705431 -0.0072449 -310.9777881 4.27
CONF53 -23.0066946 1.66 -310.9703772 -0.0074031 -310.9777803 4.27
CONF54 -23.0066490 1.69 -310.9722719 -0.0081389 -310.9804108 2.62
CONF55 -23.0066215 1.71 -310.9720817 -0.0080338 -310.9801155 2.81
CONF56 -23.0063187 1.90 -310.9713906 -0.0080262 -310.9794168 3.24
CONF57 -23.0062889 1.92 -310.9744054 -0.0078789 -310.9822844 1.44
CONF58 -23.0061728 1.99 -310.9760271 -0.0065699 -310.9825970 1.25
CONF59 -23.0061707 1.99 -310.9759465 -0.0066033 -310.9825498 1.28
CONF60 -23.0061585 2.00 -310.9758398 -0.0066532 -310.9824931 1.31
CONF61 -23.0061249 2.02 -310.9755805 -0.0067207 -310.9823012 1.43
CONF62 -23.0060951 2.04 -310.9754193 -0.0067962 -310.9822155 1.49
CONF63 -23.0059460 2.13 -310.9722754 -0.0077430 -310.9800184 2.87
CONF64 -23.0057295 2.27 -310.9745205 -0.0072754 -310.9817959 1.75
CONF65 -23.0057274 2.27 -310.9738656 -0.0082307 -310.9820962 1.56
CONF66 -23.0057248 2.27 -310.9738666 -0.0082254 -310.9820920 1.56
CONF67 -23.0057272 2.27 -310.9738672 -0.0082195 -310.9820867 1.57
CONF68 -23.0057203 2.27 -310.9738563 -0.0082409 -310.9820972 1.56
CONF69 -23.0056054 2.35 -310.9744729 -0.0076079 -310.9820808 1.57
CONF70 -23.0054560 2.44 -310.9732794 -0.0079299 -310.9812093 2.12
CONF71 -23.0054525 2.44 -310.9732799 -0.0079750 -310.9812549 2.09
CONF72 -23.0054513 2.44 -310.9732609 -0.0079122 -310.9811731 2.14
CONF73 -23.0054031 2.47 -310.9725927 -0.0082086 -310.9808013 2.37
CONF74 -23.0052804 2.55 -310.9729596 -0.0077264 -310.9806859 2.45
CONF75 -23.0052386 2.58 -310.9726279 -0.0082021 -310.9808299 2.36
CONF76 -23.0050016 2.73 -310.9724113 -0.0076527 -310.9800640 2.84
CONF77 -23.0047687 2.87 -310.9739372 -0.0071637 -310.9811008 2.19
CONF78 -23.0046820 2.93 -310.9710324 -0.0070095 -310.9780419 4.11
CONF79 -23.0046732 2.93 -310.9710915 -0.0070742 -310.9781657 4.03
CONF80 -23.0042429 3.20 -310.9726319 -0.0075430 -310.9801748 2.77
CONF83 -23.0037689 3.50 -310.9703973 -0.0075067 -310.9779040 4.19
CONF84 -23.0037636 3.50 -310.9705035 -0.0075401 -310.9780436 4.11
CONF85 -23.0034965 3.67 -310.9711983 -0.0075930 -310.9787913 3.64
--------------------------------------------------
Conformers considered further
--------------------------------------------------
Below the g_thr(1) threshold of 3.5 kcal/mol.
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF10, CONF11
CONF12, CONF13, CONF14, CONF15, CONF16, CONF17, CONF18, CONF19, CONF20, CONF21, CONF22
CONF23, CONF24, CONF25, CONF26, CONF27, CONF28, CONF29, CONF30, CONF31, CONF32, CONF33
CONF34, CONF35, CONF36, CONF38, CONF39, CONF40, CONF41, CONF42, CONF43, CONF44, CONF45
CONF46, CONF47, CONF48, CONF49, CONF50, CONF54, CONF55, CONF56, CONF57, CONF58, CONF59
CONF60, CONF61, CONF62, CONF63, CONF64, CONF65, CONF66, CONF67, CONF68, CONF69, CONF70
CONF71, CONF72, CONF73, CONF74, CONF75, CONF76, CONF77, CONF80
--------------------------------------------------
Calculating prescreening G_mRRHO with implicit solvation!
The prescreening G_mRRHO calculation was performed before for:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF10, CONF11
CONF12, CONF13, CONF14, CONF15, CONF16, CONF17, CONF18, CONF19, CONF20, CONF21, CONF22
CONF23, CONF24, CONF25, CONF26, CONF27, CONF28, CONF29, CONF30, CONF31, CONF32, CONF33
CONF34, CONF35, CONF36, CONF38, CONF39, CONF40, CONF41, CONF42, CONF43, CONF44, CONF45
CONF46, CONF47, CONF48, CONF49, CONF50, CONF54, CONF55, CONF56, CONF57, CONF58, CONF59
CONF60, CONF61, CONF62, CONF63, CONF64, CONF65, CONF66, CONF67, CONF68, CONF69, CONF70
CONF71, CONF72, CONF73, CONF74, CONF75, CONF76, CONF77, CONF80
No conformers are considered additionally.
The prescreening G_mRRHO run @ c1 was successful for CONF1/rrho_part1: 0.13266847 S_rot(sym)= 0.0000000 using= 0.1326685
The prescreening G_mRRHO run @ c1 was successful for CONF2/rrho_part1: 0.13236997 S_rot(sym)= 0.0000000 using= 0.1323700
The prescreening G_mRRHO run @ c1 was successful for CONF3/rrho_part1: 0.13238335 S_rot(sym)= 0.0000000 using= 0.1323833
The prescreening G_mRRHO run @ c1 was successful for CONF4/rrho_part1: 0.13273993 S_rot(sym)= 0.0000000 using= 0.1327399
The prescreening G_mRRHO run @ c1 was successful for CONF5/rrho_part1: 0.13207173 S_rot(sym)= 0.0000000 using= 0.1320717
The prescreening G_mRRHO run @ c1 was successful for CONF6/rrho_part1: 0.13227151 S_rot(sym)= 0.0000000 using= 0.1322715
The prescreening G_mRRHO run @ c1 was successful for CONF7/rrho_part1: 0.13203902 S_rot(sym)= 0.0000000 using= 0.1320390
The prescreening G_mRRHO run @ c1 was successful for CONF8/rrho_part1: 0.13298555 S_rot(sym)= 0.0000000 using= 0.1329856
The prescreening G_mRRHO run @ c1 was successful for CONF9/rrho_part1: 0.13208817 S_rot(sym)= 0.0000000 using= 0.1320882
The prescreening G_mRRHO run @ c1 was successful for CONF10/rrho_part1: 0.13305109 S_rot(sym)= 0.0000000 using= 0.1330511
The prescreening G_mRRHO run @ c1 was successful for CONF11/rrho_part1: 0.13208957 S_rot(sym)= 0.0000000 using= 0.1320896
The prescreening G_mRRHO run @ c1 was successful for CONF12/rrho_part1: 0.13185775 S_rot(sym)= 0.0000000 using= 0.1318577
The prescreening G_mRRHO run @ c1 was successful for CONF13/rrho_part1: 0.13172814 S_rot(sym)= 0.0000000 using= 0.1317281
The prescreening G_mRRHO run @ c1 was successful for CONF14/rrho_part1: 0.13133244 S_rot(sym)= 0.0000000 using= 0.1313324
The prescreening G_mRRHO run @ c1 was successful for CONF15/rrho_part1: 0.13158015 S_rot(sym)= 0.0000000 using= 0.1315801
The prescreening G_mRRHO run @ c1 was successful for CONF16/rrho_part1: 0.13102712 S_rot(sym)= 0.0000000 using= 0.1310271
The prescreening G_mRRHO run @ c1 was successful for CONF17/rrho_part1: 0.13152957 S_rot(sym)= 0.0000000 using= 0.1315296
The prescreening G_mRRHO run @ c1 was successful for CONF18/rrho_part1: 0.13181881 S_rot(sym)= 0.0000000 using= 0.1318188
The prescreening G_mRRHO run @ c1 was successful for CONF19/rrho_part1: 0.13179938 S_rot(sym)= 0.0000000 using= 0.1317994
The prescreening G_mRRHO run @ c1 was successful for CONF20/rrho_part1: 0.13218213 S_rot(sym)= 0.0000000 using= 0.1321821
The prescreening G_mRRHO run @ c1 was successful for CONF21/rrho_part1: 0.13180554 S_rot(sym)= 0.0000000 using= 0.1318055
The prescreening G_mRRHO run @ c1 was successful for CONF22/rrho_part1: 0.13131455 S_rot(sym)= 0.0000000 using= 0.1313146
The prescreening G_mRRHO run @ c1 was successful for CONF23/rrho_part1: 0.13173778 S_rot(sym)= 0.0000000 using= 0.1317378
The prescreening G_mRRHO run @ c1 was successful for CONF24/rrho_part1: 0.13123951 S_rot(sym)= 0.0000000 using= 0.1312395
The prescreening G_mRRHO run @ c1 was successful for CONF25/rrho_part1: 0.13121544 S_rot(sym)= 0.0000000 using= 0.1312154
The prescreening G_mRRHO run @ c1 was successful for CONF26/rrho_part1: 0.13255550 S_rot(sym)= 0.0000000 using= 0.1325555
The prescreening G_mRRHO run @ c1 was successful for CONF27/rrho_part1: 0.13109064 S_rot(sym)= 0.0000000 using= 0.1310906
The prescreening G_mRRHO run @ c1 was successful for CONF28/rrho_part1: 0.13244816 S_rot(sym)= 0.0000000 using= 0.1324482
The prescreening G_mRRHO run @ c1 was successful for CONF29/rrho_part1: 0.13249911 S_rot(sym)= 0.0000000 using= 0.1324991
The prescreening G_mRRHO run @ c1 was successful for CONF30/rrho_part1: 0.13195200 S_rot(sym)= 0.0000000 using= 0.1319520
The prescreening G_mRRHO run @ c1 was successful for CONF31/rrho_part1: 0.13188626 S_rot(sym)= 0.0000000 using= 0.1318863
The prescreening G_mRRHO run @ c1 was successful for CONF32/rrho_part1: 0.13189156 S_rot(sym)= 0.0000000 using= 0.1318916
The prescreening G_mRRHO run @ c1 was successful for CONF33/rrho_part1: 0.13068033 S_rot(sym)= 0.0000000 using= 0.1306803
The prescreening G_mRRHO run @ c1 was successful for CONF34/rrho_part1: 0.13035059 S_rot(sym)= 0.0000000 using= 0.1303506
The prescreening G_mRRHO run @ c1 was successful for CONF35/rrho_part1: 0.13212904 S_rot(sym)= 0.0000000 using= 0.1321290
The prescreening G_mRRHO run @ c1 was successful for CONF36/rrho_part1: 0.13266578 S_rot(sym)= 0.0000000 using= 0.1326658
The prescreening G_mRRHO run @ c1 was successful for CONF38/rrho_part1: 0.13097664 S_rot(sym)= 0.0000000 using= 0.1309766
The prescreening G_mRRHO run @ c1 was successful for CONF39/rrho_part1: 0.13093130 S_rot(sym)= 0.0000000 using= 0.1309313
The prescreening G_mRRHO run @ c1 was successful for CONF40/rrho_part1: 0.13122016 S_rot(sym)= 0.0000000 using= 0.1312202
The prescreening G_mRRHO run @ c1 was successful for CONF41/rrho_part1: 0.13168292 S_rot(sym)= 0.0000000 using= 0.1316829
The prescreening G_mRRHO run @ c1 was successful for CONF42/rrho_part1: 0.13140104 S_rot(sym)= 0.0000000 using= 0.1314010
The prescreening G_mRRHO run @ c1 was successful for CONF43/rrho_part1: 0.13265033 S_rot(sym)= 0.0000000 using= 0.1326503
The prescreening G_mRRHO run @ c1 was successful for CONF44/rrho_part1: 0.13077871 S_rot(sym)= 0.0000000 using= 0.1307787
The prescreening G_mRRHO run @ c1 was successful for CONF45/rrho_part1: 0.13104663 S_rot(sym)= 0.0000000 using= 0.1310466
The prescreening G_mRRHO run @ c1 was successful for CONF46/rrho_part1: 0.13046405 S_rot(sym)= 0.0000000 using= 0.1304641
The prescreening G_mRRHO run @ c1 was successful for CONF47/rrho_part1: 0.13096796 S_rot(sym)= 0.0000000 using= 0.1309680
The prescreening G_mRRHO run @ c1 was successful for CONF48/rrho_part1: 0.13272039 S_rot(sym)= 0.0000000 using= 0.1327204
The prescreening G_mRRHO run @ c1 was successful for CONF49/rrho_part1: 0.13197129 S_rot(sym)= 0.0000000 using= 0.1319713
The prescreening G_mRRHO run @ c1 was successful for CONF50/rrho_part1: 0.13190007 S_rot(sym)= 0.0000000 using= 0.1319001
The prescreening G_mRRHO run @ c1 was successful for CONF54/rrho_part1: 0.13174946 S_rot(sym)= 0.0000000 using= 0.1317495
The prescreening G_mRRHO run @ c1 was successful for CONF55/rrho_part1: 0.13173176 S_rot(sym)= 0.0000000 using= 0.1317318
The prescreening G_mRRHO run @ c1 was successful for CONF56/rrho_part1: 0.13172318 S_rot(sym)= 0.0000000 using= 0.1317232
The prescreening G_mRRHO run @ c1 was successful for CONF57/rrho_part1: 0.13179903 S_rot(sym)= 0.0000000 using= 0.1317990
The prescreening G_mRRHO run @ c1 was successful for CONF58/rrho_part1: 0.13230588 S_rot(sym)= 0.0000000 using= 0.1323059
The prescreening G_mRRHO run @ c1 was successful for CONF59/rrho_part1: 0.13223264 S_rot(sym)= 0.0000000 using= 0.1322326
The prescreening G_mRRHO run @ c1 was successful for CONF60/rrho_part1: 0.13199094 S_rot(sym)= 0.0000000 using= 0.1319909
The prescreening G_mRRHO run @ c1 was successful for CONF61/rrho_part1: 0.13168460 S_rot(sym)= 0.0000000 using= 0.1316846
The prescreening G_mRRHO run @ c1 was successful for CONF62/rrho_part1: 0.13187907 S_rot(sym)= 0.0000000 using= 0.1318791
The prescreening G_mRRHO run @ c1 was successful for CONF63/rrho_part1: 0.13193283 S_rot(sym)= 0.0000000 using= 0.1319328
The prescreening G_mRRHO run @ c1 was successful for CONF64/rrho_part1: 0.13168598 S_rot(sym)= 0.0000000 using= 0.1316860
The prescreening G_mRRHO run @ c1 was successful for CONF65/rrho_part1: 0.13143050 S_rot(sym)= 0.0000000 using= 0.1314305
The prescreening G_mRRHO run @ c1 was successful for CONF66/rrho_part1: 0.13141124 S_rot(sym)= 0.0000000 using= 0.1314112
The prescreening G_mRRHO run @ c1 was successful for CONF67/rrho_part1: 0.13143590 S_rot(sym)= 0.0000000 using= 0.1314359
The prescreening G_mRRHO run @ c1 was successful for CONF68/rrho_part1: 0.13126349 S_rot(sym)= 0.0000000 using= 0.1312635
The prescreening G_mRRHO run @ c1 was successful for CONF69/rrho_part1: 0.13171550 S_rot(sym)= 0.0000000 using= 0.1317155
The prescreening G_mRRHO run @ c1 was successful for CONF70/rrho_part1: 0.13221847 S_rot(sym)= 0.0000000 using= 0.1322185
The prescreening G_mRRHO run @ c1 was successful for CONF71/rrho_part1: 0.13204914 S_rot(sym)= 0.0000000 using= 0.1320491
The prescreening G_mRRHO run @ c1 was successful for CONF72/rrho_part1: 0.13219824 S_rot(sym)= 0.0000000 using= 0.1321982
The prescreening G_mRRHO run @ cs was successful for CONF73/rrho_part1: 0.13064925 S_rot(sym)= 0.0000000 using= 0.1306492
The prescreening G_mRRHO run @ c1 was successful for CONF74/rrho_part1: 0.13071055 S_rot(sym)= 0.0000000 using= 0.1307105
The prescreening G_mRRHO run @ c1 was successful for CONF75/rrho_part1: 0.13113590 S_rot(sym)= 0.0000000 using= 0.1311359
The prescreening G_mRRHO run @ c1 was successful for CONF76/rrho_part1: 0.13074623 S_rot(sym)= 0.0000000 using= 0.1307462
The prescreening G_mRRHO run @ c1 was successful for CONF77/rrho_part1: 0.13239450 S_rot(sym)= 0.0000000 using= 0.1323945
The prescreening G_mRRHO run @ c1 was successful for CONF80/rrho_part1: 0.13162386 S_rot(sym)= 0.0000000 using= 0.1316239
--------------------------------------------------
* Gibbs free energies of part1 *
--------------------------------------------------
CONF# G(GFNn-xTB) ΔG(GFNn-xTB) E [Eh] Gsolv [Eh] GmRRHO [Eh] Gtot ΔGtot
[a.u.] [kcal/mol] r2scan-3c COSMO-RS-normal GFN2 [Eh] [kcal/mol]
[r2scan-3c] [alpb]-bhess
CONF1 -22.8766770 0.23 -310.9770910 -0.0074950 0.1326685 -310.8519175 0.00 <------
CONF2 -22.8763699 0.43 -310.9757758 -0.0074459 0.1323700 -310.8508517 0.67
CONF3 -22.8763549 0.44 -310.9758415 -0.0074181 0.1323833 -310.8508762 0.65
CONF4 -22.8756821 0.86 -310.9758145 -0.0064451 0.1327399 -310.8495197 1.50
CONF5 -22.8763035 0.47 -310.9743875 -0.0079316 0.1320717 -310.8502473 1.05
CONF6 -22.8760337 0.64 -310.9745964 -0.0073222 0.1322715 -310.8496471 1.42
CONF7 -22.8762129 0.53 -310.9744677 -0.0079811 0.1320390 -310.8504098 0.95
CONF8 -22.8752495 1.13 -310.9757295 -0.0070120 0.1329856 -310.8497560 1.36
CONF9 -22.8760410 0.63 -310.9745570 -0.0077339 0.1320882 -310.8502027 1.08
CONF10 -22.8750444 1.26 -310.9743226 -0.0074543 0.1330511 -310.8487259 2.00
CONF11 -22.8760005 0.66 -310.9744861 -0.0076943 0.1320896 -310.8500908 1.15
CONF12 -22.8762001 0.53 -310.9741780 -0.0080687 0.1318577 -310.8503890 0.96
CONF13 -22.8762933 0.48 -310.9741538 -0.0081364 0.1317281 -310.8505620 0.85
CONF14 -22.8765524 0.31 -310.9733007 -0.0076717 0.1313324 -310.8496400 1.43
CONF15 -22.8762534 0.50 -310.9735503 -0.0077130 0.1315801 -310.8496831 1.40
CONF16 -22.8767990 0.16 -310.9731490 -0.0077013 0.1310271 -310.8498233 1.31
CONF17 -22.8762804 0.48 -310.9733525 -0.0077046 0.1315296 -310.8495276 1.50
CONF18 -22.8759386 0.70 -310.9730612 -0.0079772 0.1318188 -310.8492196 1.69
CONF19 -22.8759479 0.69 -310.9730253 -0.0079799 0.1317994 -310.8492059 1.70
CONF20 -22.8755180 0.96 -310.9743520 -0.0069489 0.1321821 -310.8491187 1.76
CONF21 -22.8758510 0.75 -310.9732578 -0.0073096 0.1318055 -310.8487618 1.98
CONF22 -22.8763409 0.45 -310.9729525 -0.0081110 0.1313146 -310.8497490 1.36
CONF23 -22.8758891 0.73 -310.9732776 -0.0077366 0.1317378 -310.8492764 1.66
CONF24 -22.8763697 0.43 -310.9728437 -0.0081699 0.1312395 -310.8497741 1.34
CONF25 -22.8763790 0.42 -310.9722243 -0.0076607 0.1312154 -310.8486695 2.04
CONF26 -22.8750959 1.23 -310.9734086 -0.0078730 0.1325555 -310.8487261 2.00
CONF27 -22.8765150 0.34 -310.9727742 -0.0081509 0.1310906 -310.8498344 1.31
CONF28 -22.8750534 1.25 -310.9733748 -0.0080349 0.1324482 -310.8489615 1.85
CONF29 -22.8750012 1.29 -310.9733671 -0.0080296 0.1324991 -310.8488977 1.89
CONF30 -22.8755216 0.96 -310.9740731 -0.0078530 0.1319520 -310.8499741 1.22
CONF31 -22.8755807 0.92 -310.9741187 -0.0076815 0.1318863 -310.8499139 1.26
CONF32 -22.8755462 0.94 -310.9741085 -0.0077049 0.1318916 -310.8499219 1.25
CONF33 -22.8767317 0.20 -310.9721512 -0.0080967 0.1306803 -310.8495676 1.47
CONF34 -22.8770512 0.00 -310.9718597 -0.0081208 0.1303506 -310.8496300 1.44
CONF35 -22.8751904 1.17 -310.9733092 -0.0074483 0.1321290 -310.8486284 2.06
CONF36 -22.8744834 1.61 -310.9738901 -0.0072987 0.1326658 -310.8485230 2.13
CONF38 -22.8760327 0.64 -310.9726276 -0.0079611 0.1309766 -310.8496121 1.45
CONF39 -22.8760598 0.62 -310.9726003 -0.0080084 0.1309313 -310.8496774 1.41
CONF40 -22.8757658 0.81 -310.9726416 -0.0080069 0.1312202 -310.8494284 1.56
CONF41 -22.8752839 1.11 -310.9727955 -0.0078562 0.1316829 -310.8489688 1.85
CONF42 -22.8755375 0.95 -310.9728220 -0.0077720 0.1314010 -310.8491929 1.71
CONF43 -22.8742642 1.75 -310.9728106 -0.0068611 0.1326503 -310.8470214 3.07
CONF44 -22.8760681 0.62 -310.9717566 -0.0079697 0.1307787 -310.8489476 1.86
CONF45 -22.8757966 0.79 -310.9718972 -0.0080084 0.1310466 -310.8488590 1.92
CONF46 -22.8763683 0.43 -310.9717215 -0.0079175 0.1304641 -310.8491750 1.72
CONF47 -22.8758561 0.75 -310.9718636 -0.0079323 0.1309680 -310.8488280 1.94
CONF48 -22.8740903 1.86 -310.9734980 -0.0075490 0.1327204 -310.8483266 2.25
CONF49 -22.8747806 1.42 -310.9729840 -0.0075986 0.1319713 -310.8486113 2.07
CONF50 -22.8748157 1.40 -310.9727818 -0.0077144 0.1319001 -310.8485961 2.08
CONF54 -22.8748996 1.35 -310.9722719 -0.0081389 0.1317495 -310.8486613 2.04
CONF55 -22.8748897 1.36 -310.9720817 -0.0080338 0.1317318 -310.8483837 2.22
CONF56 -22.8745955 1.54 -310.9713906 -0.0080262 0.1317232 -310.8476936 2.65
CONF57 -22.8744899 1.61 -310.9744054 -0.0078789 0.1317990 -310.8504853 0.90
CONF58 -22.8738669 2.00 -310.9760271 -0.0065699 0.1323059 -310.8502911 1.02
CONF59 -22.8739381 1.95 -310.9759465 -0.0066033 0.1322326 -310.8503172 1.00
CONF60 -22.8741676 1.81 -310.9758398 -0.0066532 0.1319909 -310.8505021 0.89
CONF61 -22.8744403 1.64 -310.9755805 -0.0067207 0.1316846 -310.8506166 0.82
CONF62 -22.8742160 1.78 -310.9754193 -0.0067962 0.1318791 -310.8503364 0.99
CONF63 -22.8740132 1.91 -310.9722754 -0.0077430 0.1319328 -310.8480856 2.40
CONF64 -22.8740435 1.89 -310.9745205 -0.0072754 0.1316860 -310.8501099 1.13
CONF65 -22.8742969 1.73 -310.9738656 -0.0082307 0.1314305 -310.8506657 0.79
CONF66 -22.8743136 1.72 -310.9738666 -0.0082254 0.1314112 -310.8506808 0.78
CONF67 -22.8742913 1.73 -310.9738672 -0.0082195 0.1314359 -310.8506508 0.79
CONF68 -22.8744568 1.63 -310.9738563 -0.0082409 0.1312635 -310.8508337 0.68
CONF69 -22.8738899 1.98 -310.9744729 -0.0076079 0.1317155 -310.8503653 0.97
CONF70 -22.8732375 2.39 -310.9732794 -0.0079299 0.1322185 -310.8489908 1.84
CONF71 -22.8734034 2.29 -310.9732799 -0.0079750 0.1320491 -310.8492058 1.70
CONF72 -22.8732531 2.38 -310.9732609 -0.0079122 0.1321982 -310.8489748 1.85
CONF73 -22.8747538 1.44 -310.9725927 -0.0082086 0.1306492 -310.8501521 1.11
CONF74 -22.8745698 1.56 -310.9729596 -0.0077264 0.1307105 -310.8499754 1.22
CONF75 -22.8741027 1.85 -310.9726279 -0.0082021 0.1311359 -310.8496940 1.40
CONF76 -22.8742554 1.75 -310.9724113 -0.0076527 0.1307462 -310.8493178 1.63
CONF77 -22.8723742 2.93 -310.9739372 -0.0071637 0.1323945 -310.8487064 2.02
CONF80 -22.8726191 2.78 -310.9726319 -0.0075430 0.1316239 -310.8485510 2.11
Number of conformers observed within the following ΔG windows:
ΔG [kcal/mol] #CONF sum(Boltzmann_weights)
---------------------------------------------
0 - 0.5 1 0.11
0 - 1.0 15 0.51
0 - 1.5 40 0.84
0 - 2.0 59 0.96
0 - 2.5 72 1.00
0 - 3.0 73 1.00
0 - 3.5 74 1.00
---------------------------------------------
Additional global 'fuzzy-threshold' based on the standard deviation of (G_mRRHO):
Std_dev(G_mRRHO) = 0.386 kcal/mol
Fuzzythreshold = 0.525 kcal/mol
Final sorting threshold G_thr(1) = 3.500 + 0.525 = 4.025 kcal/mol
Spearman correlation coefficient between (E + Solv) and (E + Solv + mRRHO) = 0.743
All relative (free) energies are below the initial G_thr(1) threshold of 3.5 kcal/mol.
All conformers are considered further.
Calculating Boltzmann averaged free energy of ensemble on input geometries (not DFT optimized)!
temperature /K: avE(T) /a.u. avGmRRHO(T) /a.u. avGsolv(T) /a.u. avG(T) /a.u.
----------------------------------------------------------------------------------------------------
298.15 -310.9744421 0.1318263 -0.0076646 -310.8502804 <<==part1==
----------------------------------------------------------------------------------------------------
Backing up enso_ensemble_part1.xyz to enso_ensemble_part1.xyz.1.
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>END of Part1<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
Ran part1 in 0.2158 seconds
----------------------------------------------------------------------------------------------------
CRE OPTIMIZATION - PART2
----------------------------------------------------------------------------------------------------
program: tm
functional for part2: r2scan-3c
basis set for part2: def2-mTZVPP
using the xTB-optimizer for optimization: on
using the new ensemble optimizer: on
optimize all conformers below this G_thr(opt,2) threshold: 2.5
Spearman threshold: 0.939
number of optimization iterations: 8
radsize: 10
optimization level in part2: lax
solvent: chcl3
solvent model applied in the optimization: dcosmors
solvent model for Gsolv contribution: cosmors
temperature: 298.15
evalulate at different temperatures: on
temperature range: 273.15, 278.15, 283.15, 288.15, ...
Boltzmann sum threshold G_thr(2) for sorting in part2: 95.0
calculate mRRHO contribution: on
program for mRRHO contribution: xtb
GFN version for mRRHO and/or GBSA_Gsolv: gfn2
Apply constraint to input geometry during mRRHO calculation: on
Optimizing geometries at DFT level with implicit solvation!
The optimization was performed before for:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF10, CONF11
CONF12, CONF13, CONF14, CONF15, CONF16, CONF17, CONF18, CONF19, CONF20, CONF21, CONF22
CONF23, CONF24, CONF25, CONF26, CONF27, CONF28, CONF29, CONF30, CONF31, CONF32, CONF33
CONF34, CONF35, CONF36, CONF38, CONF39, CONF40, CONF41, CONF42, CONF43, CONF44, CONF45
CONF46, CONF47, CONF48, CONF49, CONF50, CONF54, CONF55, CONF56, CONF57, CONF58, CONF59
CONF60, CONF61, CONF62, CONF63, CONF64, CONF65, CONF66, CONF67, CONF68, CONF69, CONF70
CONF71, CONF72, CONF73, CONF74, CONF75, CONF76, CONF77, CONF80
No conformers are considered additionally.
Converged optimization for CONF1 after 4 cycles: -310.9928951
Converged optimization for CONF2 after 4 cycles: -310.9915057
Converged optimization for CONF3 after 4 cycles: -310.9915272
Converged optimization for CONF4 after 11 cycles: -310.9913871
Converged optimization for CONF5 after 5 cycles: -310.9909604
Converged optimization for CONF6 after 8 cycles: -310.9911402
Converged optimization for CONF7 after 5 cycles: -310.9909887
Converged optimization for CONF8 after 9 cycles: -310.9914445
Converged optimization for CONF9 after 5 cycles: -310.9911283
Converged optimization for CONF10 after 7 cycles: -310.9903863
Converged optimization for CONF11 after 4 cycles: -310.9910044
Converged optimization for CONF12 after 5 cycles: -310.9909903
Converged optimization for CONF13 after 5 cycles: -310.9909924
Converged optimization for CONF14 after 29 cycles: -310.9910740
Converged optimization for CONF15 after 19 cycles: -310.9910621
Converged optimization for CONF16 after 6 cycles: -310.9897093
Converged optimization for CONF17 after 26 cycles: -310.9910574
Converged optimization for CONF18 after 31 cycles: -310.9909754
Converged optimization for CONF19 after 24 cycles: -310.9909336
Converged optimization for CONF20 after 23 cycles: -310.9913458
Converged optimization for CONF21 after 5 cycles: -310.9897873
Converged optimization for CONF22 after 9 cycles: -310.9899585
Converged optimization for CONF23 after 4 cycles: -310.9898048
Converged optimization for CONF24 after 32 cycles: -310.9909071
Converged optimization for CONF25 after 3 cycles: -310.9887299
Converged optimization for CONF26 after 5 cycles: -310.9899537
Converged optimization for CONF27 after 4 cycles: -310.9896269
Converged optimization for CONF28 after 3 cycles: -310.9898925
Converged optimization for CONF29 after 3 cycles: -310.9898694
Converged optimization for CONF30 after 8 cycles: -310.9908798
Converged optimization for CONF31 after 7 cycles: -310.9909051
Converged optimization for CONF32 after 8 cycles: -310.9908698
Converged optimization for CONF33 after 27 cycles: -310.9907939
Converged optimization for CONF34 after 9 cycles: -310.9888546
Converged optimization for CONF35 after 30 cycles: -310.9912973
Converged optimization for CONF36 after 8 cycles: -310.9902292
Converged optimization for CONF38 after 8 cycles: -310.9894942
Converged optimization for CONF39 after 8 cycles: -310.9894519
Converged optimization for CONF40 after 9 cycles: -310.9895310
Converged optimization for CONF41 after 8 cycles: -310.9896034
Converged optimization for CONF42 after 29 cycles: -310.9908205
Converged optimization for CONF43 after 9 cycles: -310.9888817
Converged optimization for CONF44 after 29 cycles: -310.9893961
Converged optimization for CONF45 after 10 cycles: -310.9888378
Converged optimization for CONF46 after 9 cycles: -310.9885849
Converged optimization for CONF47 after 33 cycles: -310.9907837
Converged optimization for CONF48 after 9 cycles: -310.9898909
Converged optimization for CONF49 after 22 cycles: -310.9901228
Converged optimization for CONF50 after 9 cycles: -310.9894140
Converged optimization for CONF54 after 5 cycles: -310.9890574
Converged optimization for CONF55 after 5 cycles: -310.9888447
Converged optimization for CONF56 after 26 cycles: -310.9897996
Converged optimization for CONF57 after 3 cycles: -310.9908663
Converged optimization for CONF58 after 5 cycles: -310.9912796
Converged optimization for CONF59 after 5 cycles: -310.9912368
Converged optimization for CONF60 after 5 cycles: -310.9912306
Converged optimization for CONF61 after 5 cycles: -310.9911066
Converged optimization for CONF62 after 5 cycles: -310.9910648
Converged optimization for CONF63 after 9 cycles: -310.9893066
Converged optimization for CONF64 after 8 cycles: -310.9909454
Converged optimization for CONF65 after 3 cycles: -310.9907096
Converged optimization for CONF66 after 3 cycles: -310.9907059
Converged optimization for CONF67 after 3 cycles: -310.9907099
Converged optimization for CONF68 after 3 cycles: -310.9907059
Converged optimization for CONF69 after 4 cycles: -310.9908881
Converged optimization for CONF70 after 3 cycles: -310.9897851
Converged optimization for CONF71 after 4 cycles: -310.9898168
Converged optimization for CONF72 after 3 cycles: -310.9897523
Converged optimization for CONF73 after 3 cycles: -310.9894063
Converged optimization for CONF74 after 4 cycles: -310.9894374
Converged optimization for CONF75 after 3 cycles: -310.9894370
Converged optimization for CONF76 after 9 cycles: -310.9890638
Converged optimization for CONF77 after 8 cycles: -310.9901268
Converged optimization for CONF80 after 8 cycles: -310.9890836
Calculating single-point energies and solvation contribution (G_solv)!
The low level gsolv calculation was calculated before for:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF10, CONF11
CONF12, CONF13, CONF14, CONF15, CONF16, CONF17, CONF18, CONF19, CONF20, CONF21, CONF22
CONF23, CONF24, CONF25, CONF26, CONF27, CONF28, CONF29, CONF30, CONF31, CONF32, CONF33
CONF34, CONF35, CONF36, CONF38, CONF39, CONF40, CONF41, CONF42, CONF43, CONF44, CONF45
CONF46, CONF47, CONF48, CONF49, CONF50, CONF54, CONF55, CONF56, CONF57, CONF58, CONF59
CONF60, CONF61, CONF62, CONF63, CONF64, CONF65, CONF66, CONF67, CONF68, CONF69, CONF70
CONF71, CONF72, CONF73, CONF74, CONF75, CONF76, CONF77, CONF80
No conformers are considered additionally.
lowlevel COSMO-RS calculation was successful for CONF1/r2scan-3c/COSMO: -0.0078485
lowlevel COSMO-RS calculation was successful for CONF2/r2scan-3c/COSMO: -0.0077870
lowlevel COSMO-RS calculation was successful for CONF3/r2scan-3c/COSMO: -0.0077611
lowlevel COSMO-RS calculation was successful for CONF4/r2scan-3c/COSMO: -0.0069825
lowlevel COSMO-RS calculation was successful for CONF5/r2scan-3c/COSMO: -0.0082178
lowlevel COSMO-RS calculation was successful for CONF6/r2scan-3c/COSMO: -0.0077038
lowlevel COSMO-RS calculation was successful for CONF7/r2scan-3c/COSMO: -0.0082944
lowlevel COSMO-RS calculation was successful for CONF8/r2scan-3c/COSMO: -0.0073678
lowlevel COSMO-RS calculation was successful for CONF9/r2scan-3c/COSMO: -0.0080528
lowlevel COSMO-RS calculation was successful for CONF10/r2scan-3c/COSMO: -0.0078398
lowlevel COSMO-RS calculation was successful for CONF11/r2scan-3c/COSMO: -0.0080269
lowlevel COSMO-RS calculation was successful for CONF12/r2scan-3c/COSMO: -0.0083661
lowlevel COSMO-RS calculation was successful for CONF13/r2scan-3c/COSMO: -0.0084110
lowlevel COSMO-RS calculation was successful for CONF14/r2scan-3c/COSMO: -0.0079817
lowlevel COSMO-RS calculation was successful for CONF15/r2scan-3c/COSMO: -0.0080490
lowlevel COSMO-RS calculation was successful for CONF16/r2scan-3c/COSMO: -0.0080047
lowlevel COSMO-RS calculation was successful for CONF17/r2scan-3c/COSMO: -0.0080391
lowlevel COSMO-RS calculation was successful for CONF18/r2scan-3c/COSMO: -0.0082018
lowlevel COSMO-RS calculation was successful for CONF19/r2scan-3c/COSMO: -0.0082043
lowlevel COSMO-RS calculation was successful for CONF20/r2scan-3c/COSMO: -0.0071146
lowlevel COSMO-RS calculation was successful for CONF21/r2scan-3c/COSMO: -0.0076613
lowlevel COSMO-RS calculation was successful for CONF22/r2scan-3c/COSMO: -0.0083778
lowlevel COSMO-RS calculation was successful for CONF23/r2scan-3c/COSMO: -0.0080352
lowlevel COSMO-RS calculation was successful for CONF24/r2scan-3c/COSMO: -0.0084452
lowlevel COSMO-RS calculation was successful for CONF25/r2scan-3c/COSMO: -0.0079553
lowlevel COSMO-RS calculation was successful for CONF26/r2scan-3c/COSMO: -0.0081645
lowlevel COSMO-RS calculation was successful for CONF27/r2scan-3c/COSMO: -0.0084651
lowlevel COSMO-RS calculation was successful for CONF28/r2scan-3c/COSMO: -0.0082860
lowlevel COSMO-RS calculation was successful for CONF29/r2scan-3c/COSMO: -0.0082883
lowlevel COSMO-RS calculation was successful for CONF30/r2scan-3c/COSMO: -0.0081326
lowlevel COSMO-RS calculation was successful for CONF31/r2scan-3c/COSMO: -0.0079707
lowlevel COSMO-RS calculation was successful for CONF32/r2scan-3c/COSMO: -0.0079108
lowlevel COSMO-RS calculation was successful for CONF33/r2scan-3c/COSMO: -0.0083655
lowlevel COSMO-RS calculation was successful for CONF34/r2scan-3c/COSMO: -0.0084222
lowlevel COSMO-RS calculation was successful for CONF35/r2scan-3c/COSMO: -0.0074835
lowlevel COSMO-RS calculation was successful for CONF36/r2scan-3c/COSMO: -0.0076260
lowlevel COSMO-RS calculation was successful for CONF38/r2scan-3c/COSMO: -0.0082618
lowlevel COSMO-RS calculation was successful for CONF39/r2scan-3c/COSMO: -0.0083128
lowlevel COSMO-RS calculation was successful for CONF40/r2scan-3c/COSMO: -0.0082993
lowlevel COSMO-RS calculation was successful for CONF41/r2scan-3c/COSMO: -0.0081040
lowlevel COSMO-RS calculation was successful for CONF42/r2scan-3c/COSMO: -0.0081352
lowlevel COSMO-RS calculation was successful for CONF43/r2scan-3c/COSMO: -0.0074285
lowlevel COSMO-RS calculation was successful for CONF44/r2scan-3c/COSMO: -0.0083126
lowlevel COSMO-RS calculation was successful for CONF45/r2scan-3c/COSMO: -0.0083314
lowlevel COSMO-RS calculation was successful for CONF46/r2scan-3c/COSMO: -0.0082362
lowlevel COSMO-RS calculation was successful for CONF47/r2scan-3c/COSMO: -0.0081979
lowlevel COSMO-RS calculation was successful for CONF48/r2scan-3c/COSMO: -0.0077822
lowlevel COSMO-RS calculation was successful for CONF49/r2scan-3c/COSMO: -0.0077532
lowlevel COSMO-RS calculation was successful for CONF50/r2scan-3c/COSMO: -0.0079805
lowlevel COSMO-RS calculation was successful for CONF54/r2scan-3c/COSMO: -0.0084644
lowlevel COSMO-RS calculation was successful for CONF55/r2scan-3c/COSMO: -0.0083328
lowlevel COSMO-RS calculation was successful for CONF56/r2scan-3c/COSMO: -0.0083829
lowlevel COSMO-RS calculation was successful for CONF57/r2scan-3c/COSMO: -0.0080724
lowlevel COSMO-RS calculation was successful for CONF58/r2scan-3c/COSMO: -0.0068709
lowlevel COSMO-RS calculation was successful for CONF59/r2scan-3c/COSMO: -0.0068729
lowlevel COSMO-RS calculation was successful for CONF60/r2scan-3c/COSMO: -0.0069148
lowlevel COSMO-RS calculation was successful for CONF61/r2scan-3c/COSMO: -0.0070052
lowlevel COSMO-RS calculation was successful for CONF62/r2scan-3c/COSMO: -0.0070689
lowlevel COSMO-RS calculation was successful for CONF63/r2scan-3c/COSMO: -0.0078661
lowlevel COSMO-RS calculation was successful for CONF64/r2scan-3c/COSMO: -0.0076605
lowlevel COSMO-RS calculation was successful for CONF65/r2scan-3c/COSMO: -0.0084410
lowlevel COSMO-RS calculation was successful for CONF66/r2scan-3c/COSMO: -0.0084406
lowlevel COSMO-RS calculation was successful for CONF67/r2scan-3c/COSMO: -0.0084297
lowlevel COSMO-RS calculation was successful for CONF68/r2scan-3c/COSMO: -0.0084359
lowlevel COSMO-RS calculation was successful for CONF69/r2scan-3c/COSMO: -0.0078726
lowlevel COSMO-RS calculation was successful for CONF70/r2scan-3c/COSMO: -0.0081421
lowlevel COSMO-RS calculation was successful for CONF71/r2scan-3c/COSMO: -0.0082042
lowlevel COSMO-RS calculation was successful for CONF72/r2scan-3c/COSMO: -0.0081331
lowlevel COSMO-RS calculation was successful for CONF73/r2scan-3c/COSMO: -0.0084100
lowlevel COSMO-RS calculation was successful for CONF74/r2scan-3c/COSMO: -0.0080078
lowlevel COSMO-RS calculation was successful for CONF75/r2scan-3c/COSMO: -0.0083993
lowlevel COSMO-RS calculation was successful for CONF76/r2scan-3c/COSMO: -0.0079706
lowlevel COSMO-RS calculation was successful for CONF77/r2scan-3c/COSMO: -0.0073916
lowlevel COSMO-RS calculation was successful for CONF80/r2scan-3c/COSMO: -0.0078527
Calculating lowlevel G_mRRHO with implicit solvation on DFT geometry!
The G_mRRHO calculation was performed before for:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF10, CONF11
CONF12, CONF13, CONF14, CONF15, CONF16, CONF17, CONF18, CONF19, CONF20, CONF21, CONF22
CONF23, CONF24, CONF25, CONF26, CONF27, CONF28, CONF29, CONF30, CONF31, CONF32, CONF33
CONF34, CONF35, CONF36, CONF38, CONF39, CONF40, CONF41, CONF42, CONF43, CONF44, CONF45
CONF46, CONF47, CONF48, CONF49, CONF50, CONF54, CONF55, CONF56, CONF57, CONF58, CONF59
CONF60, CONF61, CONF62, CONF63, CONF64, CONF65, CONF66, CONF67, CONF68, CONF69, CONF70
CONF71, CONF72, CONF73, CONF74, CONF75, CONF76, CONF77, CONF80
No conformers are considered additionally.
The lowlevel G_mRRHO calculation @ c1 was successful for CONF1/rrho_part2: 0.13267798 S_rot(sym)= 0.0000000 using= 0.1326780
The lowlevel G_mRRHO calculation @ c1 was successful for CONF2/rrho_part2: 0.13228210 S_rot(sym)= 0.0000000 using= 0.1322821
The lowlevel G_mRRHO calculation @ c1 was successful for CONF3/rrho_part2: 0.13228741 S_rot(sym)= 0.0000000 using= 0.1322874
The lowlevel G_mRRHO calculation @ c1 was successful for CONF4/rrho_part2: 0.13266443 S_rot(sym)= 0.0000000 using= 0.1326644
The lowlevel G_mRRHO calculation @ c1 was successful for CONF5/rrho_part2: 0.13209144 S_rot(sym)= 0.0000000 using= 0.1320914
The lowlevel G_mRRHO calculation @ c1 was successful for CONF6/rrho_part2: 0.13224841 S_rot(sym)= 0.0000000 using= 0.1322484
The lowlevel G_mRRHO calculation @ c1 was successful for CONF7/rrho_part2: 0.13211950 S_rot(sym)= 0.0000000 using= 0.1321195
The lowlevel G_mRRHO calculation @ c1 was successful for CONF8/rrho_part2: 0.13294841 S_rot(sym)= 0.0000000 using= 0.1329484
The lowlevel G_mRRHO calculation @ c1 was successful for CONF9/rrho_part2: 0.13206130 S_rot(sym)= 0.0000000 using= 0.1320613
The lowlevel G_mRRHO calculation @ c1 was successful for CONF10/rrho_part2: 0.13305963 S_rot(sym)= 0.0000000 using= 0.1330596
The lowlevel G_mRRHO calculation @ c1 was successful for CONF11/rrho_part2: 0.13211958 S_rot(sym)= 0.0000000 using= 0.1321196
The lowlevel G_mRRHO calculation @ c1 was successful for CONF12/rrho_part2: 0.13187169 S_rot(sym)= 0.0000000 using= 0.1318717
The lowlevel G_mRRHO calculation @ c1 was successful for CONF13/rrho_part2: 0.13174523 S_rot(sym)= 0.0000000 using= 0.1317452
The lowlevel G_mRRHO calculation @ c1 was successful for CONF14/rrho_part2: 0.13210247 S_rot(sym)= 0.0000000 using= 0.1321025
The lowlevel G_mRRHO calculation @ c1 was successful for CONF15/rrho_part2: 0.13204381 S_rot(sym)= 0.0000000 using= 0.1320438
The lowlevel G_mRRHO calculation @ c1 was successful for CONF16/rrho_part2: 0.13099766 S_rot(sym)= 0.0000000 using= 0.1309977
The lowlevel G_mRRHO calculation @ c1 was successful for CONF17/rrho_part2: 0.13199645 S_rot(sym)= 0.0000000 using= 0.1319964
The lowlevel G_mRRHO calculation @ c1 was successful for CONF18/rrho_part2: 0.13219580 S_rot(sym)= 0.0000000 using= 0.1321958
The lowlevel G_mRRHO calculation @ c1 was successful for CONF19/rrho_part2: 0.13216578 S_rot(sym)= 0.0000000 using= 0.1321658
The lowlevel G_mRRHO calculation @ c1 was successful for CONF20/rrho_part2: 0.13224679 S_rot(sym)= 0.0000000 using= 0.1322468
The lowlevel G_mRRHO calculation @ c1 was successful for CONF21/rrho_part2: 0.13172718 S_rot(sym)= 0.0000000 using= 0.1317272
The lowlevel G_mRRHO calculation @ c1 was successful for CONF22/rrho_part2: 0.13133611 S_rot(sym)= 0.0000000 using= 0.1313361
The lowlevel G_mRRHO calculation @ c1 was successful for CONF23/rrho_part2: 0.13189489 S_rot(sym)= 0.0000000 using= 0.1318949
The lowlevel G_mRRHO calculation @ c1 was successful for CONF24/rrho_part2: 0.13168408 S_rot(sym)= 0.0000000 using= 0.1316841
The lowlevel G_mRRHO calculation @ c1 was successful for CONF25/rrho_part2: 0.13133376 S_rot(sym)= 0.0000000 using= 0.1313338
The lowlevel G_mRRHO calculation @ c1 was successful for CONF26/rrho_part2: 0.13255296 S_rot(sym)= 0.0000000 using= 0.1325530
The lowlevel G_mRRHO calculation @ c1 was successful for CONF27/rrho_part2: 0.13116594 S_rot(sym)= 0.0000000 using= 0.1311659
The lowlevel G_mRRHO calculation @ c1 was successful for CONF28/rrho_part2: 0.13241540 S_rot(sym)= 0.0000000 using= 0.1324154
The lowlevel G_mRRHO calculation @ c1 was successful for CONF29/rrho_part2: 0.13247407 S_rot(sym)= 0.0000000 using= 0.1324741
The lowlevel G_mRRHO calculation @ c1 was successful for CONF30/rrho_part2: 0.13194576 S_rot(sym)= 0.0000000 using= 0.1319458
The lowlevel G_mRRHO calculation @ c1 was successful for CONF31/rrho_part2: 0.13190564 S_rot(sym)= 0.0000000 using= 0.1319056
The lowlevel G_mRRHO calculation @ c1 was successful for CONF32/rrho_part2: 0.13186994 S_rot(sym)= 0.0000000 using= 0.1318699
The lowlevel G_mRRHO calculation @ c1 was successful for CONF33/rrho_part2: 0.13176311 S_rot(sym)= 0.0000000 using= 0.1317631
The lowlevel G_mRRHO calculation @ c1 was successful for CONF34/rrho_part2: 0.13088131 S_rot(sym)= 0.0000000 using= 0.1308813
The lowlevel G_mRRHO calculation @ c1 was successful for CONF35/rrho_part2: 0.13281166 S_rot(sym)= 0.0000000 using= 0.1328117
The lowlevel G_mRRHO calculation @ c1 was successful for CONF36/rrho_part2: 0.13268291 S_rot(sym)= 0.0000000 using= 0.1326829
The lowlevel G_mRRHO calculation @ c1 was successful for CONF38/rrho_part2: 0.13130501 S_rot(sym)= 0.0000000 using= 0.1313050
The lowlevel G_mRRHO calculation @ c1 was successful for CONF39/rrho_part2: 0.13099672 S_rot(sym)= 0.0000000 using= 0.1309967
The lowlevel G_mRRHO calculation @ c1 was successful for CONF40/rrho_part2: 0.13122440 S_rot(sym)= 0.0000000 using= 0.1312244
The lowlevel G_mRRHO calculation @ c1 was successful for CONF41/rrho_part2: 0.13173201 S_rot(sym)= 0.0000000 using= 0.1317320
The lowlevel G_mRRHO calculation @ c1 was successful for CONF42/rrho_part2: 0.13191230 S_rot(sym)= 0.0000000 using= 0.1319123
The lowlevel G_mRRHO calculation @ c1 was successful for CONF43/rrho_part2: 0.13263792 S_rot(sym)= 0.0000000 using= 0.1326379
The lowlevel G_mRRHO calculation @ c1 was successful for CONF44/rrho_part2: 0.13086578 S_rot(sym)= 0.0000000 using= 0.1308658
The lowlevel G_mRRHO calculation @ c1 was successful for CONF45/rrho_part2: 0.13103817 S_rot(sym)= 0.0000000 using= 0.1310382
The lowlevel G_mRRHO calculation @ c1 was successful for CONF46/rrho_part2: 0.13006721 S_rot(sym)= 0.0000000 using= 0.1300672
The lowlevel G_mRRHO calculation @ c1 was successful for CONF47/rrho_part2: 0.13183812 S_rot(sym)= 0.0000000 using= 0.1318381
The lowlevel G_mRRHO calculation @ c1 was successful for CONF48/rrho_part2: 0.13273601 S_rot(sym)= 0.0000000 using= 0.1327360
The lowlevel G_mRRHO calculation @ c1 was successful for CONF49/rrho_part2: 0.13250143 S_rot(sym)= 0.0000000 using= 0.1325014
The lowlevel G_mRRHO calculation @ c1 was successful for CONF50/rrho_part2: 0.13201737 S_rot(sym)= 0.0000000 using= 0.1320174
The lowlevel G_mRRHO calculation @ c1 was successful for CONF54/rrho_part2: 0.13103567 S_rot(sym)= 0.0000000 using= 0.1310357
The lowlevel G_mRRHO calculation @ c1 was successful for CONF55/rrho_part2: 0.13175546 S_rot(sym)= 0.0000000 using= 0.1317555
The lowlevel G_mRRHO calculation @ c1 was successful for CONF56/rrho_part2: 0.13222114 S_rot(sym)= 0.0000000 using= 0.1322211
The lowlevel G_mRRHO calculation @ c1 was successful for CONF57/rrho_part2: 0.13183572 S_rot(sym)= 0.0000000 using= 0.1318357
The lowlevel G_mRRHO calculation @ c1 was successful for CONF58/rrho_part2: 0.13233692 S_rot(sym)= 0.0000000 using= 0.1323369
The lowlevel G_mRRHO calculation @ c1 was successful for CONF59/rrho_part2: 0.13229538 S_rot(sym)= 0.0000000 using= 0.1322954
The lowlevel G_mRRHO calculation @ c1 was successful for CONF60/rrho_part2: 0.13218498 S_rot(sym)= 0.0000000 using= 0.1321850
The lowlevel G_mRRHO calculation @ c1 was successful for CONF61/rrho_part2: 0.13186221 S_rot(sym)= 0.0000000 using= 0.1318622
The lowlevel G_mRRHO calculation @ c1 was successful for CONF62/rrho_part2: 0.13203598 S_rot(sym)= 0.0000000 using= 0.1320360
The lowlevel G_mRRHO calculation @ c1 was successful for CONF63/rrho_part2: 0.13227617 S_rot(sym)= 0.0000000 using= 0.1322762
The lowlevel G_mRRHO calculation @ c1 was successful for CONF64/rrho_part2: 0.13170436 S_rot(sym)= 0.0000000 using= 0.1317044
The lowlevel G_mRRHO calculation @ c1 was successful for CONF65/rrho_part2: 0.13142100 S_rot(sym)= 0.0000000 using= 0.1314210
The lowlevel G_mRRHO calculation @ c1 was successful for CONF66/rrho_part2: 0.13139665 S_rot(sym)= 0.0000000 using= 0.1313966
The lowlevel G_mRRHO calculation @ c1 was successful for CONF67/rrho_part2: 0.13142137 S_rot(sym)= 0.0000000 using= 0.1314214
The lowlevel G_mRRHO calculation @ c1 was successful for CONF68/rrho_part2: 0.13130488 S_rot(sym)= 0.0000000 using= 0.1313049
The lowlevel G_mRRHO calculation @ c1 was successful for CONF69/rrho_part2: 0.13171196 S_rot(sym)= 0.0000000 using= 0.1317120
The lowlevel G_mRRHO calculation @ c1 was successful for CONF70/rrho_part2: 0.13219741 S_rot(sym)= 0.0000000 using= 0.1321974
The lowlevel G_mRRHO calculation @ c1 was successful for CONF71/rrho_part2: 0.13212971 S_rot(sym)= 0.0000000 using= 0.1321297
The lowlevel G_mRRHO calculation @ c1 was successful for CONF72/rrho_part2: 0.13217932 S_rot(sym)= 0.0000000 using= 0.1321793
The lowlevel G_mRRHO calculation @ cs was successful for CONF73/rrho_part2: 0.13062682 S_rot(sym)= 0.0000000 using= 0.1306268
The lowlevel G_mRRHO calculation @ c1 was successful for CONF74/rrho_part2: 0.13057213 S_rot(sym)= 0.0000000 using= 0.1305721
The lowlevel G_mRRHO calculation @ c1 was successful for CONF75/rrho_part2: 0.13120600 S_rot(sym)= 0.0000000 using= 0.1312060
The lowlevel G_mRRHO calculation @ c1 was successful for CONF76/rrho_part2: 0.13134242 S_rot(sym)= 0.0000000 using= 0.1313424
The lowlevel G_mRRHO calculation @ c1 was successful for CONF77/rrho_part2: 0.13238152 S_rot(sym)= 0.0000000 using= 0.1323815
The lowlevel G_mRRHO calculation @ c1 was successful for CONF80/rrho_part2: 0.13138485 S_rot(sym)= 0.0000000 using= 0.1313849
--------------------------------------------------
* Gibbs free energies of part2 *
--------------------------------------------------
CONF# E(GFNn-xTB) ΔE(GFNn-xTB) E [Eh] Gsolv [Eh] GmRRHO [Eh] Gtot ΔGtot Boltzmannweight
[a.u.] [kcal/mol] r2scan-3c COSMO-RS-normal GFN2 [Eh] [kcal/mol] % at 298.15 K
[r2scan-3c] [alpb]-bhess
CONF1 -23.0093455 0.00 -310.9779421 -0.0078485 0.1326780 -310.8531126 0.00 10.23 <------
CONF2 -23.0087399 0.38 -310.9766733 -0.0077870 0.1322821 -310.8521781 0.59 3.80
CONF3 -23.0087382 0.38 -310.9767072 -0.0077611 0.1322874 -310.8521809 0.58 3.81
CONF4 -23.0084220 0.58 -310.9769910 -0.0069825 0.1326644 -310.8513091 1.13 1.51
CONF5 -23.0083752 0.61 -310.9752433 -0.0082178 0.1320914 -310.8513697 1.09 1.62
CONF6 -23.0083052 0.65 -310.9755626 -0.0077038 0.1322484 -310.8510180 1.31 1.11
CONF7 -23.0082519 0.69 -310.9752725 -0.0082944 0.1321195 -310.8514474 1.04 1.75
CONF8 -23.0082350 0.70 -310.9769749 -0.0073678 0.1329484 -310.8513943 1.08 1.66
CONF9 -23.0081291 0.76 -310.9754631 -0.0080528 0.1320613 -310.8514546 1.04 1.77
CONF10 -23.0080955 0.78 -310.9754106 -0.0078398 0.1330596 -310.8501908 1.83 0.46
CONF11 -23.0080901 0.79 -310.9753819 -0.0080269 0.1321196 -310.8512892 1.14 1.48
CONF12 -23.0080578 0.81 -310.9749503 -0.0083661 0.1318717 -310.8514447 1.05 1.75
CONF13 -23.0080214 0.83 -310.9749185 -0.0084110 0.1317452 -310.8515843 0.96 2.03
CONF14 -23.0078848 0.92 -310.9754401 -0.0079817 0.1321025 -310.8513194 1.13 1.53
CONF15 -23.0078336 0.95 -310.9753877 -0.0080490 0.1320438 -310.8513929 1.08 1.66
CONF16 -23.0078261 0.95 -310.9740923 -0.0080047 0.1309977 -310.8510993 1.26 1.21
CONF17 -23.0078099 0.96 -310.9754046 -0.0080391 0.1319964 -310.8514473 1.04 1.75
CONF18 -23.0077574 1.00 -310.9753558 -0.0082018 0.1321958 -310.8513618 1.10 1.60
CONF19 -23.0077473 1.00 -310.9752809 -0.0082043 0.1321658 -310.8513194 1.13 1.53
CONF20 -23.0077001 1.03 -310.9766845 -0.0071146 0.1322468 -310.8515524 0.98 1.96
CONF21 -23.0076566 1.06 -310.9741790 -0.0076613 0.1317272 -310.8501131 1.88 0.43
CONF22 -23.0076554 1.06 -310.9738935 -0.0083778 0.1313361 -310.8509352 1.37 1.02
CONF23 -23.0076269 1.08 -310.9741656 -0.0080352 0.1318949 -310.8503059 1.76 0.52
CONF24 -23.0076092 1.09 -310.9748194 -0.0084452 0.1316841 -310.8515806 0.96 2.02
CONF25 -23.0075945 1.10 -310.9731415 -0.0079553 0.1313338 -310.8497630 2.10 0.29
CONF26 -23.0076514 1.06 -310.9742541 -0.0081645 0.1325530 -310.8498657 2.04 0.33
CONF27 -23.0076056 1.09 -310.9735485 -0.0084651 0.1311659 -310.8508477 1.42 0.93
CONF28 -23.0075015 1.16 -310.9741802 -0.0082860 0.1324154 -310.8500508 1.92 0.40
CONF29 -23.0075003 1.16 -310.9741720 -0.0082883 0.1324741 -310.8499863 1.96 0.37
CONF30 -23.0074736 1.17 -310.9751637 -0.0081326 0.1319458 -310.8513505 1.11 1.58
CONF31 -23.0074670 1.18 -310.9751571 -0.0079707 0.1319056 -310.8512222 1.19 1.38
CONF32 -23.0074378 1.20 -310.9751719 -0.0079108 0.1318699 -310.8512127 1.19 1.37
CONF33 -23.0074120 1.21 -310.9747496 -0.0083655 0.1317631 -310.8513519 1.10 1.59
CONF34 -23.0074018 1.22 -310.9727905 -0.0084222 0.1308813 -310.8503314 1.75 0.54
CONF35 -23.0073194 1.27 -310.9767663 -0.0074835 0.1328117 -310.8514381 1.05 1.74
CONF36 -23.0071492 1.38 -310.9751062 -0.0076260 0.1326829 -310.8500493 1.92 0.40
CONF38 -23.0070094 1.47 -310.9737012 -0.0082618 0.1313050 -310.8506580 1.54 0.76
CONF39 -23.0069911 1.48 -310.9736356 -0.0083128 0.1309967 -310.8509517 1.36 1.04
CONF40 -23.0069860 1.48 -310.9736982 -0.0082993 0.1312244 -310.8507731 1.47 0.86
CONF41 -23.0069668 1.49 -310.9739328 -0.0081040 0.1317320 -310.8503048 1.76 0.52
CONF42 -23.0069385 1.51 -310.9750954 -0.0081352 0.1319123 -310.8513183 1.13 1.53
CONF43 -23.0069145 1.53 -310.9743753 -0.0074285 0.1326379 -310.8491658 2.48 0.16
CONF44 -23.0068468 1.57 -310.9735814 -0.0083126 0.1308658 -310.8510282 1.31 1.12
CONF45 -23.0068432 1.57 -310.9730178 -0.0083314 0.1310382 -310.8503110 1.76 0.53
CONF46 -23.0068324 1.58 -310.9728206 -0.0082362 0.1300672 -310.8509896 1.33 1.08
CONF47 -23.0068240 1.58 -310.9750111 -0.0081979 0.1318381 -310.8513709 1.09 1.62
CONF48 -23.0068107 1.59 -310.9746133 -0.0077822 0.1327360 -310.8496595 2.17 0.26
CONF49 -23.0067519 1.63 -310.9748531 -0.0077532 0.1325014 -310.8501049 1.89 0.42
CONF50 -23.0067158 1.65 -310.9739287 -0.0079805 0.1320174 -310.8498918 2.02 0.34
CONF54 -23.0066490 1.69 -310.9730529 -0.0084644 0.1310357 -310.8504816 1.65 0.63
CONF55 -23.0066215 1.71 -310.9728914 -0.0083328 0.1317555 -310.8494688 2.29 0.22
CONF56 -23.0063187 1.90 -310.9738816 -0.0083829 0.1322211 -310.8500434 1.93 0.40
CONF57 -23.0062889 1.92 -310.9751917 -0.0080724 0.1318357 -310.8514284 1.06 1.72
CONF58 -23.0061728 1.99 -310.9769417 -0.0068709 0.1323369 -310.8514757 1.03 1.81
CONF59 -23.0061707 1.99 -310.9768786 -0.0068729 0.1322954 -310.8514561 1.04 1.77
CONF60 -23.0061585 2.00 -310.9767960 -0.0069148 0.1321850 -310.8515258 1.00 1.91
CONF61 -23.0061249 2.02 -310.9765424 -0.0070052 0.1318622 -310.8516854 0.90 2.26
CONF62 -23.0060951 2.04 -310.9763744 -0.0070689 0.1320360 -310.8514073 1.07 1.68
CONF63 -23.0059460 2.13 -310.9738060 -0.0078661 0.1322762 -310.8493960 2.33 0.20
CONF64 -23.0057295 2.27 -310.9754112 -0.0076605 0.1317044 -310.8513673 1.10 1.61
CONF65 -23.0057274 2.27 -310.9746075 -0.0084410 0.1314210 -310.8516274 0.93 2.12
CONF66 -23.0057248 2.27 -310.9746089 -0.0084406 0.1313966 -310.8516528 0.92 2.18
CONF67 -23.0057272 2.27 -310.9746088 -0.0084297 0.1314214 -310.8516172 0.94 2.10
CONF68 -23.0057203 2.27 -310.9745980 -0.0084359 0.1313049 -310.8517290 0.87 2.36
CONF69 -23.0056054 2.35 -310.9753119 -0.0078726 0.1317120 -310.8514726 1.03 1.80
CONF70 -23.0054560 2.44 -310.9740677 -0.0081421 0.1321974 -310.8500123 1.95 0.38
CONF71 -23.0054525 2.44 -310.9740465 -0.0082042 0.1321297 -310.8501210 1.88 0.43
CONF72 -23.0054513 2.44 -310.9740503 -0.0081331 0.1321793 -310.8500041 1.95 0.38
CONF73 -23.0054031 2.47 -310.9733414 -0.0084100 0.1306268 -310.8511246 1.25 1.25
CONF74 -23.0052804 2.55 -310.9738313 -0.0080078 0.1305721 -310.8512669 1.16 1.45
CONF75 -23.0052386 2.58 -310.9733849 -0.0083993 0.1312060 -310.8505781 1.59 0.70
CONF76 -23.0050016 2.73 -310.9734415 -0.0079706 0.1313424 -310.8500696 1.91 0.41
CONF77 -23.0047687 2.87 -310.9751204 -0.0073916 0.1323815 -310.8501305 1.87 0.43
CONF80 -23.0042429 3.20 -310.9736616 -0.0078527 0.1313849 -310.8501295 1.87 0.43
Number of conformers observed within the following ΔG windows:
ΔG [kcal/mol] #CONF sum(Boltzmann_weights)
---------------------------------------------
0 - 0.5 1 0.10
0 - 1.0 12 0.37
0 - 1.5 47 0.89
0 - 2.0 67 0.98
0 - 2.5 74 1.00
---------------------------------------------
Calculating ALPB_Gsolv values for evaluation of the std. dev. of Gsolv.
Constructed folders!
ALPB_GSOLV calculation was successful for CONF1/alpb_gsolv: -0.01285004
ALPB_GSOLV calculation was successful for CONF2/alpb_gsolv: -0.01280002
ALPB_GSOLV calculation was successful for CONF3/alpb_gsolv: -0.01277819
ALPB_GSOLV calculation was successful for CONF4/alpb_gsolv: -0.01274925
ALPB_GSOLV calculation was successful for CONF5/alpb_gsolv: -0.01313694
ALPB_GSOLV calculation was successful for CONF6/alpb_gsolv: -0.01328493
ALPB_GSOLV calculation was successful for CONF7/alpb_gsolv: -0.01318346
ALPB_GSOLV calculation was successful for CONF8/alpb_gsolv: -0.01263519
ALPB_GSOLV calculation was successful for CONF9/alpb_gsolv: -0.01330525
ALPB_GSOLV calculation was successful for CONF10/alpb_gsolv: -0.01277941
ALPB_GSOLV calculation was successful for CONF11/alpb_gsolv: -0.01328256
ALPB_GSOLV calculation was successful for CONF12/alpb_gsolv: -0.01336548
ALPB_GSOLV calculation was successful for CONF13/alpb_gsolv: -0.01336002
ALPB_GSOLV calculation was successful for CONF14/alpb_gsolv: -0.01328312
ALPB_GSOLV calculation was successful for CONF15/alpb_gsolv: -0.01326772
ALPB_GSOLV calculation was successful for CONF16/alpb_gsolv: -0.01329061
ALPB_GSOLV calculation was successful for CONF17/alpb_gsolv: -0.01327680
ALPB_GSOLV calculation was successful for CONF18/alpb_gsolv: -0.01297745
ALPB_GSOLV calculation was successful for CONF19/alpb_gsolv: -0.01304245
ALPB_GSOLV calculation was successful for CONF20/alpb_gsolv: -0.01292181
ALPB_GSOLV calculation was successful for CONF21/alpb_gsolv: -0.01328324
ALPB_GSOLV calculation was successful for CONF22/alpb_gsolv: -0.01337805
ALPB_GSOLV calculation was successful for CONF23/alpb_gsolv: -0.01329834
ALPB_GSOLV calculation was successful for CONF24/alpb_gsolv: -0.01335163
ALPB_GSOLV calculation was successful for CONF25/alpb_gsolv: -0.01322471
ALPB_GSOLV calculation was successful for CONF26/alpb_gsolv: -0.01309176
ALPB_GSOLV calculation was successful for CONF27/alpb_gsolv: -0.01340553
ALPB_GSOLV calculation was successful for CONF28/alpb_gsolv: -0.01306177
ALPB_GSOLV calculation was successful for CONF29/alpb_gsolv: -0.01305185
ALPB_GSOLV calculation was successful for CONF30/alpb_gsolv: -0.01326722
ALPB_GSOLV calculation was successful for CONF31/alpb_gsolv: -0.01327232
ALPB_GSOLV calculation was successful for CONF32/alpb_gsolv: -0.01322292
ALPB_GSOLV calculation was successful for CONF33/alpb_gsolv: -0.01335019
ALPB_GSOLV calculation was successful for CONF34/alpb_gsolv: -0.01334335
ALPB_GSOLV calculation was successful for CONF35/alpb_gsolv: -0.01271707
ALPB_GSOLV calculation was successful for CONF36/alpb_gsolv: -0.01306942
ALPB_GSOLV calculation was successful for CONF38/alpb_gsolv: -0.01333341
ALPB_GSOLV calculation was successful for CONF39/alpb_gsolv: -0.01333772
ALPB_GSOLV calculation was successful for CONF40/alpb_gsolv: -0.01334631
ALPB_GSOLV calculation was successful for CONF41/alpb_gsolv: -0.01326067
ALPB_GSOLV calculation was successful for CONF42/alpb_gsolv: -0.01327910
ALPB_GSOLV calculation was successful for CONF43/alpb_gsolv: -0.01293696
ALPB_GSOLV calculation was successful for CONF44/alpb_gsolv: -0.01333919
ALPB_GSOLV calculation was successful for CONF45/alpb_gsolv: -0.01333254
ALPB_GSOLV calculation was successful for CONF46/alpb_gsolv: -0.01330277
ALPB_GSOLV calculation was successful for CONF47/alpb_gsolv: -0.01330074
ALPB_GSOLV calculation was successful for CONF48/alpb_gsolv: -0.01287387
ALPB_GSOLV calculation was successful for CONF49/alpb_gsolv: -0.01316727
ALPB_GSOLV calculation was successful for CONF50/alpb_gsolv: -0.01319151
ALPB_GSOLV calculation was successful for CONF54/alpb_gsolv: -0.01328004
ALPB_GSOLV calculation was successful for CONF55/alpb_gsolv: -0.01327608
ALPB_GSOLV calculation was successful for CONF56/alpb_gsolv: -0.01324014
ALPB_GSOLV calculation was successful for CONF57/alpb_gsolv: -0.01315803
ALPB_GSOLV calculation was successful for CONF58/alpb_gsolv: -0.01284263
ALPB_GSOLV calculation was successful for CONF59/alpb_gsolv: -0.01286644
ALPB_GSOLV calculation was successful for CONF60/alpb_gsolv: -0.01292228
ALPB_GSOLV calculation was successful for CONF61/alpb_gsolv: -0.01300958
ALPB_GSOLV calculation was successful for CONF62/alpb_gsolv: -0.01306527
ALPB_GSOLV calculation was successful for CONF63/alpb_gsolv: -0.01308700
ALPB_GSOLV calculation was successful for CONF64/alpb_gsolv: -0.01328019
ALPB_GSOLV calculation was successful for CONF65/alpb_gsolv: -0.01336510
ALPB_GSOLV calculation was successful for CONF66/alpb_gsolv: -0.01336521
ALPB_GSOLV calculation was successful for CONF67/alpb_gsolv: -0.01336498
ALPB_GSOLV calculation was successful for CONF68/alpb_gsolv: -0.01337017
ALPB_GSOLV calculation was successful for CONF69/alpb_gsolv: -0.01332467
ALPB_GSOLV calculation was successful for CONF70/alpb_gsolv: -0.01303270
ALPB_GSOLV calculation was successful for CONF71/alpb_gsolv: -0.01308196
ALPB_GSOLV calculation was successful for CONF72/alpb_gsolv: -0.01302229
ALPB_GSOLV calculation was successful for CONF73/alpb_gsolv: -0.01337897
ALPB_GSOLV calculation was successful for CONF74/alpb_gsolv: -0.01332813
ALPB_GSOLV calculation was successful for CONF75/alpb_gsolv: -0.01336873
ALPB_GSOLV calculation was successful for CONF76/alpb_gsolv: -0.01328551
ALPB_GSOLV calculation was successful for CONF77/alpb_gsolv: -0.01304175
ALPB_GSOLV calculation was successful for CONF80/alpb_gsolv: -0.01315765
SD of solvation models @ 298.15 K (all units in kcal/mol):
CONFX ΔG(COSMO-RS) ΔG(DCOSMO-RS_gsolv) ΔG(ALPB_gsolv) SD(COSMO-RS 40%, DCOSMO-RS_gsolv 40%, ALPB_gsolv 20%)
----------------------------------------------------------------------------------------------------
CONF1 0.00 0.00 0.00 0.00
CONF2 0.04 0.08 0.03 0.02
CONF3 0.05 0.08 0.05 0.02
CONF4 0.54 0.35 0.06 0.22
CONF5 -0.23 -0.48 -0.18 0.16
CONF6 0.09 -0.39 -0.27 0.27
CONF7 -0.28 -0.48 -0.21 0.14
CONF8 0.30 0.30 0.13 0.08
CONF9 -0.13 -0.45 -0.29 0.17
CONF10 0.01 -0.01 0.04 0.03
CONF11 -0.11 -0.42 -0.27 0.17
CONF12 -0.32 -0.68 -0.32 0.21
CONF13 -0.35 -0.70 -0.32 0.22
CONF14 -0.08 -0.43 -0.27 0.19
CONF15 -0.13 -0.45 -0.26 0.18
CONF16 -0.10 -0.42 -0.28 0.17
CONF17 -0.12 -0.44 -0.27 0.18
CONF18 -0.22 -0.42 -0.08 0.16
CONF19 -0.22 -0.44 -0.12 0.16
CONF20 0.46 0.18 -0.05 0.24
CONF21 0.12 -0.41 -0.27 0.30
CONF22 -0.33 -0.70 -0.33 0.22
CONF23 -0.12 -0.43 -0.28 0.17
CONF24 -0.37 -0.71 -0.31 0.22
CONF25 -0.07 -0.40 -0.24 0.18
CONF26 -0.20 -0.47 -0.15 0.17
CONF27 -0.39 -0.71 -0.35 0.20
CONF28 -0.27 -0.48 -0.13 0.16
CONF29 -0.28 -0.47 -0.13 0.16
CONF30 -0.18 -0.48 -0.26 0.17
CONF31 -0.08 -0.50 -0.26 0.23
CONF32 -0.04 -0.47 -0.23 0.23
CONF33 -0.32 -0.68 -0.31 0.22
CONF34 -0.36 -0.70 -0.31 0.21
CONF35 0.23 0.26 0.08 0.08
CONF36 0.14 -0.11 -0.14 0.15
CONF38 -0.26 -0.53 -0.30 0.15
CONF39 -0.29 -0.54 -0.31 0.15
CONF40 -0.28 -0.55 -0.31 0.16
CONF41 -0.16 -0.45 -0.26 0.16
CONF42 -0.18 -0.48 -0.27 0.17
CONF43 0.26 0.28 -0.05 0.16
CONF44 -0.29 -0.54 -0.31 0.15
CONF45 -0.30 -0.54 -0.30 0.14
CONF46 -0.24 -0.51 -0.28 0.15
CONF47 -0.22 -0.51 -0.28 0.17
CONF48 0.04 -0.20 -0.01 0.14
CONF49 0.06 -0.20 -0.20 0.16
CONF50 -0.08 -0.33 -0.21 0.14
CONF54 -0.39 -0.66 -0.27 0.19
CONF55 -0.30 -0.63 -0.27 0.20
CONF56 -0.34 -0.61 -0.24 0.18
CONF57 -0.14 -0.45 -0.19 0.18
CONF58 0.61 0.39 0.00 0.27
CONF59 0.61 0.37 -0.01 0.28
CONF60 0.59 0.33 -0.05 0.28
CONF61 0.53 0.24 -0.10 0.29
CONF62 0.49 0.16 -0.14 0.29
CONF63 -0.01 -0.34 -0.15 0.18
CONF64 0.12 -0.36 -0.27 0.27
CONF65 -0.37 -0.72 -0.32 0.22
CONF66 -0.37 -0.72 -0.32 0.22
CONF67 -0.36 -0.72 -0.32 0.22
CONF68 -0.37 -0.72 -0.33 0.22
CONF69 -0.02 -0.39 -0.30 0.21
CONF70 -0.18 -0.48 -0.11 0.19
CONF71 -0.22 -0.51 -0.15 0.19
CONF72 -0.18 -0.47 -0.11 0.19
CONF73 -0.35 -0.70 -0.33 0.21
CONF74 -0.10 -0.41 -0.30 0.17
CONF75 -0.35 -0.69 -0.33 0.21
CONF76 -0.08 -0.42 -0.27 0.19
CONF77 0.29 -0.03 -0.12 0.21
CONF80 -0.00 -0.29 -0.19 0.16
----------------------------------------------------------------------------------------------------
Calculating Boltzmann averaged free energy of ensemble!
temperature /K: avE(T) /a.u. avGmRRHO(T) /a.u. avGsolv(T) /a.u. avG(T) /a.u.
----------------------------------------------------------------------------------------------------
273.15 -310.9756033 0.1355882 -0.0091311 -310.8491461
278.15 -310.9755795 0.1348675 -0.0088841 -310.8495962
283.15 -310.9755569 0.1341424 -0.0086406 -310.8500551
288.15 -310.9755353 0.1334132 -0.0084008 -310.8505229
293.15 -310.9755150 0.1326797 -0.0081645 -310.8509998
298.15 -310.9754955 0.1319419 -0.0079319 -310.8514855 <<==part2==
303.15 -310.9754769 0.1312001 -0.0077029 -310.8519798
308.15 -310.9754593 0.1304538 -0.0074775 -310.8524829
313.15 -310.9754424 0.1297036 -0.0072557 -310.8529945
318.15 -310.9754261 0.1289490 -0.0070375 -310.8535146
323.15 -310.9754107 0.1281902 -0.0068228 -310.8540433
328.15 -310.9753957 0.1274272 -0.0066117 -310.8545801
333.15 -310.9753812 0.1266601 -0.0064040 -310.8551251
338.15 -310.9753673 0.1258888 -0.0061997 -310.8556783
343.15 -310.9753540 0.1251132 -0.0059988 -310.8562396
348.15 -310.9753408 0.1243334 -0.0058013 -310.8568087
353.15 -310.9753284 0.1235495 -0.0056070 -310.8573859
358.15 -310.9753162 0.1227614 -0.0054158 -310.8579706
363.15 -310.9753043 0.1219691 -0.0052278 -310.8585630
368.15 -310.9752929 0.1211727 -0.0050428 -310.8591631
373.15 -310.9752817 0.1203720 -0.0048609 -310.8597706
----------------------------------------------------------------------------------------------------
--------------------------------------------------
Conformers considered further
--------------------------------------------------
Conformers that are below the Boltzmann threshold G_thr(2) of 95.0%:
CONF1, CONF3, CONF2, CONF68, CONF61, CONF66, CONF65, CONF67, CONF13, CONF24, CONF20
CONF60, CONF58, CONF69, CONF59, CONF9, CONF7, CONF17, CONF12, CONF35, CONF57, CONF62
CONF8, CONF15, CONF47, CONF5, CONF64, CONF18, CONF33, CONF30, CONF14, CONF19, CONF42
CONF4, CONF11, CONF74, CONF31, CONF32, CONF73, CONF16, CONF44, CONF6, CONF46, CONF39
CONF22, CONF27, CONF40, CONF38, CONF75, CONF54, CONF34, CONF45, CONF23, CONF41, CONF10
CONF77, CONF80, CONF71
Backing up enso_ensemble_part2.xyz to enso_ensemble_part2.xyz.1.
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>END of Part2<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
Ran part2 in 0.2050 seconds
----------------------------------------------------------------------------------------------------
NMR MODE - PART4
----------------------------------------------------------------------------------------------------
calculate coupling constants: on
prog4J - program for coupling constant calculation: tm
funcJ - functional for coupling constant calculation: pbe0
basisJ - basis for coupling constant calculation: def2-TZVP
sm4J - solvent model for the coupling calculation: dcosmors
calculate shielding constants σ: on
prog4S - program for shielding constant calculation: tm
funcS - functional for shielding constant calculation: pbe0
basisS - basis for shielding constant calculation: def2-TZVP
sm4S - solvent model for the shielding calculation: dcosmors
Calculating proton spectrum: on
reference for 1H: TMS
spectrometer frequency: 300.0
Considering the following 58 conformers:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF10, CONF11
CONF12, CONF13, CONF14, CONF15, CONF16, CONF17, CONF18, CONF19, CONF20, CONF22, CONF23
CONF24, CONF27, CONF30, CONF31, CONF32, CONF33, CONF34, CONF35, CONF38, CONF39, CONF40
CONF41, CONF42, CONF44, CONF45, CONF46, CONF47, CONF54, CONF57, CONF58, CONF59, CONF60
CONF61, CONF62, CONF64, CONF65, CONF66, CONF67, CONF68, CONF69, CONF71, CONF73, CONF74
CONF75, CONF77, CONF80
--------------------------------------------------
* Gibbs free energies used in part4 *
--------------------------------------------------
CONF# E [Eh] Gsolv [Eh] GmRRHO [Eh] Gtot ΔGtot Boltzmannweight
r2scan-3c COSMO-RS-normal GFN2 [Eh] [kcal/mol] % at 298.15 K
[r2scan-3c] [alpb]-bhess
CONF1 -310.9779421 -0.0078485 0.1326780 -310.8531126 0.00 10.81 <------
CONF2 -310.9766733 -0.0077870 0.1322821 -310.8521781 0.59 4.02
CONF3 -310.9767072 -0.0077611 0.1322874 -310.8521809 0.58 4.03
CONF4 -310.9769910 -0.0069825 0.1326644 -310.8513091 1.13 1.60
CONF5 -310.9752433 -0.0082178 0.1320914 -310.8513697 1.09 1.71
CONF6 -310.9755626 -0.0077038 0.1322484 -310.8510180 1.31 1.18
CONF7 -310.9752725 -0.0082944 0.1321195 -310.8514474 1.04 1.85
CONF8 -310.9769749 -0.0073678 0.1329484 -310.8513943 1.08 1.75
CONF9 -310.9754631 -0.0080528 0.1320613 -310.8514546 1.04 1.87
CONF10 -310.9754106 -0.0078398 0.1330596 -310.8501908 1.83 0.49
CONF11 -310.9753819 -0.0080269 0.1321196 -310.8512892 1.14 1.57
CONF12 -310.9749503 -0.0083661 0.1318717 -310.8514447 1.05 1.85
CONF13 -310.9749185 -0.0084110 0.1317452 -310.8515843 0.96 2.14
CONF14 -310.9754401 -0.0079817 0.1321025 -310.8513194 1.13 1.62
CONF15 -310.9753877 -0.0080490 0.1320438 -310.8513929 1.08 1.75
CONF16 -310.9740923 -0.0080047 0.1309977 -310.8510993 1.26 1.28
CONF17 -310.9754046 -0.0080391 0.1319964 -310.8514473 1.04 1.85
CONF18 -310.9753558 -0.0082018 0.1321958 -310.8513618 1.10 1.69
CONF19 -310.9752809 -0.0082043 0.1321658 -310.8513194 1.13 1.62
CONF20 -310.9766845 -0.0071146 0.1322468 -310.8515524 0.98 2.07
CONF22 -310.9738935 -0.0083778 0.1313361 -310.8509352 1.37 1.08
CONF23 -310.9741656 -0.0080352 0.1318949 -310.8503059 1.76 0.55
CONF24 -310.9748194 -0.0084452 0.1316841 -310.8515806 0.96 2.13
CONF27 -310.9735485 -0.0084651 0.1311659 -310.8508477 1.42 0.98
CONF30 -310.9751637 -0.0081326 0.1319458 -310.8513505 1.11 1.67
CONF31 -310.9751571 -0.0079707 0.1319056 -310.8512222 1.19 1.46
CONF32 -310.9751719 -0.0079108 0.1318699 -310.8512127 1.19 1.45
CONF33 -310.9747496 -0.0083655 0.1317631 -310.8513519 1.10 1.68
CONF34 -310.9727905 -0.0084222 0.1308813 -310.8503314 1.75 0.57
CONF35 -310.9767663 -0.0074835 0.1328117 -310.8514381 1.05 1.84
CONF38 -310.9737012 -0.0082618 0.1313050 -310.8506580 1.54 0.80
CONF39 -310.9736356 -0.0083128 0.1309967 -310.8509517 1.36 1.10
CONF40 -310.9736982 -0.0082993 0.1312244 -310.8507731 1.47 0.91
CONF41 -310.9739328 -0.0081040 0.1317320 -310.8503048 1.76 0.55
CONF42 -310.9750954 -0.0081352 0.1319123 -310.8513183 1.13 1.62
CONF44 -310.9735814 -0.0083126 0.1308658 -310.8510282 1.31 1.19
CONF45 -310.9730178 -0.0083314 0.1310382 -310.8503110 1.76 0.56
CONF46 -310.9728206 -0.0082362 0.1300672 -310.8509896 1.33 1.14
CONF47 -310.9750111 -0.0081979 0.1318381 -310.8513709 1.09 1.71
CONF54 -310.9730529 -0.0084644 0.1310357 -310.8504816 1.65 0.67
CONF57 -310.9751917 -0.0080724 0.1318357 -310.8514284 1.06 1.82
CONF58 -310.9769417 -0.0068709 0.1323369 -310.8514757 1.03 1.91
CONF59 -310.9768786 -0.0068729 0.1322954 -310.8514561 1.04 1.87
CONF60 -310.9767960 -0.0069148 0.1321850 -310.8515258 1.00 2.01
CONF61 -310.9765424 -0.0070052 0.1318622 -310.8516854 0.90 2.38
CONF62 -310.9763744 -0.0070689 0.1320360 -310.8514073 1.07 1.78
CONF64 -310.9754112 -0.0076605 0.1317044 -310.8513673 1.10 1.70
CONF65 -310.9746075 -0.0084410 0.1314210 -310.8516274 0.93 2.24
CONF66 -310.9746089 -0.0084406 0.1313966 -310.8516528 0.92 2.30
CONF67 -310.9746088 -0.0084297 0.1314214 -310.8516172 0.94 2.22
CONF68 -310.9745980 -0.0084359 0.1313049 -310.8517290 0.87 2.50
CONF69 -310.9753119 -0.0078726 0.1317120 -310.8514726 1.03 1.90
CONF71 -310.9740465 -0.0082042 0.1321297 -310.8501210 1.88 0.45
CONF73 -310.9733414 -0.0084100 0.1306268 -310.8511246 1.25 1.32
CONF74 -310.9738313 -0.0080078 0.1305721 -310.8512669 1.16 1.53
CONF75 -310.9733849 -0.0083993 0.1312060 -310.8505781 1.59 0.74
CONF77 -310.9751204 -0.0073916 0.1323815 -310.8501305 1.87 0.46
CONF80 -310.9736616 -0.0078527 0.1313849 -310.8501295 1.87 0.46
Conformers that are below the Boltzmann-thr of 95.0:
CONF1, CONF3, CONF2, CONF68, CONF61, CONF66, CONF65, CONF67, CONF13, CONF24, CONF20
CONF60, CONF58, CONF69, CONF59, CONF9, CONF7, CONF17, CONF12, CONF35, CONF57, CONF62
CONF8, CONF15, CONF47, CONF5, CONF64, CONF18, CONF33, CONF30, CONF14, CONF19, CONF42
CONF4, CONF11, CONF74, CONF31, CONF32, CONF73, CONF16, CONF44, CONF6, CONF46, CONF39
CONF22, CONF27, CONF40, CONF38, CONF75, CONF54, CONF34, CONF45, CONF23, CONF41, CONF10
CONF77, CONF80, CONF71
Constructed folders!
Performing coupling constant calculations:
Coupling constant calculation was successful for CONF1/NMR
Coupling constant calculation was successful for CONF3/NMR
Coupling constant calculation was successful for CONF2/NMR
Coupling constant calculation was successful for CONF68/NMR
Coupling constant calculation was successful for CONF61/NMR
Coupling constant calculation was successful for CONF66/NMR
Coupling constant calculation was successful for CONF65/NMR
Coupling constant calculation was successful for CONF67/NMR
Coupling constant calculation was successful for CONF13/NMR
Coupling constant calculation was successful for CONF24/NMR
Coupling constant calculation was successful for CONF20/NMR
Coupling constant calculation was successful for CONF60/NMR
Coupling constant calculation was successful for CONF58/NMR
Coupling constant calculation was successful for CONF69/NMR
Coupling constant calculation was successful for CONF59/NMR
Coupling constant calculation was successful for CONF9/NMR
Coupling constant calculation was successful for CONF7/NMR
Coupling constant calculation was successful for CONF17/NMR
Coupling constant calculation was successful for CONF12/NMR
Coupling constant calculation was successful for CONF35/NMR
Coupling constant calculation was successful for CONF57/NMR
Coupling constant calculation was successful for CONF62/NMR
Coupling constant calculation was successful for CONF8/NMR
Coupling constant calculation was successful for CONF15/NMR
Coupling constant calculation was successful for CONF47/NMR
Coupling constant calculation was successful for CONF5/NMR
Coupling constant calculation was successful for CONF64/NMR
Coupling constant calculation was successful for CONF18/NMR
Coupling constant calculation was successful for CONF33/NMR
Coupling constant calculation was successful for CONF30/NMR
Coupling constant calculation was successful for CONF14/NMR
Coupling constant calculation was successful for CONF19/NMR
Coupling constant calculation was successful for CONF42/NMR
Coupling constant calculation was successful for CONF4/NMR
Coupling constant calculation was successful for CONF11/NMR
Coupling constant calculation was successful for CONF74/NMR
Coupling constant calculation was successful for CONF31/NMR
Coupling constant calculation was successful for CONF32/NMR
Coupling constant calculation was successful for CONF73/NMR
Coupling constant calculation was successful for CONF16/NMR
Coupling constant calculation was successful for CONF44/NMR
Coupling constant calculation was successful for CONF6/NMR
Coupling constant calculation was successful for CONF46/NMR
Coupling constant calculation was successful for CONF39/NMR
Coupling constant calculation was successful for CONF22/NMR
Coupling constant calculation was successful for CONF27/NMR
Coupling constant calculation was successful for CONF40/NMR
Coupling constant calculation was successful for CONF38/NMR
Coupling constant calculation was successful for CONF75/NMR
Coupling constant calculation was successful for CONF54/NMR
Coupling constant calculation was successful for CONF34/NMR
Coupling constant calculation was successful for CONF45/NMR
Coupling constant calculation was successful for CONF23/NMR
Coupling constant calculation was successful for CONF41/NMR
Coupling constant calculation was successful for CONF10/NMR
Coupling constant calculation was successful for CONF77/NMR
Coupling constant calculation was successful for CONF80/NMR
Coupling constant calculation was successful for CONF71/NMR
Performing shielding constant calculations:
Shielding constant calculation was successful for CONF1/NMR
Shielding constant calculation was successful for CONF3/NMR
Shielding constant calculation was successful for CONF2/NMR
Shielding constant calculation was successful for CONF68/NMR
Shielding constant calculation was successful for CONF61/NMR
Shielding constant calculation was successful for CONF66/NMR
Shielding constant calculation was successful for CONF65/NMR
Shielding constant calculation was successful for CONF67/NMR
Shielding constant calculation was successful for CONF13/NMR
Shielding constant calculation was successful for CONF24/NMR
Shielding constant calculation was successful for CONF20/NMR
Shielding constant calculation was successful for CONF60/NMR
Shielding constant calculation was successful for CONF58/NMR
Shielding constant calculation was successful for CONF69/NMR
Shielding constant calculation was successful for CONF59/NMR
Shielding constant calculation was successful for CONF9/NMR
Shielding constant calculation was successful for CONF7/NMR
Shielding constant calculation was successful for CONF17/NMR
Shielding constant calculation was successful for CONF12/NMR
Shielding constant calculation was successful for CONF35/NMR
Shielding constant calculation was successful for CONF57/NMR
Shielding constant calculation was successful for CONF62/NMR
Shielding constant calculation was successful for CONF8/NMR
Shielding constant calculation was successful for CONF15/NMR
Shielding constant calculation was successful for CONF47/NMR
Shielding constant calculation was successful for CONF5/NMR
Shielding constant calculation was successful for CONF64/NMR
Shielding constant calculation was successful for CONF18/NMR
Shielding constant calculation was successful for CONF33/NMR
Shielding constant calculation was successful for CONF30/NMR
Shielding constant calculation was successful for CONF14/NMR
Shielding constant calculation was successful for CONF19/NMR
Shielding constant calculation was successful for CONF42/NMR
Shielding constant calculation was successful for CONF4/NMR
Shielding constant calculation was successful for CONF11/NMR
Shielding constant calculation was successful for CONF74/NMR
Shielding constant calculation was successful for CONF31/NMR
Shielding constant calculation was successful for CONF32/NMR
Shielding constant calculation was successful for CONF73/NMR
Shielding constant calculation was successful for CONF16/NMR
Shielding constant calculation was successful for CONF44/NMR
Shielding constant calculation was successful for CONF6/NMR
Shielding constant calculation was successful for CONF46/NMR
Shielding constant calculation was successful for CONF39/NMR
Shielding constant calculation was successful for CONF22/NMR
Shielding constant calculation was successful for CONF27/NMR
Shielding constant calculation was successful for CONF40/NMR
Shielding constant calculation was successful for CONF38/NMR
Shielding constant calculation was successful for CONF75/NMR
Shielding constant calculation was successful for CONF54/NMR
Shielding constant calculation was successful for CONF34/NMR
Shielding constant calculation was successful for CONF45/NMR
Shielding constant calculation was successful for CONF23/NMR
Shielding constant calculation was successful for CONF41/NMR
Shielding constant calculation was successful for CONF10/NMR
Shielding constant calculation was successful for CONF77/NMR
Shielding constant calculation was successful for CONF80/NMR
Shielding constant calculation was successful for CONF71/NMR
Generating file anmr_enso for processing with the ANMR program.
Writing .anmrrc!
Generating plain nmrprop.dat files for each populated conformer.
These files contain all calculated shielding and coupling constants.
The files can be read by ANMR using the keyword '-plain'.
Tasks completed!
Averaged shielding constants:
# in coord element σ(sigma) SD(σ based on SD Gsolv) SD(σ by 0.4 kcal/mol) shift σ_ref
---------------------------------------------------------------------------------------------------------
5 h 29.34 0.006872 0.017630 2.29 31.636
6 h 29.34 0.006872 0.017630 2.29 31.636
8 h 29.99 0.003893 0.015362 1.64 31.636
10 h 27.83 0.003181 0.008456 3.80 31.636
12 h 30.67 0.018049 0.103016 0.96 31.636
13 h 27.83 0.003181 0.008456 3.80 31.636
14 h 29.99 0.003893 0.015362 1.64 31.636
15 h 25.72 0.006823 0.027844 5.91 31.636
16 h 25.63 0.003137 0.008056 6.01 31.636
17 h 29.88 0.001245 0.008133 1.75 31.636
18 h 29.88 0.001245 0.008133 1.75 31.636
19 h 29.88 0.001245 0.008133 1.75 31.636
---------------------------------------------------------------------------------------------------------
# in coord element σ(sigma) min(σ)* CONFX max(σ)* CONFX Δ(max-min)
---------------------------------------------------------------------------------------------------------
5 h 29.34 29.19 CONF20 29.63 CONF34 0.44
6 h 29.34 29.19 CONF20 29.63 CONF34 0.44
8 h 29.99 29.82 CONF2 30.19 CONF11 0.38
10 h 27.83 27.58 CONF80 28.03 CONF5 0.44
12 h 30.67 29.45 CONF1 31.64 CONF20 2.19
13 h 27.83 27.58 CONF80 28.03 CONF5 0.44
14 h 29.99 29.82 CONF2 30.19 CONF11 0.38
15 h 25.72 25.27 CONF10 26.00 CONF62 0.73
16 h 25.63 25.40 CONF10 25.71 CONF14 0.31
17 h 29.88 29.78 CONF1 29.99 CONF34 0.21
18 h 29.88 29.78 CONF1 29.99 CONF34 0.21
19 h 29.88 29.78 CONF1 29.99 CONF34 0.21
---------------------------------------------------------------------------------------------------------
* min(σ) and max(σ) are averaged over the chemical equivalent atoms, but not Boltzmann weighted.
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>END of Part4<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
Ran part4 in 4.4708 seconds
Part : #conf time time (including restarts)
-----------------------------------------------------------------------
Input : 86 - -
Part0_all : 83 0.19 s 49.26 s
Part1_initial_sort : 74 0.00 s 299.03 s
Part1_all : 74 0.22 s 312.69 s
Part2_opt : 74 0.00 s 3663.38 s
Part2_all : 58 0.20 s 3955.92 s
Part4 : 58 4.47 s 4404.81 s
-----------------------------------------------------------------------
All parts : - 5.08 s 8722.68 s
CENSO all done!
$CENSO global configuration file: .censorc
$VERSION:1.1.2
ORCA: /home/$USER/orca_5_0_1_linux_x86-64_openmpi411
ORCA version: 5.0.1
GFN-xTB: /home/$USER/bin/xtb
CREST: /home/$USER/bin/crest
mpshift: /home/$USER/TURBOMOLE.7.5/bin/em64t-unknown-linux-gnu/mpshift
escf: /home/$USER/TURBOMOLE.7.5/bin/em64t-unknown-linux-gnu/escf
#COSMO-RS
ctd = BP_TZVP_C30_1601.ctd cdir = "/home/$USER/COSMOthermX16/COSMOtherm/CTDATA-FILES" ldir = "/home/$USER/COSMOthermX16/COSMOtherm/CTDATA-FILES"
$ENDPROGRAMS
$CRE SORTING SETTINGS:
$GENERAL SETTINGS:
nconf: all # ['all', 'number e.g. 10 up to all conformers']
charge: 0 # ['number e.g. 0']
unpaired: 0 # ['number e.g. 0']
solvent: gas # ['gas', 'acetone', 'acetonitrile', 'aniline', 'benzaldehyde', 'benzene', 'ccl4', '...']
prog_rrho: xtb # ['xtb']
temperature: 298.15 # ['temperature in K e.g. 298.15']
trange: [273.15, 378.15, 5] # ['temperature range [start, end, step]']
multitemp: on # ['on', 'off']
evaluate_rrho: on # ['on', 'off']
consider_sym: on # ['on', 'off']
bhess: on # ['on', 'off']
imagthr: automatic # ['automatic or e.g., -100 # in cm-1']
sthr: automatic # ['automatic or e.g., 50 # in cm-1']
scale: automatic # ['automatic or e.g., 1.0 ']
rmsdbias: off # ['on', 'off']
sm_rrho: alpb # ['alpb', 'gbsa']
progress: off # possibilities
check: on # ['on', 'off']
prog: tm # ['tm', 'orca']
func: r2scan-3c # ['b3-lyp', 'b3lyp', 'b3lyp-3c', 'b3lyp-d3', 'b3lyp-d3(0)', 'b3lyp-d4', 'b3lyp-nl', '...']
basis: automatic # ['automatic', 'def2-TZVP', 'def2-mSVP', 'def2-mSVP', 'def2-mSVP', 'def2-mSVP', '...']
maxthreads: 7 # ['number of threads e.g. 2']
omp: 4 # ['number cores per thread e.g. 4']
balance: off # possibilities
cosmorsparam: automatic # ['automatic', '12-fine', '12-normal', '13-fine', '13-normal', '14-fine', '...']
$PART0 - CHEAP-PRESCREENING - SETTINGS:
part0: on # ['on', 'off']
func0: b97-d # ['b3-lyp', 'b3lyp', 'b3lyp-3c', 'b3lyp-d3', 'b3lyp-d3(0)', 'b3lyp-d4', '...']
basis0: def2-SV(P) # ['automatic', 'def2-SV(P)', 'def2-TZVP', 'def2-mSVP', 'def2-mSVP', 'def2-mSVP', '...']
part0_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
part0_threshold: 4.0 # ['number e.g. 4.0']
$PART1 - PRESCREENING - SETTINGS:
# func and basis is set under GENERAL SETTINGS
part1: on # ['on', 'off']
smgsolv1: cosmors # ['alpb_gsolv', 'cosmo', 'cosmors', 'cosmors-fine', 'cpcm', 'dcosmors', '...']
part1_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
part1_threshold: 3.5 # ['number e.g. 5.0']
$PART2 - OPTIMIZATION - SETTINGS:
# func and basis is set under GENERAL SETTINGS
part2: on # ['on', 'off']
opt_limit: 2.5 # ['number e.g. 4.0']
sm2: default # ['cosmo', 'cpcm', 'dcosmors', 'default', 'smd']
smgsolv2: cosmors # ['alpb_gsolv', 'cosmo', 'cosmors', 'cosmors-fine', 'cpcm', 'dcosmors', '...']
part2_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
ancopt: on # ['on']
hlow: 0.01 # ['lowest force constant in ANC generation, e.g. 0.01']
opt_spearman: on # ['on', 'off']
part2_threshold: 99 # ['Boltzmann sum threshold in %. e.g. 95 (between 1 and 100)']
optlevel2: automatic # ['crude', 'sloppy', 'loose', 'lax', 'normal', 'tight', 'vtight', 'extreme', '...']
optcycles: 8 # ['number e.g. 5 or 10']
spearmanthr: -4.0 # ['value between -1 and 1, if outside set automatically']
radsize: 10 # ['number e.g. 8 or 10']
crestcheck: off # ['on', 'off']
$PART3 - REFINEMENT - SETTINGS:
part3: off # ['on', 'off']
prog3: prog # ['tm', 'orca', 'prog']
func3: pw6b95 # ['b3-lyp', 'b3lyp', 'b3lyp-3c', 'b3lyp-d3', 'b3lyp-d3(0)', 'b3lyp-d4', 'b3lyp-nl', '...']
basis3: def2-TZVPD # ['DZ', 'QZV', 'QZVP', 'QZVPP', 'SV(P)', 'SVP', 'TZVP', 'TZVPP', 'aug-cc-pV5Z', '...']
smgsolv3: cosmors # ['alpb_gsolv', 'cosmo', 'cosmors', 'cosmors-fine', 'cpcm', 'dcosmors', '...']
part3_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
part3_threshold: 99 # ['Boltzmann sum threshold in %. e.g. 95 (between 1 and 100)']
$NMR PROPERTY SETTINGS:
$PART4 SETTINGS:
part4: off # ['on', 'off']
couplings: on # ['on', 'off']
progJ: prog # ['tm', 'orca', 'adf', 'prog']
funcJ: pbe0 # ['b3-lyp', 'b3lyp', 'b3lyp-3c', 'b3lyp-d3', 'b3lyp-d3(0)', 'b3lyp-d4', 'b3lyp-nl', '...']
basisJ: def2-TZVP # ['DZ', 'QZV', 'QZVP', 'QZVPP', 'SV(P)', 'SVP', 'TZVP', 'TZVPP', 'aug-cc-pV5Z', '...']
sm4J: default # ['cosmo', 'cpcm', 'dcosmors', 'smd']
shieldings: on # ['on', 'off']
progS: prog # ['tm', 'orca', 'adf', 'prog']
funcS: pbe0 # ['b3-lyp', 'b3lyp', 'b3lyp-3c', 'b3lyp-d3', 'b3lyp-d3(0)', 'b3lyp-d4', 'b3lyp-nl', '...']
basisS: def2-TZVP # ['DZ', 'QZV', 'QZVP', 'QZVPP', 'SV(P)', 'SVP', 'TZVP', 'TZVPP', 'aug-cc-pV5Z', '...']
sm4S: default # ['cosmo', 'cpcm', 'dcosmors', 'smd']
reference_1H: TMS # ['TMS']
reference_13C: TMS # ['TMS']
reference_19F: CFCl3 # ['CFCl3']
reference_29Si: TMS # ['TMS']
reference_31P: TMP # ['TMP', 'PH3']
1H_active: on # ['on', 'off']
13C_active: on # ['on', 'off']
19F_active: off # ['on', 'off']
29Si_active: off # ['on', 'off']
31P_active: off # ['on', 'off']
resonance_frequency: 300.0 # ['MHz number of your experimental spectrometer setup']
$OPTICAL ROTATION PROPERTY SETTINGS:
$PART5 SETTINGS:
optical_rotation: off # ['on', 'off']
funcOR: pbe # ['functional for opt_rot e.g. pbe']
funcOR_SCF: r2scan-3c # ['functional for SCF in opt_rot e.g. r2scan-3c']
basisOR: def2-SVPD # ['basis set for opt_rot e.g. def2-SVPD']
frequency_optical_rot: [589.0] # ['list of freq in nm to evaluate opt rot at e.g. [589, 700]']
$END CENSORC
Now all information is present and ANMR can be called to calculate the full
NMR spectrum.
$ anmr --plain --mf 400 > anmr.out 2> error.anmr &
+--------------------------------------+
| A N M R |
| S. Grimme |
| Universitaet Bonn, MCTC |
| 1989-2019 |
| version 3.5.1 |
| Sat Feb 9 06:41:57 CET 2019 |
+--------------------------------------+
Based on a TurboPascal program written
in 1989 which was translated to F90 in
2005 and re-activated in 2017.
Please cite work employing this code as:
ANMR Ver. 3.5: An automatic, QC based
coupled NMR spectra simulation program.
S. Grimme, Universitaet Bonn, 2019
S. Grimme, C. Bannwarth, S. Dohm, A. Hansen
J. Pisarek, P. Pracht, J. Seibert, F. Neese
Angew. Chem. Int. Ed. 2017, 56, 14763-14769.
DOI:10.1002/anie.201708266
=============================
# OMP threads = 4
=============================
reading <.anmrrc> for standard data
1 31.6360000000000 0.000000000000000E+000 1
1H resonance frequency (-mf <real>) : 400.00
line width (-lw <real>) : 1.00
number of all conformers :58
temperature in K : 298.15
remove J couplings to OH groups : T
maximum spin system size in a fragment :14
fragmentation type (0=none,1=at,2=mol) : 2
chemical shift scalings a,b : 1.00 0.00
spin-spin coupling scal factor : 1.07
plot offset : 0.00
Active nuclei :H
reading from anmr_enso
conformational energies: 58
0.000 0.775 0.796 2.098 0.878 2.092 2.092 2.092 1.897 1.960
0.789 0.719 0.628 1.650 0.667 1.556 1.675 1.592 1.877 0.738
1.726 0.984 0.607 1.603 1.839 1.694 1.588 1.623 2.003 1.743
1.570 1.670 1.786 0.597 1.607 2.580 1.748 1.738 2.887 2.416
2.736 1.493 3.214 2.702 2.541 2.757 2.663 2.661 2.860 3.068
3.233 3.090 2.370 2.516 1.589 1.771 2.686 2.445
conformational RRHO energies:
0.000 -0.245 -0.248 -0.862 -0.512 -0.804 -0.789 -0.789 -0.585 -0.624
-0.271 -0.309 -0.214 -0.606 -0.240 -0.387 -0.350 -0.428 -0.506 0.084
-0.529 -0.403 0.170 -0.398 -0.527 -0.368 -0.611 -0.303 -0.574 -0.459
-0.361 -0.321 -0.480 -0.009 -0.350 -1.321 -0.485 -0.507 -1.287 -1.054
-1.137 -0.270 -1.638 -1.055 -0.842 -0.949 -0.912 -0.862 -0.924 -1.031
-1.127 -1.029 -0.491 -0.594 0.239 -0.186 -0.811 -0.344
conformational Gsolv free energies:
0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000
0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000
0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000
0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000
0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000
0.000 0.000 0.000 0.000 0.000 0.000 0.000 0.000
conformational free energies:
0.000 0.530 0.548 1.237 0.366 1.288 1.304 1.303 1.312 1.336
0.519 0.410 0.414 1.044 0.427 1.169 1.325 1.165 1.371 0.822
1.197 0.581 0.777 1.205 1.312 1.325 0.977 1.320 1.429 1.284
1.209 1.349 1.306 0.588 1.256 1.258 1.263 1.231 1.600 1.361
1.599 1.224 1.575 1.647 1.698 1.808 1.751 1.800 1.936 2.037
2.105 2.061 1.878 1.922 1.828 1.585 1.875 2.100
gi per conformer:
1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00
1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00
1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00
1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00
1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00
1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00
conformational populations calculated by anmr:
0.103 0.042 0.041 0.013 0.055 0.012 0.011 0.011 0.011 0.011
0.043 0.051 0.051 0.018 0.050 0.014 0.011 0.014 0.010 0.026
0.014 0.038 0.028 0.013 0.011 0.011 0.020 0.011 0.009 0.012
0.013 0.011 0.011 0.038 0.012 0.012 0.012 0.013 0.007 0.010
0.007 0.013 0.007 0.006 0.006 0.005 0.005 0.005 0.004 0.003
0.003 0.003 0.004 0.004 0.005 0.007 0.004 0.003
conformational populations calculated by enso:
These are used:
0.108 0.040 0.040 0.025 0.024 0.023 0.022 0.022 0.021 0.021
0.021 0.020 0.019 0.019 0.019 0.019 0.018 0.018 0.018 0.018
0.018 0.018 0.018 0.018 0.017 0.017 0.017 0.017 0.017 0.017
0.016 0.016 0.016 0.016 0.016 0.015 0.015 0.015 0.013 0.013
0.012 0.012 0.011 0.011 0.011 0.010 0.009 0.008 0.007 0.007
0.006 0.006 0.005 0.005 0.005 0.005 0.005 0.004
1( 1) 10.8
2( 3) 4.0
3( 2) 4.0
4( 68) 2.5
5( 61) 2.4
6( 66) 2.3
7( 65) 2.2
8( 67) 2.2
9( 13) 2.1
10( 24) 2.1
11( 20) 2.1
12( 60) 2.0
13( 58) 1.9
14( 69) 1.9
15( 59) 1.9
16( 9) 1.9
17( 7) 1.8
18( 17) 1.8
19( 12) 1.8
20( 35) 1.8
21( 57) 1.8
22( 62) 1.8
23( 8) 1.8
24( 15) 1.8
25( 47) 1.7
26( 5) 1.7
27( 64) 1.7
28( 18) 1.7
29( 33) 1.7
30( 30) 1.7
31( 14) 1.6
32( 19) 1.6
33( 42) 1.6
34( 4) 1.6
35( 11) 1.6
36( 74) 1.5
37( 31) 1.5
38( 32) 1.5
39( 73) 1.3
40( 16) 1.3
41( 44) 1.2
42( 6) 1.2
43( 46) 1.1
44( 39) 1.1
45( 22) 1.1
46( 27) 1.0
47( 40) 0.9
48( 38) 0.8
49( 75) 0.7
50( 54) 0.7
51( 34) 0.6
52( 45) 0.6
53( 23) 0.5
54( 41) 0.5
55( 10) 0.5
56( 77) 0.5
57( 80) 0.5
58( 71) 0.4
ensemble average free energy (Eh) : -310.843646
ensemble entropy (cal/mol K) : 7.658
ensemble total free energy (kcal/mol) : -310.844731
removing acidic XH 12
reading rotamer data from <anmr_rotamer>
number of unique conformers from CREST 86
number of unique conformers from enso 58
conformer 1 averaging over 6 rotamers
conformer 2 averaging over 3 rotamers
conformer 3 averaging over 1 rotamers
conformer 4 averaging over 6 rotamers
conformer 5 averaging over 6 rotamers
conformer 6 averaging over 6 rotamers
conformer 7 averaging over 6 rotamers
conformer 8 averaging over 6 rotamers
conformer 9 averaging over 6 rotamers
conformer 10 averaging over 6 rotamers
conformer 11 averaging over 1 rotamers
conformer 12 averaging over 7 rotamers
conformer 13 averaging over 5 rotamers
conformer 14 averaging over 3 rotamers
conformer 15 averaging over 2 rotamers
conformer 16 averaging over 1 rotamers
conformer 17 averaging over 2 rotamers
conformer 18 averaging over 1 rotamers
conformer 19 averaging over 1 rotamers
conformer 20 averaging over 1 rotamers
conformer 21 averaging over 1 rotamers
conformer 21 not in anmr_enso, skipping
conformer 22 averaging over 5 rotamers
conformer 23 averaging over 6 rotamers
conformer 24 averaging over 1 rotamers
conformer 25 averaging over 1 rotamers
conformer 25 not in anmr_enso, skipping
conformer 26 averaging over 2 rotamers
conformer 26 not in anmr_enso, skipping
conformer 27 averaging over 1 rotamers
conformer 28 averaging over 5 rotamers
conformer 28 not in anmr_enso, skipping
conformer 29 averaging over 1 rotamers
conformer 29 not in anmr_enso, skipping
conformer 30 averaging over 6 rotamers
conformer 31 averaging over 6 rotamers
conformer 32 averaging over 1 rotamers
conformer 33 averaging over 1 rotamers
conformer 34 averaging over 2 rotamers
conformer 35 averaging over 1 rotamers
conformer 36 averaging over 6 rotamers
conformer 36 not in anmr_enso, skipping
conformer 37 averaging over 6 rotamers
conformer 37 not in anmr_enso, skipping
conformer 38 averaging over 3 rotamers
conformer 39 averaging over 1 rotamers
conformer 40 averaging over 1 rotamers
conformer 41 averaging over 1 rotamers
conformer 42 averaging over 1 rotamers
conformer 43 averaging over 6 rotamers
conformer 43 not in anmr_enso, skipping
conformer 44 averaging over 3 rotamers
conformer 45 averaging over 1 rotamers
conformer 46 averaging over 2 rotamers
conformer 47 averaging over 2 rotamers
conformer 48 averaging over 6 rotamers
conformer 48 not in anmr_enso, skipping
conformer 49 averaging over 1 rotamers
conformer 49 not in anmr_enso, skipping
conformer 50 averaging over 1 rotamers
conformer 50 not in anmr_enso, skipping
conformer 51 averaging over 1 rotamers
conformer 51 not in anmr_enso, skipping
conformer 52 averaging over 1 rotamers
conformer 52 not in anmr_enso, skipping
conformer 53 averaging over 1 rotamers
conformer 53 not in anmr_enso, skipping
conformer 54 averaging over 1 rotamers
conformer 55 averaging over 1 rotamers
conformer 55 not in anmr_enso, skipping
conformer 56 averaging over 1 rotamers
conformer 56 not in anmr_enso, skipping
conformer 57 averaging over 6 rotamers
conformer 58 averaging over 3 rotamers
conformer 59 averaging over 1 rotamers
conformer 60 averaging over 1 rotamers
conformer 61 averaging over 1 rotamers
conformer 62 averaging over 1 rotamers
conformer 63 averaging over 1 rotamers
conformer 63 not in anmr_enso, skipping
conformer 64 averaging over 4 rotamers
conformer 65 averaging over 4 rotamers
conformer 66 averaging over 2 rotamers
conformer 67 averaging over 1 rotamers
conformer 68 averaging over 1 rotamers
conformer 69 averaging over 6 rotamers
conformer 70 averaging over 3 rotamers
conformer 70 not in anmr_enso, skipping
conformer 71 averaging over 2 rotamers
conformer 72 averaging over 1 rotamers
conformer 72 not in anmr_enso, skipping
conformer 73 averaging over 1 rotamers
conformer 74 averaging over 1 rotamers
conformer 75 averaging over 1 rotamers
conformer 76 averaging over 1 rotamers
conformer 76 not in anmr_enso, skipping
conformer 77 averaging over 6 rotamers
conformer 78 averaging over 4 rotamers
conformer 78 not in anmr_enso, skipping
conformer 79 averaging over 1 rotamers
conformer 79 not in anmr_enso, skipping
conformer 80 averaging over 1 rotamers
conformer 81 averaging over 1 rotamers
conformer 81 not in anmr_enso, skipping
conformer 82 averaging over 1 rotamers
conformer 82 not in anmr_enso, skipping
conformer 83 averaging over 3 rotamers
conformer 83 not in anmr_enso, skipping
conformer 84 averaging over 1 rotamers
conformer 84 not in anmr_enso, skipping
conformer 85 averaging over 1 rotamers
conformer 85 not in anmr_enso, skipping
conformer 86 averaging over 1 rotamers
conformer 86 not in anmr_enso, skipping
average over 165 in CRE
====================================================
reading J/sigma data for conformer 1
====================================================
reading from CONF1/NMR/nmrprop.dat
reading from CONF1/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.436
2 6 1 2.436
3 8 1 1.796
4 10 1 3.853
5 13 1 3.853
6 14 1 1.796
7 15 1 6.197
8 16 1 6.060
9 17 3 1.856
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -13.67040 0.00000
3 2.76599 6.85478 0.00000
4 -0.22109 0.49409 5.91314 0.00000
5 0.15160 -0.22500 1.58791 -11.38440 0.00000
6 8.73554 2.67079 -14.11040 2.04535 7.66190 0.00000
7 8.44440 6.45120 -0.30680 0.08936 0.04854 -0.28550
8 -1.61077 -2.25943 0.28100 -0.07075 -0.04005 0.20780
9 0.66623 1.00136 0.04434 0.06734 0.04569 0.03071
7 8 9
7 0.00000
8 10.08350 0.00000
9 -1.88014 8.26294 0.00000
writing spin system data for this confomer to tmpanmr.1
====================================================
reading J/sigma data for conformer 3
====================================================
reading from CONF3/NMR/nmrprop.dat
reading from CONF3/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.421
2 6 1 2.421
3 8 1 1.817
4 10 1 3.863
5 13 1 3.863
6 14 1 1.817
7 15 1 6.283
8 16 1 6.109
9 17 3 1.808
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -13.78920 0.00000
3 2.53297 8.87557 0.00000
4 -0.21076 0.11884 7.65418 0.00000
5 0.45133 -0.20630 2.04117 -11.38290 0.00000
6 7.17992 2.48495 -14.07960 1.69635 5.97570 0.00000
7 6.32259 8.33571 -0.28331 0.05059 0.09401 -0.30529
8 -2.34610 -1.58860 0.19949 -0.04433 -0.07907 0.28121
9 2.14133 0.93346 0.06896 0.05212 0.10158 0.12852
7 8 9
7 0.00000
8 9.72840 0.00000
9 -2.62903 6.06336 0.00000
writing spin system data for this confomer to tmpanmr.3
====================================================
reading J/sigma data for conformer 2
====================================================
reading from CONF2/NMR/nmrprop.dat
reading from CONF2/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.424
2 6 1 2.424
3 8 1 1.819
4 10 1 3.859
5 13 1 3.859
6 14 1 1.819
7 15 1 6.269
8 16 1 6.110
9 17 3 1.803
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -13.95210 0.00000
3 2.64011 7.02231 0.00000
4 -0.22744 0.45451 5.85144 0.00000
5 0.12812 -0.23219 1.71695 -11.39500 0.00000
6 8.85465 2.56163 -14.07420 2.19172 7.67539 0.00000
7 8.26741 6.21989 -0.30384 0.09889 0.05261 -0.28256
8 -1.64874 -2.43416 0.28379 -0.07910 -0.04400 0.19861
9 0.91159 1.17300 0.05753 0.06777 0.05302 0.09777
7 8 9
7 0.00000
8 9.72580 0.00000
9 -1.65107 6.72730 0.00000
writing spin system data for this confomer to tmpanmr.2
====================================================
reading J/sigma data for conformer 68
====================================================
reading from CONF68/NMR/nmrprop.dat
reading from CONF68/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.080
2 6 1 2.080
3 8 1 1.607
4 10 1 3.851
5 13 1 3.851
6 14 1 1.607
7 15 1 5.937
8 16 1 5.950
9 17 3 1.735
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -12.73380 0.00000
3 11.84459 3.66772 0.00000
4 -0.24748 -0.28547 4.62120 0.00000
5 -0.28102 -0.25234 11.03822 -7.64950 0.00000
6 3.61149 12.05310 -11.68820 10.96201 4.57247 0.00000
7 8.36250 6.45700 -0.26872 0.07101 0.10889 -0.26228
8 -1.43424 -1.97396 0.13989 -0.00301 -0.00369 0.16541
9 0.51403 0.82080 -0.05086 -0.00722 -0.00765 -0.04665
7 8 9
7 0.00000
8 9.95390 0.00000
9 -1.91431 8.18934 0.00000
writing spin system data for this confomer to tmpanmr.68
====================================================
reading J/sigma data for conformer 61
====================================================
reading from CONF61/NMR/nmrprop.dat
reading from CONF61/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.433
2 6 1 2.433
3 8 1 1.663
4 10 1 3.803
5 13 1 3.803
6 14 1 1.663
7 15 1 5.653
8 16 1 6.086
9 17 3 1.767
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -13.53910 0.00000
3 2.65141 6.88075 0.00000
4 -0.06222 -0.43978 8.04884 0.00000
5 -0.48280 0.15839 2.48462 -8.16800 0.00000
6 9.00738 2.70206 -14.04120 2.10496 5.98208 0.00000
7 8.41522 6.49468 -0.27545 -0.00799 -0.01181 -0.27315
8 -1.51413 -2.10777 0.24689 0.03125 0.02625 0.18471
9 0.82196 1.57108 0.09387 -0.02487 -0.02470 0.07986
7 8 9
7 0.00000
8 9.97670 0.00000
9 -2.21979 6.53866 0.00000
writing spin system data for this confomer to tmpanmr.61
====================================================
reading J/sigma data for conformer 66
====================================================
reading from CONF66/NMR/nmrprop.dat
reading from CONF66/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.070
2 6 1 2.070
3 8 1 1.592
4 10 1 3.847
5 13 1 3.847
6 14 1 1.592
7 15 1 5.965
8 16 1 5.959
9 17 3 1.737
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -12.44700 0.00000
3 11.90501 3.75249 0.00000
4 -0.25137 -0.28035 4.65176 0.00000
5 -0.28411 -0.24737 11.04210 -7.64580 0.00000
6 3.55637 11.95773 -11.71170 10.95369 4.53775 0.00000
7 6.70292 8.43298 -0.25962 0.11899 0.07871 -0.26728
8 -1.77491 -1.33159 0.17150 -0.00045 0.00115 0.14960
9 0.66847 0.39043 -0.05495 -0.01077 -0.01038 -0.03862
7 8 9
7 0.00000
8 9.93230 0.00000
9 -1.90079 8.20957 0.00000
writing spin system data for this confomer to tmpanmr.66
====================================================
reading J/sigma data for conformer 65
====================================================
reading from CONF65/NMR/nmrprop.dat
reading from CONF65/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.068
2 6 1 2.068
3 8 1 1.592
4 10 1 3.846
5 13 1 3.846
6 14 1 1.592
7 15 1 5.967
8 16 1 5.959
9 17 3 1.736
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -12.46150 0.00000
3 11.86714 3.65730 0.00000
4 -0.24715 -0.28511 4.62754 0.00000
5 -0.28170 -0.25124 11.03765 -7.65160 0.00000
6 3.64217 12.02459 -11.71160 10.95860 4.55910 0.00000
7 8.43644 6.71716 -0.26786 0.07928 0.11932 -0.26004
8 -1.32811 -1.76799 0.15028 0.00081 -0.00061 0.17262
9 0.66721 1.42242 -0.02959 -0.01480 -0.01533 -0.07295
7 8 9
7 0.00000
8 9.92670 0.00000
9 -2.40312 6.33934 0.00000
writing spin system data for this confomer to tmpanmr.65
====================================================
reading J/sigma data for conformer 67
====================================================
reading from CONF67/NMR/nmrprop.dat
reading from CONF67/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.069
2 6 1 2.069
3 8 1 1.591
4 10 1 3.846
5 13 1 3.846
6 14 1 1.591
7 15 1 5.965
8 16 1 5.959
9 17 3 1.736
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -12.45580 0.00000
3 11.86122 3.64805 0.00000
4 -0.24697 -0.28533 4.63255 0.00000
5 -0.28160 -0.25131 11.03908 -7.64760 0.00000
6 3.65026 12.02217 -11.71250 10.95546 4.55451 0.00000
7 8.43429 6.71421 -0.26771 0.07972 0.11948 -0.25999
8 -1.32928 -1.77012 0.15010 0.00073 -0.00053 0.17220
9 0.69387 1.44604 -0.07598 -0.01295 -0.01414 -0.09577
7 8 9
7 0.00000
8 9.92840 0.00000
9 -2.58646 5.96030 0.00000
writing spin system data for this confomer to tmpanmr.67
====================================================
reading J/sigma data for conformer 13
====================================================
reading from CONF13/NMR/nmrprop.dat
reading from CONF13/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.047
2 6 1 2.047
3 8 1 1.530
4 10 1 3.720
5 13 1 3.720
6 14 1 1.530
7 15 1 5.907
8 16 1 5.945
9 17 3 1.729
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -12.47670 0.00000
3 11.70556 3.76919 0.00000
4 -0.24288 -0.28201 4.51980 0.00000
5 -0.28474 -0.23097 11.06302 -10.74900 0.00000
6 3.48531 11.69925 -11.50210 11.11378 4.44270 0.00000
7 6.75544 8.46726 -0.25593 0.11736 0.07834 -0.26277
8 -1.76118 -1.32512 0.16908 -0.00461 -0.00769 0.14862
9 0.66657 0.39659 -0.05346 -0.00871 -0.00750 -0.03739
7 8 9
7 0.00000
8 9.94230 0.00000
9 -1.89388 8.24243 0.00000
writing spin system data for this confomer to tmpanmr.13
====================================================
reading J/sigma data for conformer 24
====================================================
reading from CONF24/NMR/nmrprop.dat
reading from CONF24/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.067
2 6 1 2.067
3 8 1 1.576
4 10 1 3.724
5 13 1 3.724
6 14 1 1.576
7 15 1 5.855
8 16 1 5.935
9 17 3 1.730
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -13.15630 0.00000
3 11.63527 3.69918 0.00000
4 -0.22997 -0.28391 4.48292 0.00000
5 -0.28120 -0.24182 11.15199 -10.79360 0.00000
6 3.54693 11.94231 -11.46280 10.97233 4.48937 0.00000
7 8.18504 6.10586 -0.26654 0.05826 0.09244 -0.26086
8 -1.58555 -2.21205 0.12553 -0.01660 -0.01240 0.15327
9 0.87993 1.88191 -0.07444 0.00511 -0.00152 -0.08709
7 8 9
7 0.00000
8 9.97910 0.00000
9 -2.62534 5.93610 0.00000
writing spin system data for this confomer to tmpanmr.24
====================================================
reading J/sigma data for conformer 20
====================================================
reading from CONF20/NMR/nmrprop.dat
reading from CONF20/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.450
2 6 1 2.450
3 8 1 1.580
4 10 1 3.695
5 13 1 3.695
6 14 1 1.580
7 15 1 5.654
8 16 1 6.113
9 17 3 1.757
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -13.16660 0.00000
3 2.71275 9.04789 0.00000
4 0.28507 -0.47778 5.60118 0.00000
5 -0.44839 0.02120 2.19438 -11.34470 0.00000
6 6.99929 2.56007 -14.23030 2.63735 7.94619 0.00000
7 6.57960 8.40680 -0.27320 -0.00633 -0.00447 -0.27770
8 -1.94727 -1.42623 0.18494 0.02163 0.02877 0.23886
9 1.43817 0.69960 0.01258 -0.01919 -0.02059 0.08695
7 8 9
7 0.00000
8 9.96880 0.00000
9 -2.43663 6.23488 0.00000
writing spin system data for this confomer to tmpanmr.20
====================================================
reading J/sigma data for conformer 60
====================================================
reading from CONF60/NMR/nmrprop.dat
reading from CONF60/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.425
2 6 1 2.425
3 8 1 1.635
4 10 1 3.799
5 13 1 3.799
6 14 1 1.635
7 15 1 5.669
8 16 1 6.095
9 17 3 1.773
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -13.21890 0.00000
3 2.61682 6.93007 0.00000
4 -0.07587 -0.44397 8.02479 0.00000
5 -0.48436 0.13319 2.49063 -8.16580 0.00000
6 9.05131 2.64340 -13.98060 2.11196 5.95671 0.00000
7 8.50976 6.75054 -0.28105 -0.00754 -0.01146 -0.27605
8 -1.39882 -1.90148 0.23528 0.03386 0.02724 0.18362
9 0.54631 0.85324 0.06661 -0.01689 -0.01683 0.03444
7 8 9
7 0.00000
8 9.94060 0.00000
9 -1.93291 8.05311 0.00000
writing spin system data for this confomer to tmpanmr.60
====================================================
reading J/sigma data for conformer 58
====================================================
reading from CONF58/NMR/nmrprop.dat
reading from CONF58/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.421
2 6 1 2.421
3 8 1 1.618
4 10 1 3.806
5 13 1 3.806
6 14 1 1.618
7 15 1 5.697
8 16 1 6.109
9 17 3 1.783
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -13.07890 0.00000
3 2.59658 6.97055 0.00000
4 -0.08274 -0.44516 8.01581 0.00000
5 -0.48456 0.12147 2.45618 -8.21390 0.00000
6 9.10685 2.60872 -13.99120 2.08534 5.95828 0.00000
7 8.52727 6.91743 -0.28503 -0.00832 -0.01268 -0.27897
8 -1.32133 -1.74367 0.22357 0.03639 0.02851 0.18063
9 0.74809 1.59235 0.16397 -0.01565 -0.01285 0.04541
7 8 9
7 0.00000
8 9.88760 0.00000
9 -2.65950 5.75579 0.00000
writing spin system data for this confomer to tmpanmr.58
====================================================
reading J/sigma data for conformer 69
====================================================
reading from CONF69/NMR/nmrprop.dat
reading from CONF69/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.341
2 6 1 2.341
3 8 1 1.523
4 10 1 3.955
5 13 1 3.955
6 14 1 1.523
7 15 1 5.919
8 16 1 5.944
9 17 3 1.729
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -12.68660 0.00000
3 11.82647 3.78033 0.00000
4 -0.56130 -0.30097 5.72893 0.00000
5 -0.30423 -0.44270 2.24733 -8.29780 0.00000
6 3.50792 11.59788 -13.79340 2.76985 7.85619 0.00000
7 6.70027 8.49813 -0.25586 0.06376 0.08444 -0.26064
8 -1.87065 -1.37935 0.16969 0.00042 0.00658 0.13901
9 0.71466 0.43376 -0.05448 0.03223 0.01848 -0.03974
7 8 9
7 0.00000
8 9.99280 0.00000
9 -1.88347 8.20761 0.00000
writing spin system data for this confomer to tmpanmr.69
====================================================
reading J/sigma data for conformer 59
====================================================
reading from CONF59/NMR/nmrprop.dat
reading from CONF59/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.424
2 6 1 2.424
3 8 1 1.621
4 10 1 3.802
5 13 1 3.802
6 14 1 1.621
7 15 1 5.686
8 16 1 6.107
9 17 3 1.777
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -13.15960 0.00000
3 2.59337 9.11527 0.00000
4 0.13972 -0.49481 5.82714 0.00000
5 -0.45489 -0.06762 2.08720 -8.16740 0.00000
6 7.11662 2.43954 -13.97630 2.57659 7.99827 0.00000
7 6.84592 8.51928 -0.27766 -0.01212 -0.00778 -0.28334
8 -1.80900 -1.35030 0.18088 0.02821 0.03549 0.22762
9 0.71713 0.47129 0.06239 -0.01477 -0.01410 0.06591
7 8 9
7 0.00000
8 9.89300 0.00000
9 -1.86433 8.05827 0.00000
writing spin system data for this confomer to tmpanmr.59
====================================================
reading J/sigma data for conformer 9
====================================================
reading from CONF9/NMR/nmrprop.dat
reading from CONF9/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.360
2 6 1 2.360
3 8 1 1.455
4 10 1 3.820
5 13 1 3.820
6 14 1 1.455
7 15 1 5.875
8 16 1 5.949
9 17 3 1.733
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -12.78550 0.00000
3 11.80253 3.88041 0.00000
4 -0.53153 -0.30535 5.51562 0.00000
5 -0.30026 -0.42296 2.42770 -11.23510 0.00000
6 3.45663 11.59814 -14.18160 2.90177 7.84151 0.00000
7 6.75555 8.49905 -0.25137 0.06703 0.08827 -0.25623
8 -1.83275 -1.36155 0.16347 -0.00447 0.00567 0.13593
9 1.37196 0.66577 -0.08217 0.07006 0.02818 -0.08267
7 8 9
7 0.00000
8 9.98330 0.00000
9 -2.44186 6.23273 0.00000
writing spin system data for this confomer to tmpanmr.9
====================================================
reading J/sigma data for conformer 7
====================================================
reading from CONF7/NMR/nmrprop.dat
reading from CONF7/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.277
2 6 1 2.277
3 8 1 1.715
4 10 1 3.614
5 13 1 3.614
6 14 1 1.715
7 15 1 5.750
8 16 1 5.959
9 17 3 1.676
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -13.80590 0.00000
3 2.71751 9.07607 0.00000
4 -0.47133 -0.31085 4.56513 0.00000
5 -0.34660 -0.37922 11.30201 -10.98080 0.00000
6 7.22801 2.56512 -11.74690 11.43281 4.52886 0.00000
7 6.28224 8.28486 -0.27991 0.00865 0.00315 -0.28449
8 -2.27546 -1.62654 0.20620 -0.02019 -0.01681 0.27120
9 1.67386 0.80879 -0.02331 0.01427 0.03425 0.01149
7 8 9
7 0.00000
8 10.07900 0.00000
9 -2.46145 6.28195 0.00000
writing spin system data for this confomer to tmpanmr.7
====================================================
reading J/sigma data for conformer 17
====================================================
reading from CONF17/NMR/nmrprop.dat
reading from CONF17/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.371
2 6 1 2.371
3 8 1 1.490
4 10 1 3.821
5 13 1 3.821
6 14 1 1.490
7 15 1 5.843
8 16 1 5.937
9 17 3 1.742
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -13.43160 0.00000
3 11.99737 3.70298 0.00000
4 -0.56363 -0.28113 5.47151 0.00000
5 -0.28320 -0.43644 2.62134 -11.43900 0.00000
6 3.60357 11.69678 -14.08260 3.09255 7.85270 0.00000
7 6.20514 8.26756 -0.25933 0.03750 0.05910 -0.26407
8 -2.20573 -1.57987 0.15611 0.00340 0.00820 0.12179
9 1.70809 0.80906 -0.07694 0.07024 0.03248 -0.08507
7 8 9
7 0.00000
8 10.00910 0.00000
9 -2.48432 6.23408 0.00000
writing spin system data for this confomer to tmpanmr.17
====================================================
reading J/sigma data for conformer 12
====================================================
reading from CONF12/NMR/nmrprop.dat
reading from CONF12/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.051
2 6 1 2.051
3 8 1 1.534
4 10 1 3.724
5 13 1 3.724
6 14 1 1.534
7 15 1 5.946
8 16 1 5.937
9 17 3 1.719
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -12.47740 0.00000
3 11.68781 3.65295 0.00000
4 -0.23635 -0.28215 4.51269 0.00000
5 -0.27916 -0.23294 11.04240 -10.73980 0.00000
6 3.58105 11.78149 -11.49000 11.09634 4.42898 0.00000
7 8.46130 6.72780 -0.26587 0.07768 0.12222 -0.25753
8 -1.32417 -1.76103 0.15048 0.00057 -0.00147 0.17382
9 0.69625 1.45025 -0.07834 -0.01306 -0.01387 -0.09624
7 8 9
7 0.00000
8 9.94400 0.00000
9 -2.59009 5.97032 0.00000
writing spin system data for this confomer to tmpanmr.12
====================================================
reading J/sigma data for conformer 35
====================================================
reading from CONF35/NMR/nmrprop.dat
reading from CONF35/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.403
2 6 1 2.403
3 8 1 1.706
4 10 1 3.729
5 13 1 3.729
6 14 1 1.706
7 15 1 5.737
8 16 1 6.083
9 17 3 1.803
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -14.76950 0.00000
3 4.42572 5.79518 0.00000
4 0.00991 -0.50625 7.87362 0.00000
5 -0.52003 0.17997 3.70252 -11.79570 0.00000
6 8.32469 4.63212 -13.47200 3.15348 5.41637 0.00000
7 8.31202 6.03768 -0.29170 -0.01583 -0.03227 -0.27130
8 -1.78672 -2.48888 0.26635 0.00740 0.01490 0.18835
9 1.01834 1.85043 0.03369 0.01121 -0.00029 0.01080
7 8 9
7 0.00000
8 10.12330 0.00000
9 -2.30187 6.45876 0.00000
writing spin system data for this confomer to tmpanmr.35
====================================================
reading J/sigma data for conformer 57
====================================================
reading from CONF57/NMR/nmrprop.dat
reading from CONF57/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.306
2 6 1 2.306
3 8 1 1.783
4 10 1 3.737
5 13 1 3.737
6 14 1 1.783
7 15 1 5.775
8 16 1 5.941
9 17 3 1.680
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -14.02330 0.00000
3 2.79341 7.09086 0.00000
4 -0.36903 -0.34357 4.77227 0.00000
5 -0.31190 -0.45060 11.40930 -7.92390 0.00000
6 9.10331 2.83082 -11.94380 11.21626 4.62177 0.00000
7 8.21145 6.18645 -0.29195 0.01532 0.01668 -0.28545
8 -1.67036 -2.33724 0.28895 -0.03272 -0.03408 0.21585
9 0.65741 0.99776 0.01715 0.02832 0.02087 0.00951
7 8 9
7 0.00000
8 10.05100 0.00000
9 -1.91738 8.19076 0.00000
writing spin system data for this confomer to tmpanmr.57
====================================================
reading J/sigma data for conformer 62
====================================================
reading from CONF62/NMR/nmrprop.dat
reading from CONF62/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.434
2 6 1 2.434
3 8 1 1.679
4 10 1 3.804
5 13 1 3.804
6 14 1 1.679
7 15 1 5.639
8 16 1 6.078
9 17 3 1.764
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -13.74510 0.00000
3 2.70230 9.00644 0.00000
4 0.18836 -0.49425 5.86975 0.00000
5 -0.45005 -0.03996 2.10003 -8.18330 0.00000
6 7.01142 2.50454 -14.05940 2.60054 8.05028 0.00000
7 6.31734 8.32256 -0.27047 -0.01285 -0.00905 -0.26993
8 -2.23670 -1.59830 0.18523 0.02553 0.02967 0.25337
9 0.89647 0.60265 0.03487 -0.01996 -0.01870 0.03472
7 8 9
7 0.00000
8 10.01050 0.00000
9 -1.87650 8.14328 0.00000
writing spin system data for this confomer to tmpanmr.62
====================================================
reading J/sigma data for conformer 8
====================================================
reading from CONF8/NMR/nmrprop.dat
reading from CONF8/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.393
2 6 1 2.393
3 8 1 1.677
4 10 1 3.714
5 13 1 3.714
6 14 1 1.677
7 15 1 5.719
8 16 1 6.073
9 17 3 1.803
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -14.02200 0.00000
3 3.97948 6.14574 0.00000
4 -0.03430 -0.50906 7.84865 0.00000
5 -0.52217 0.11673 3.60350 -11.75420 0.00000
6 8.61642 4.12065 -13.46480 3.06582 5.42213 0.00000
7 8.62091 6.53429 -0.29839 -0.00820 -0.02200 -0.28031
8 -1.53138 -2.12402 0.24556 0.01279 0.01661 0.18384
9 0.57058 0.87226 0.02833 0.00046 0.00083 0.00330
7 8 9
7 0.00000
8 10.10790 0.00000
9 -1.90697 8.24533 0.00000
writing spin system data for this confomer to tmpanmr.8
====================================================
reading J/sigma data for conformer 15
====================================================
reading from CONF15/NMR/nmrprop.dat
reading from CONF15/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.370
2 6 1 2.370
3 8 1 1.490
4 10 1 3.833
5 13 1 3.833
6 14 1 1.490
7 15 1 5.837
8 16 1 5.937
9 17 3 1.744
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -13.49850 0.00000
3 11.95790 3.71959 0.00000
4 -0.56573 -0.28625 5.43961 0.00000
5 -0.28752 -0.43690 2.64976 -11.28020 0.00000
6 3.58679 11.65302 -14.05520 3.12254 7.81209 0.00000
7 6.16953 8.24427 -0.25915 0.03533 0.05627 -0.26355
8 -2.23604 -1.60006 0.15377 0.00448 0.00852 0.11963
9 0.87150 0.56089 -0.05030 0.03902 0.02401 -0.03391
7 8 9
7 0.00000
8 10.00970 0.00000
9 -1.88522 8.23299 0.00000
writing spin system data for this confomer to tmpanmr.15
====================================================
reading J/sigma data for conformer 47
====================================================
reading from CONF47/NMR/nmrprop.dat
reading from CONF47/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.200
2 6 1 2.200
3 8 1 1.554
4 10 1 3.837
5 13 1 3.837
6 14 1 1.554
7 15 1 5.854
8 16 1 5.954
9 17 3 1.735
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -12.86330 0.00000
3 11.68698 3.78118 0.00000
4 -0.41885 -0.26905 7.73397 0.00000
5 -0.27565 -0.54705 2.73879 -11.72430 0.00000
6 3.43057 12.31097 -13.54720 2.26899 5.71226 0.00000
7 8.07686 5.98394 -0.26912 0.05085 0.02815 -0.26268
8 -1.63929 -2.28631 0.12545 0.01586 0.01614 0.16285
9 0.90235 1.72811 -0.03070 0.02199 0.05080 -0.06527
7 8 9
7 0.00000
8 9.98680 0.00000
9 -2.34784 6.40196 0.00000
writing spin system data for this confomer to tmpanmr.47
====================================================
reading J/sigma data for conformer 5
====================================================
reading from CONF5/NMR/nmrprop.dat
reading from CONF5/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.284
2 6 1 2.284
3 8 1 1.699
4 10 1 3.610
5 13 1 3.610
6 14 1 1.699
7 15 1 5.738
8 16 1 5.940
9 17 3 1.676
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -13.77180 0.00000
3 2.74189 8.95428 0.00000
4 -0.47993 -0.29265 4.55441 0.00000
5 -0.33160 -0.38912 11.42772 -10.99780 0.00000
6 7.15570 2.58113 -11.77450 11.32297 4.55141 0.00000
7 6.36310 8.32420 -0.28807 0.01796 0.01254 -0.29593
8 -2.24053 -1.60177 0.21660 -0.03046 -0.02654 0.28530
9 1.64398 0.79188 -0.02166 0.02116 0.04233 0.01699
7 8 9
7 0.00000
8 10.04940 0.00000
9 -2.45539 6.27149 0.00000
writing spin system data for this confomer to tmpanmr.5
====================================================
reading J/sigma data for conformer 64
====================================================
reading from CONF64/NMR/nmrprop.dat
reading from CONF64/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.336
2 6 1 2.336
3 8 1 1.507
4 10 1 3.943
5 13 1 3.943
6 14 1 1.507
7 15 1 5.939
8 16 1 5.940
9 17 3 1.741
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -12.47750 0.00000
3 11.67550 3.49874 0.00000
4 -0.50032 -0.26479 5.58839 0.00000
5 -0.26873 -0.40117 2.54861 -8.21960 0.00000
6 3.73646 11.55660 -13.61000 3.05906 7.80014 0.00000
7 8.59148 6.92792 -0.24353 -0.00466 -0.00464 -0.23297
8 -1.28295 -1.65715 0.14094 0.05387 0.04403 0.15326
9 0.64390 1.35192 -0.02307 0.02574 0.00045 -0.06856
7 8 9
7 0.00000
8 9.91360 0.00000
9 -2.39346 6.34365 0.00000
writing spin system data for this confomer to tmpanmr.64
====================================================
reading J/sigma data for conformer 18
====================================================
reading from CONF18/NMR/nmrprop.dat
reading from CONF18/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.270
2 6 1 2.270
3 8 1 1.686
4 10 1 3.631
5 13 1 3.631
6 14 1 1.686
7 15 1 5.830
8 16 1 5.995
9 17 3 1.695
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -13.12520 0.00000
3 2.77934 9.16022 0.00000
4 -0.46950 -0.30595 4.54686 0.00000
5 -0.33958 -0.37927 11.19842 -10.94690 0.00000
6 7.23198 2.68576 -11.68410 11.29991 4.45482 0.00000
7 6.84567 8.51223 -0.29100 0.01225 0.00615 -0.30120
8 -1.79958 -1.35932 0.21059 -0.00753 -0.00957 0.25371
9 1.12611 0.62100 -0.00266 0.01314 0.04656 0.04842
7 8 9
7 0.00000
8 10.01520 0.00000
9 -2.30196 6.45890 0.00000
writing spin system data for this confomer to tmpanmr.18
====================================================
reading J/sigma data for conformer 33
====================================================
reading from CONF33/NMR/nmrprop.dat
reading from CONF33/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.062
2 6 1 2.062
3 8 1 1.569
4 10 1 3.729
5 13 1 3.729
6 14 1 1.569
7 15 1 5.908
8 16 1 5.931
9 17 3 1.765
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -12.78550 0.00000
3 11.68508 3.77299 0.00000
4 -0.23545 -0.28026 4.47970 0.00000
5 -0.27855 -0.23134 11.01427 -10.99720 0.00000
6 3.50904 11.86210 -11.46910 11.10666 4.46467 0.00000
7 8.26275 6.23065 -0.26495 0.06310 0.10480 -0.25785
8 -1.48835 -2.07025 0.13506 -0.00471 -0.00539 0.16374
9 0.81207 1.78190 -0.04975 -0.00876 -0.01183 -0.08330
7 8 9
7 0.00000
8 9.93550 0.00000
9 -2.59610 6.08208 0.00000
writing spin system data for this confomer to tmpanmr.33
====================================================
reading J/sigma data for conformer 30
====================================================
reading from CONF30/NMR/nmrprop.dat
reading from CONF30/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.157
2 6 1 2.157
3 8 1 1.495
4 10 1 3.830
5 13 1 3.830
6 14 1 1.495
7 15 1 5.928
8 16 1 5.982
9 17 3 1.727
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -11.90610 0.00000
3 11.73013 3.64144 0.00000
4 -0.41325 -0.27542 7.68071 0.00000
5 -0.28774 -0.53449 2.77718 -11.68280 0.00000
6 3.51406 12.19746 -13.51170 2.29764 5.65037 0.00000
7 8.44617 6.81513 -0.26467 0.10464 0.08666 -0.25843
8 -1.30020 -1.70550 0.15158 0.00276 -0.00806 0.18212
9 0.42313 0.68221 -0.05913 0.02448 0.03899 -0.05308
7 8 9
7 0.00000
8 9.95020 0.00000
9 -1.89717 8.20275 0.00000
writing spin system data for this confomer to tmpanmr.30
====================================================
reading J/sigma data for conformer 14
====================================================
reading from CONF14/NMR/nmrprop.dat
reading from CONF14/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.400
2 6 1 2.400
3 8 1 1.494
4 10 1 3.848
5 13 1 3.848
6 14 1 1.494
7 15 1 5.878
8 16 1 5.924
9 17 3 1.752
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -13.35310 0.00000
3 11.58991 3.79753 0.00000
4 -0.39200 -0.24720 7.80689 0.00000
5 -0.22625 -0.48345 2.72157 -11.37950 0.00000
6 3.42634 11.80142 -14.10350 2.39758 5.61175 0.00000
7 6.29765 8.37875 -0.23849 -0.00914 -0.00826 -0.24831
8 -2.15350 -1.54840 0.13182 0.06059 0.08241 0.11298
9 1.61200 0.77074 -0.07245 -0.00154 0.01894 -0.07885
7 8 9
7 0.00000
8 9.94880 0.00000
9 -2.45442 6.26766 0.00000
writing spin system data for this confomer to tmpanmr.14
====================================================
reading J/sigma data for conformer 19
====================================================
reading from CONF19/NMR/nmrprop.dat
reading from CONF19/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.255
2 6 1 2.255
3 8 1 1.694
4 10 1 3.613
5 13 1 3.613
6 14 1 1.694
7 15 1 5.795
8 16 1 5.969
9 17 3 1.648
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -13.20790 0.00000
3 2.65259 9.11538 0.00000
4 -0.48134 -0.30243 4.60860 0.00000
5 -0.33229 -0.38604 11.23323 -11.08180 0.00000
6 7.27573 2.54030 -11.75900 11.33697 4.43820 0.00000
7 6.65217 8.44883 -0.28412 0.01137 0.00453 -0.29058
8 -1.94785 -1.42955 0.20635 -0.01371 -0.01259 0.25485
9 1.68491 0.68076 0.00799 0.01154 0.02743 0.09791
7 8 9
7 0.00000
8 10.03060 0.00000
9 -2.69725 5.86213 0.00000
writing spin system data for this confomer to tmpanmr.19
====================================================
reading J/sigma data for conformer 42
====================================================
reading from CONF42/NMR/nmrprop.dat
reading from CONF42/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.184
2 6 1 2.184
3 8 1 1.513
4 10 1 3.844
5 13 1 3.844
6 14 1 1.513
7 15 1 5.910
8 16 1 5.965
9 17 3 1.737
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -12.08400 0.00000
3 12.06059 3.50386 0.00000
4 -0.55258 -0.26233 5.51432 0.00000
5 -0.25550 -0.42449 2.38662 -11.70600 0.00000
6 3.66214 11.73950 -13.47360 2.96752 7.67624 0.00000
7 6.49809 8.37851 -0.26024 0.06484 0.08466 -0.26596
8 -1.90325 -1.39375 0.17963 -0.00525 0.00365 0.14487
9 1.24542 0.63267 -0.08105 0.06374 0.02734 -0.09273
7 8 9
7 0.00000
8 9.97050 0.00000
9 -2.34355 6.42723 0.00000
writing spin system data for this confomer to tmpanmr.42
====================================================
reading J/sigma data for conformer 4
====================================================
reading from CONF4/NMR/nmrprop.dat
reading from CONF4/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.436
2 6 1 2.436
3 8 1 1.543
4 10 1 3.697
5 13 1 3.697
6 14 1 1.543
7 15 1 5.718
8 16 1 6.131
9 17 3 1.751
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -13.05110 0.00000
3 2.30805 9.21361 0.00000
4 0.28764 -0.45059 5.64280 0.00000
5 -0.42899 0.01984 2.10899 -11.41040 0.00000
6 7.28003 2.17981 -14.27210 2.54572 7.96249 0.00000
7 6.91201 8.44649 -0.27356 -0.00410 -0.00290 -0.27504
8 -1.67704 -1.28886 0.17500 0.01908 0.02912 0.21150
9 0.66484 0.43972 0.08031 0.00157 -0.00167 0.08432
7 8 9
7 0.00000
8 9.82420 0.00000
9 -1.86052 8.03652 0.00000
writing spin system data for this confomer to tmpanmr.4
====================================================
reading J/sigma data for conformer 11
====================================================
reading from CONF11/NMR/nmrprop.dat
reading from CONF11/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.344
2 6 1 2.344
3 8 1 1.442
4 10 1 3.814
5 13 1 3.814
6 14 1 1.442
7 15 1 5.942
8 16 1 5.972
9 17 3 1.729
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -12.46440 0.00000
3 11.79789 3.96595 0.00000
4 -0.52473 -0.30617 5.52487 0.00000
5 -0.30032 -0.42038 2.37901 -11.21090 0.00000
6 3.49510 11.67425 -14.25770 2.84994 7.83668 0.00000
7 7.10294 8.46416 -0.24855 0.10695 0.12185 -0.25645
8 -1.49031 -1.22729 0.17380 -0.01838 -0.00232 0.15190
9 0.58397 0.34739 -0.05706 0.04205 0.02342 -0.04245
7 8 9
7 0.00000
8 9.91730 0.00000
9 -1.88530 8.21164 0.00000
writing spin system data for this confomer to tmpanmr.11
====================================================
reading J/sigma data for conformer 74
====================================================
reading from CONF74/NMR/nmrprop.dat
reading from CONF74/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.411
2 6 1 2.411
3 8 1 1.725
4 10 1 3.954
5 13 1 3.954
6 14 1 1.725
7 15 1 5.752
8 16 1 5.960
9 17 3 1.736
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -17.88340 0.00000
3 12.30850 4.22321 0.00000
4 -0.39857 -0.30110 7.86092 0.00000
5 -0.28853 -0.49491 2.72244 -8.34090 0.00000
6 4.07331 12.63418 -13.54810 2.32650 5.82025 0.00000
7 4.60555 4.83445 -0.27705 -0.03441 -0.04109 -0.28085
8 -3.37896 -3.31654 0.04926 0.07520 0.12410 0.05334
9 2.67628 2.43595 -0.05531 0.01176 0.00437 -0.05319
7 8 9
7 0.00000
8 9.66040 0.00000
9 -2.53660 6.18245 0.00000
writing spin system data for this confomer to tmpanmr.74
====================================================
reading J/sigma data for conformer 31
====================================================
reading from CONF31/NMR/nmrprop.dat
reading from CONF31/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.178
2 6 1 2.178
3 8 1 1.494
4 10 1 3.816
5 13 1 3.816
6 14 1 1.494
7 15 1 5.932
8 16 1 5.952
9 17 3 1.732
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -11.87520 0.00000
3 11.60527 3.79505 0.00000
4 -0.37636 -0.24940 7.71334 0.00000
5 -0.25411 -0.47312 2.75130 -11.61010 0.00000
6 3.34215 11.99814 -13.53210 2.30748 5.67618 0.00000
7 6.70117 8.26973 -0.23663 -0.00650 -0.00620 -0.24767
8 -1.71028 -1.29582 0.15259 0.04532 0.05838 0.14061
9 1.42964 0.59532 -0.07483 0.00016 0.03055 -0.01922
7 8 9
7 0.00000
8 9.91000 0.00000
9 -2.54660 6.07673 0.00000
writing spin system data for this confomer to tmpanmr.31
====================================================
reading J/sigma data for conformer 32
====================================================
reading from CONF32/NMR/nmrprop.dat
reading from CONF32/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.178
2 6 1 2.178
3 8 1 1.481
4 10 1 3.812
5 13 1 3.812
6 14 1 1.481
7 15 1 5.983
8 16 1 5.965
9 17 3 1.736
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -11.71260 0.00000
3 11.90656 3.44999 0.00000
4 -0.48242 -0.25441 5.48583 0.00000
5 -0.24428 -0.38228 2.45150 -11.71260 0.00000
6 3.76915 11.67910 -13.52780 3.00815 7.68802 0.00000
7 8.26788 6.97132 -0.24895 0.00019 -0.00259 -0.23595
8 -1.21595 -1.48075 0.15322 0.04125 0.03565 0.16078
9 0.65557 1.19262 -0.06161 0.04749 0.01016 -0.09235
7 8 9
7 0.00000
8 9.86070 0.00000
9 -2.57372 5.97218 0.00000
writing spin system data for this confomer to tmpanmr.32
====================================================
reading J/sigma data for conformer 73
====================================================
reading from CONF73/NMR/nmrprop.dat
reading from CONF73/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.135
2 6 1 2.135
3 8 1 1.801
4 10 1 3.821
5 13 1 3.821
6 14 1 1.801
7 15 1 5.778
8 16 1 5.953
9 17 3 1.708
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -17.54450 0.00000
3 12.69788 4.27713 0.00000
4 -0.29039 -0.31021 4.64600 0.00000
5 -0.31060 -0.29020 11.00184 -7.75800 0.00000
6 4.16500 12.79649 -11.45840 10.92368 4.56677 0.00000
7 4.60973 4.63847 -0.28801 0.02065 0.02055 -0.28839
8 -3.31601 -3.30529 0.07001 -0.04412 -0.04428 0.06969
9 2.47774 2.57510 -0.04929 0.04323 0.04833 -0.06000
7 8 9
7 0.00000
8 9.61690 0.00000
9 -2.58086 6.01463 0.00000
writing spin system data for this confomer to tmpanmr.73
====================================================
reading J/sigma data for conformer 16
====================================================
reading from CONF16/NMR/nmrprop.dat
reading from CONF16/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.429
2 6 1 2.429
3 8 1 1.661
4 10 1 3.821
5 13 1 3.821
6 14 1 1.661
7 15 1 5.717
8 16 1 5.969
9 17 3 1.742
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -17.90870 0.00000
3 12.24312 4.16252 0.00000
4 -0.38986 -0.29661 7.84512 0.00000
5 -0.29380 -0.48853 2.82103 -11.33520 0.00000
6 4.13230 12.67505 -13.92210 2.46981 5.62013 0.00000
7 4.92355 4.55475 -0.28259 -0.02748 -0.03182 -0.27831
8 -3.24225 -3.40525 0.04752 0.07479 0.12241 0.05718
9 2.47336 2.50653 -0.04962 0.01368 0.00992 -0.05045
7 8 9
7 0.00000
8 9.69740 0.00000
9 -2.44657 6.26932 0.00000
writing spin system data for this confomer to tmpanmr.16
====================================================
reading J/sigma data for conformer 44
====================================================
reading from CONF44/NMR/nmrprop.dat
reading from CONF44/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.250
2 6 1 2.250
3 8 1 1.696
4 10 1 3.818
5 13 1 3.818
6 14 1 1.696
7 15 1 5.732
8 16 1 5.988
9 17 3 1.756
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -16.85570 0.00000
3 12.43967 4.23523 0.00000
4 -0.37194 -0.26233 7.71588 0.00000
5 -0.25857 -0.46736 2.58934 -11.73760 0.00000
6 3.92526 12.78154 -13.41140 2.15352 5.75956 0.00000
7 4.35518 4.84652 -0.27857 -0.02646 -0.03364 -0.28853
8 -3.36645 -3.14765 0.05664 0.07925 0.13315 0.05586
9 2.41117 2.03171 -0.05139 0.00548 -0.00066 -0.03878
7 8 9
7 0.00000
8 9.63920 0.00000
9 -2.25373 6.51235 0.00000
writing spin system data for this confomer to tmpanmr.44
====================================================
reading J/sigma data for conformer 6
====================================================
reading from CONF6/NMR/nmrprop.dat
reading from CONF6/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.380
2 6 1 2.380
3 8 1 1.460
4 10 1 3.830
5 13 1 3.830
6 14 1 1.460
7 15 1 5.935
8 16 1 5.938
9 17 3 1.739
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -12.76270 0.00000
3 11.73840 3.59639 0.00000
4 -0.48843 -0.24521 5.47882 0.00000
5 -0.25582 -0.39594 2.41435 -11.39790 0.00000
6 3.64061 11.66170 -14.03860 2.85432 7.79051 0.00000
7 8.60302 6.75328 -0.24317 -0.00632 -0.00738 -0.23243
8 -1.34412 -1.79848 0.13193 0.06265 0.05015 0.14927
9 0.68493 1.46285 -0.02678 0.03078 0.00220 -0.07008
7 8 9
7 0.00000
8 9.93210 0.00000
9 -2.41404 6.30644 0.00000
writing spin system data for this confomer to tmpanmr.6
====================================================
reading J/sigma data for conformer 46
====================================================
reading from CONF46/NMR/nmrprop.dat
reading from CONF46/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.127
2 6 1 2.127
3 8 1 1.739
4 10 1 3.837
5 13 1 3.837
6 14 1 1.739
7 15 1 5.918
8 16 1 6.123
9 17 3 1.661
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -17.30690 0.00000
3 12.68563 3.93762 0.00000
4 -0.48852 -0.28157 5.50715 0.00000
5 -0.26890 -0.37591 2.47350 -11.81190 0.00000
6 4.05699 12.38026 -13.30900 3.05131 7.69785 0.00000
7 4.46580 5.26290 -0.27849 -0.02758 -0.01632 -0.28441
8 -3.57840 -3.25320 0.05742 0.11640 0.06720 0.04078
9 2.61591 2.02973 -0.04396 0.01651 0.01745 -0.03318
7 8 9
7 0.00000
8 9.59010 0.00000
9 -2.36512 6.55645 0.00000
writing spin system data for this confomer to tmpanmr.46
====================================================
reading J/sigma data for conformer 39
====================================================
reading from CONF39/NMR/nmrprop.dat
reading from CONF39/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.242
2 6 1 2.242
3 8 1 1.699
4 10 1 3.816
5 13 1 3.816
6 14 1 1.699
7 15 1 5.721
8 16 1 5.987
9 17 3 1.737
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -16.83570 0.00000
3 12.44833 4.19093 0.00000
4 -0.36444 -0.26936 7.72957 0.00000
5 -0.27597 -0.46383 2.73940 -11.73320 0.00000
6 3.97231 12.78633 -13.33900 2.28165 5.71257 0.00000
7 4.35549 4.78171 -0.27750 -0.02370 -0.03060 -0.28680
8 -3.34148 -3.16222 0.05602 0.07636 0.12944 0.05708
9 2.53819 2.17122 -0.05447 0.00727 0.00059 -0.04656
7 8 9
7 0.00000
8 9.67700 0.00000
9 -2.41379 6.36173 0.00000
writing spin system data for this confomer to tmpanmr.39
====================================================
reading J/sigma data for conformer 22
====================================================
reading from CONF22/NMR/nmrprop.dat
reading from CONF22/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.098
2 6 1 2.098
3 8 1 1.705
4 10 1 3.702
5 13 1 3.702
6 14 1 1.705
7 15 1 5.779
8 16 1 5.942
9 17 3 1.714
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -16.15060 0.00000
3 12.09696 4.12515 0.00000
4 -0.25017 -0.28968 4.48104 0.00000
5 -0.28502 -0.25023 10.98283 -11.05840 0.00000
6 3.84600 12.37839 -11.29650 11.07208 4.45505 0.00000
7 6.16560 4.55940 -0.28357 0.02948 0.04672 -0.26863
8 -2.63151 -3.19529 0.07539 -0.02940 -0.02910 0.09681
9 1.81678 2.66718 -0.05480 0.02867 0.02640 -0.06566
7 8 9
7 0.00000
8 9.80810 0.00000
9 -2.63865 5.99336 0.00000
writing spin system data for this confomer to tmpanmr.22
====================================================
reading J/sigma data for conformer 27
====================================================
reading from CONF27/NMR/nmrprop.dat
reading from CONF27/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.118
2 6 1 2.118
3 8 1 1.738
4 10 1 3.698
5 13 1 3.698
6 14 1 1.738
7 15 1 5.744
8 16 1 5.957
9 17 3 1.709
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -17.32530 0.00000
3 12.34669 4.16999 0.00000
4 -0.26097 -0.30275 4.52929 0.00000
5 -0.29686 -0.27033 11.18262 -10.82800 0.00000
6 4.09788 12.64364 -11.29460 10.97772 4.47247 0.00000
7 5.15148 4.41782 -0.29004 0.02965 0.03015 -0.28136
8 -3.10392 -3.37288 0.06656 -0.04343 -0.03947 0.07394
9 2.25962 2.39279 -0.04917 0.04092 0.03305 -0.04803
7 8 9
7 0.00000
8 9.68110 0.00000
9 -2.34024 6.40329 0.00000
writing spin system data for this confomer to tmpanmr.27
====================================================
reading J/sigma data for conformer 40
====================================================
reading from CONF40/NMR/nmrprop.dat
reading from CONF40/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.251
2 6 1 2.251
3 8 1 1.699
4 10 1 3.823
5 13 1 3.823
6 14 1 1.699
7 15 1 5.736
8 16 1 5.982
9 17 3 1.746
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -16.40830 0.00000
3 12.56231 4.03560 0.00000
4 -0.46198 -0.26635 5.62528 0.00000
5 -0.26062 -0.36595 2.24851 -11.75320 0.00000
6 4.01052 12.52867 -13.42610 2.81002 7.73639 0.00000
7 5.30145 4.29235 -0.28487 -0.02888 -0.02332 -0.26903
8 -2.92822 -3.31898 0.05631 0.13408 0.08022 0.06279
9 2.07506 2.68628 -0.05036 -0.00283 0.00510 -0.06185
7 8 9
7 0.00000
8 9.72550 0.00000
9 -2.58839 6.07535 0.00000
writing spin system data for this confomer to tmpanmr.40
====================================================
reading J/sigma data for conformer 38
====================================================
reading from CONF38/NMR/nmrprop.dat
reading from CONF38/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.250
2 6 1 2.250
3 8 1 1.697
4 10 1 3.820
5 13 1 3.820
6 14 1 1.697
7 15 1 5.730
8 16 1 5.988
9 17 3 1.744
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -16.81420 0.00000
3 12.73069 4.02779 0.00000
4 -0.48619 -0.28070 5.51171 0.00000
5 -0.26860 -0.37581 2.46820 -11.80820 0.00000
6 4.12951 12.44411 -13.29270 3.04775 7.70515 0.00000
7 4.33679 4.94101 -0.28161 -0.02904 -0.01946 -0.28709
8 -3.35025 -3.13985 0.07072 0.10762 0.06248 0.05618
9 2.58407 2.18032 -0.05610 0.01709 0.02071 -0.04638
7 8 9
7 0.00000
8 9.67870 0.00000
9 -2.46896 6.29837 0.00000
writing spin system data for this confomer to tmpanmr.38
====================================================
reading J/sigma data for conformer 75
====================================================
reading from CONF75/NMR/nmrprop.dat
reading from CONF75/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.065
2 6 1 2.065
3 8 1 1.616
4 10 1 3.853
5 13 1 3.853
6 14 1 1.616
7 15 1 5.992
8 16 1 6.000
9 17 3 1.689
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -12.66260 0.00000
3 12.01182 3.76441 0.00000
4 -0.25587 -0.28078 4.61867 0.00000
5 -0.28702 -0.24992 11.02596 -7.64770 0.00000
6 3.58322 11.96834 -11.67760 10.96146 4.56812 0.00000
7 6.27461 8.24569 -0.25945 0.10035 0.06575 -0.26505
8 -2.01238 -1.37622 0.17441 -0.00545 -0.00585 0.13639
9 0.85783 0.65843 -0.02914 -0.00604 -0.00517 0.01518
7 8 9
7 0.00000
8 9.61780 0.00000
9 -1.66335 6.72287 0.00000
writing spin system data for this confomer to tmpanmr.75
====================================================
reading J/sigma data for conformer 54
====================================================
reading from CONF54/NMR/nmrprop.dat
reading from CONF54/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.246
2 6 1 2.246
3 8 1 1.634
4 10 1 3.831
5 13 1 3.831
6 14 1 1.634
7 15 1 5.932
8 16 1 5.942
9 17 3 1.716
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -15.09630 0.00000
3 1.82225 7.60534 0.00000
4 -0.43198 -0.36147 4.63426 0.00000
5 -0.35048 -0.51987 11.48884 -10.80340 0.00000
6 9.44821 1.67470 -11.86810 11.07982 4.49068 0.00000
7 5.67601 8.08159 -0.30963 -0.05742 -0.09958 -0.29877
8 -2.57574 -1.90106 0.28747 -0.00753 -0.00657 0.27323
9 1.02715 0.71616 0.05237 0.00343 0.00483 0.01751
7 8 9
7 0.00000
8 10.03610 0.00000
9 -1.91186 8.28355 0.00000
writing spin system data for this confomer to tmpanmr.54
====================================================
reading J/sigma data for conformer 34
====================================================
reading from CONF34/NMR/nmrprop.dat
reading from CONF34/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.010
2 6 1 2.010
3 8 1 1.749
4 10 1 3.701
5 13 1 3.701
6 14 1 1.749
7 15 1 5.880
8 16 1 6.062
9 17 3 1.644
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -17.48230 0.00000
3 12.36860 4.15779 0.00000
4 -0.24887 -0.29509 4.43357 0.00000
5 -0.28988 -0.26167 11.08456 -11.10760 0.00000
6 3.99350 12.61941 -11.39530 10.93946 4.55190 0.00000
7 5.45984 4.51206 -0.28444 0.03061 0.03519 -0.27426
8 -3.12911 -3.54079 0.05327 -0.04271 -0.04189 0.07083
9 2.41162 3.06631 -0.04451 0.04201 0.03858 -0.05757
7 8 9
7 0.00000
8 9.50950 0.00000
9 -2.90563 5.75603 0.00000
writing spin system data for this confomer to tmpanmr.34
====================================================
reading J/sigma data for conformer 45
====================================================
reading from CONF45/NMR/nmrprop.dat
reading from CONF45/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.180
2 6 1 2.180
3 8 1 1.684
4 10 1 3.833
5 13 1 3.833
6 14 1 1.684
7 15 1 5.879
8 16 1 6.062
9 17 3 1.684
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -15.88760 0.00000
3 12.32686 3.96852 0.00000
4 -0.35798 -0.25462 7.75148 0.00000
5 -0.25590 -0.45230 2.64726 -11.73650 0.00000
6 3.75888 12.53204 -13.52620 2.19970 5.76476 0.00000
7 4.58829 6.13441 -0.25928 -0.02176 -0.02684 -0.27502
8 -3.34025 -2.65855 0.06975 0.09066 0.14884 0.04915
9 2.47032 1.90020 -0.03192 -0.00037 -0.00584 -0.00140
7 8 9
7 0.00000
8 9.55160 0.00000
9 -2.61644 6.38216 0.00000
writing spin system data for this confomer to tmpanmr.45
====================================================
reading J/sigma data for conformer 23
====================================================
reading from CONF23/NMR/nmrprop.dat
reading from CONF23/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.361
2 6 1 2.361
3 8 1 1.495
4 10 1 3.837
5 13 1 3.837
6 14 1 1.495
7 15 1 5.916
8 16 1 5.995
9 17 3 1.694
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -13.12510 0.00000
3 11.58842 3.56757 0.00000
4 -0.42289 -0.30953 7.88832 0.00000
5 -0.31420 -0.53688 2.78265 -11.23770 0.00000
6 3.73918 12.06123 -14.19870 2.41435 5.66218 0.00000
7 8.30636 6.32344 -0.25605 0.06143 0.04067 -0.25255
8 -1.42679 -2.09781 0.12600 0.01062 0.00418 0.16820
9 0.89005 1.87304 -0.04373 0.02854 0.07184 -0.07493
7 8 9
7 0.00000
8 9.67710 0.00000
9 -2.71953 5.91565 0.00000
writing spin system data for this confomer to tmpanmr.23
====================================================
reading J/sigma data for conformer 41
====================================================
reading from CONF41/NMR/nmrprop.dat
reading from CONF41/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.141
2 6 1 2.141
3 8 1 1.515
4 10 1 3.843
5 13 1 3.843
6 14 1 1.515
7 15 1 5.974
8 16 1 6.028
9 17 3 1.685
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -11.82880 0.00000
3 11.72405 3.67497 0.00000
4 -0.41593 -0.27549 7.69343 0.00000
5 -0.28789 -0.54318 2.80588 -11.67520 0.00000
6 3.49442 12.23976 -13.46850 2.32579 5.65050 0.00000
7 8.36968 6.55082 -0.26007 0.08786 0.06954 -0.25533
8 -1.25246 -1.79154 0.15127 0.00224 -0.00974 0.19443
9 0.69681 1.47459 -0.00595 0.02477 0.06388 -0.05573
7 8 9
7 0.00000
8 9.67010 0.00000
9 -2.34134 6.42225 0.00000
writing spin system data for this confomer to tmpanmr.41
====================================================
reading J/sigma data for conformer 10
====================================================
reading from CONF10/NMR/nmrprop.dat
reading from CONF10/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.355
2 6 1 2.355
3 8 1 1.815
4 10 1 3.899
5 13 1 3.899
6 14 1 1.815
7 15 1 6.364
8 16 1 6.234
9 17 3 1.845
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -13.01560 0.00000
3 4.32748 6.00798 0.00000
4 -0.28706 0.53241 6.19556 0.00000
5 0.17999 -0.27544 1.27749 -11.08210 0.00000
6 8.42857 4.24548 -13.95960 1.67805 7.70051 0.00000
7 7.45855 7.68045 -0.16577 0.03610 0.02710 -0.19223
8 -0.96464 -1.50766 0.17974 -0.08421 -0.05519 0.17626
9 0.71436 1.05660 -0.00217 0.08549 0.07724 -0.03633
7 8 9
7 0.00000
8 9.82800 0.00000
9 -2.15674 6.54381 0.00000
writing spin system data for this confomer to tmpanmr.10
====================================================
reading J/sigma data for conformer 77
====================================================
reading from CONF77/NMR/nmrprop.dat
reading from CONF77/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.400
2 6 1 2.400
3 8 1 1.693
4 10 1 3.888
5 13 1 3.888
6 14 1 1.693
7 15 1 6.212
8 16 1 6.092
9 17 3 1.744
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -12.83980 0.00000
3 4.08210 6.06131 0.00000
4 -0.11295 -0.52471 8.08303 0.00000
5 -0.54768 0.03424 3.26793 -8.46860 0.00000
6 8.61112 4.23787 -13.28240 2.77706 5.77438 0.00000
7 7.32062 7.90288 -0.07675 0.16330 0.16540 -0.14505
8 -0.99854 -1.26796 0.23053 0.01219 0.00971 0.21837
9 0.49893 0.38920 0.00804 -0.01551 -0.01643 -0.03076
7 8 9
7 0.00000
8 9.85230 0.00000
9 -1.89864 8.17182 0.00000
writing spin system data for this confomer to tmpanmr.77
====================================================
reading J/sigma data for conformer 80
====================================================
reading from CONF80/NMR/nmrprop.dat
reading from CONF80/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.439
2 6 1 2.439
3 8 1 1.672
4 10 1 4.054
5 13 1 4.054
6 14 1 1.672
7 15 1 6.105
8 16 1 5.951
9 17 3 1.747
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -15.34130 0.00000
3 2.43509 7.02694 0.00000
4 -0.12414 -0.55103 8.04404 0.00000
5 -0.55248 0.06796 2.92442 -9.00490 0.00000
6 9.35286 2.34291 -13.93180 2.41742 5.86262 0.00000
7 5.39805 7.69485 -0.30183 -0.23883 -0.18477 -0.28707
8 -2.72983 -2.04007 0.32894 0.02296 0.01484 0.28196
9 2.16658 1.18501 0.09426 0.02081 0.02715 -0.00173
7 8 9
7 0.00000
8 10.04950 0.00000
9 -2.56030 6.00770 0.00000
writing spin system data for this confomer to tmpanmr.80
====================================================
reading J/sigma data for conformer 71
====================================================
reading from CONF71/NMR/nmrprop.dat
reading from CONF71/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 5 1 2.241
2 6 1 2.241
3 8 1 1.746
4 10 1 3.845
5 13 1 3.845
6 14 1 1.746
7 15 1 5.968
8 16 1 6.018
9 17 3 1.766
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -12.99260 0.00000
3 3.33788 8.97902 0.00000
4 -0.46982 -0.29033 4.70020 0.00000
5 -0.31676 -0.38359 11.29696 -7.59810 0.00000
6 6.88352 3.18338 -11.38960 11.33242 4.48032 0.00000
7 8.09893 7.41507 -0.17098 0.16629 0.09751 -0.12702
8 -1.29063 -0.99957 0.21887 -0.02393 -0.02417 0.22513
9 0.80704 0.89444 -0.04466 0.03967 0.04213 0.03581
7 8 9
7 0.00000
8 9.82330 0.00000
9 -2.55474 6.22824 0.00000
writing spin system data for this confomer to tmpanmr.71
=====================================================
average over conformers
=====================================================
conformer 1 (normalized) population : 0.108
conformer 3 (normalized) population : 0.040
conformer 2 (normalized) population : 0.040
conformer 68 (normalized) population : 0.025
conformer 61 (normalized) population : 0.024
conformer 66 (normalized) population : 0.023
conformer 65 (normalized) population : 0.022
conformer 67 (normalized) population : 0.022
conformer 13 (normalized) population : 0.021
conformer 24 (normalized) population : 0.021
conformer 20 (normalized) population : 0.021
conformer 60 (normalized) population : 0.020
conformer 58 (normalized) population : 0.019
conformer 69 (normalized) population : 0.019
conformer 59 (normalized) population : 0.019
conformer 9 (normalized) population : 0.019
conformer 7 (normalized) population : 0.018
conformer 17 (normalized) population : 0.018
conformer 12 (normalized) population : 0.018
conformer 35 (normalized) population : 0.018
conformer 57 (normalized) population : 0.018
conformer 62 (normalized) population : 0.018
conformer 8 (normalized) population : 0.018
conformer 15 (normalized) population : 0.018
conformer 47 (normalized) population : 0.017
conformer 5 (normalized) population : 0.017
conformer 64 (normalized) population : 0.017
conformer 18 (normalized) population : 0.017
conformer 33 (normalized) population : 0.017
conformer 30 (normalized) population : 0.017
conformer 14 (normalized) population : 0.016
conformer 19 (normalized) population : 0.016
conformer 42 (normalized) population : 0.016
conformer 4 (normalized) population : 0.016
conformer 11 (normalized) population : 0.016
conformer 74 (normalized) population : 0.015
conformer 31 (normalized) population : 0.015
conformer 32 (normalized) population : 0.015
conformer 73 (normalized) population : 0.013
conformer 16 (normalized) population : 0.013
conformer 44 (normalized) population : 0.012
conformer 6 (normalized) population : 0.012
conformer 46 (normalized) population : 0.011
conformer 39 (normalized) population : 0.011
conformer 22 (normalized) population : 0.011
conformer 27 (normalized) population : 0.010
conformer 40 (normalized) population : 0.009
conformer 38 (normalized) population : 0.008
conformer 75 (normalized) population : 0.007
conformer 54 (normalized) population : 0.007
conformer 34 (normalized) population : 0.006
conformer 45 (normalized) population : 0.006
conformer 23 (normalized) population : 0.005
conformer 41 (normalized) population : 0.005
conformer 10 (normalized) population : 0.005
conformer 77 (normalized) population : 0.005
conformer 80 (normalized) population : 0.005
conformer 71 (normalized) population : 0.004
# in coord file # nucs delta(ppm)
1 5 1 2.292 +/- 0.02
2 6 1 2.292 +/- 0.02
3 8 1 1.645 +/- 0.02
4 10 1 3.804 +/- 0.01
5 13 1 3.804 +/- 0.01
6 14 1 1.645 +/- 0.02
7 15 1 5.911 +/- 0.02
8 16 1 6.009 +/- 0.01
9 17 3 1.754 +/- 0.01
MATRIX PRINTED: conformer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -13.67188 0.00000
3 7.62604 5.63680 0.00000
4 -0.27586 -0.18585 5.94972 0.00000
5 -0.22604 -0.26286 5.11228 -10.34731 0.00000
6 5.79907 7.64421 -13.17452 5.17721 6.18257 0.00000
7 7.22604 7.01310 -0.27432 0.03870 0.03706 -0.27136
8 -1.90134 -1.97233 0.18174 -0.00105 0.00500 0.17439
9 1.12921 1.15750 -0.00787 0.02117 0.01874 -0.00766
7 8 9
7 0.00000
8 9.91643 0.00000
9 -2.22730 6.95894 0.00000
====================================================
building sub-systems
====================================================
J neighbors of spin 1 : 2 3 4 5 6 7 8 9
J neighbors of spin 2 : 1 3 4 5 6 7 8 9
J neighbors of spin 3 : 1 2 4 5 6 7 8
J neighbors of spin 4 : 1 2 3 5 6
J neighbors of spin 5 : 1 2 3 4 6
J neighbors of spin 6 : 1 2 3 4 5 7 8
J neighbors of spin 7 : 1 2 3 6 8 9
J neighbors of spin 8 : 1 2 3 6 7 9
J neighbors of spin 9 : 1 2 7 8
1:14 max/av. J neglected : 0.000 0.000
2:13 max/av. J neglected : 0.000 0.000
3:12 max/av. J neglected : 0.000 0.000
4:11 max/av. J neglected : 0.000 0.000
5:10 max/av. J neglected : 0.000 0.000
6: 9 max/av. J neglected : 0.000 0.000
7: 8 max/av. J neglected : 0.276 0.138
8: 7 max/av. J neglected : 0.276 0.138
9: 6 max/av. J neglected : 0.276 0.099
selected scheme 6
system largest neglected JAB nuclei (dummy atoms are negative)
1 0.000 1 2 3 4 5 6 7 8 9
=== FRAGMENTED SYSTEM ===
====================================================
solving (J/sigma) averaged spin Hamiltonian
====================================================
spinsystem 1 with 9 spins
1024 product functions 12 Mt blocks, largest is 210
1( 1) 2( 9) 3( 37) 4( 93) 7( 210) 10( 37) 11( 9) 12( 1) 5( 162) 6( 210) 8( 162) 9( 93)
first maxtrix multiply, sparsity in % 99.536 ...
second maxtrix multiply, sparsity in % 94.792 ...
512 product functions 10 Mt blocks, largest is 126
1( 1) 4( 84) 7( 84) 9( 9) 2( 9) 3( 36) 10( 1) 5( 126) 8( 36) 6( 126)
first maxtrix multiply, sparsity in % 99.121 ...
second maxtrix multiply, sparsity in % 91.890 ...
done.
12462 non-zero transitions.
spectrum on <anmr.dat>
Range (delta in ppm) 0.539825439453125 7.05391174316406
Range (delta in Hz) 215.930175781250 2821.56469726562
Min/max Int. ) 4.179619316900532E-003
computing spectrum ...
done.
writing output file ...
done.
After ANMR finished computing, the file anmr.dat is written and it contains
the spectrum (intensity vs shift) the user can plot:
$ nmrplot.py -i anmr.dat exp.dat -start 0 -end 6.5 -o 1Hspectrum -orientation 1 -1
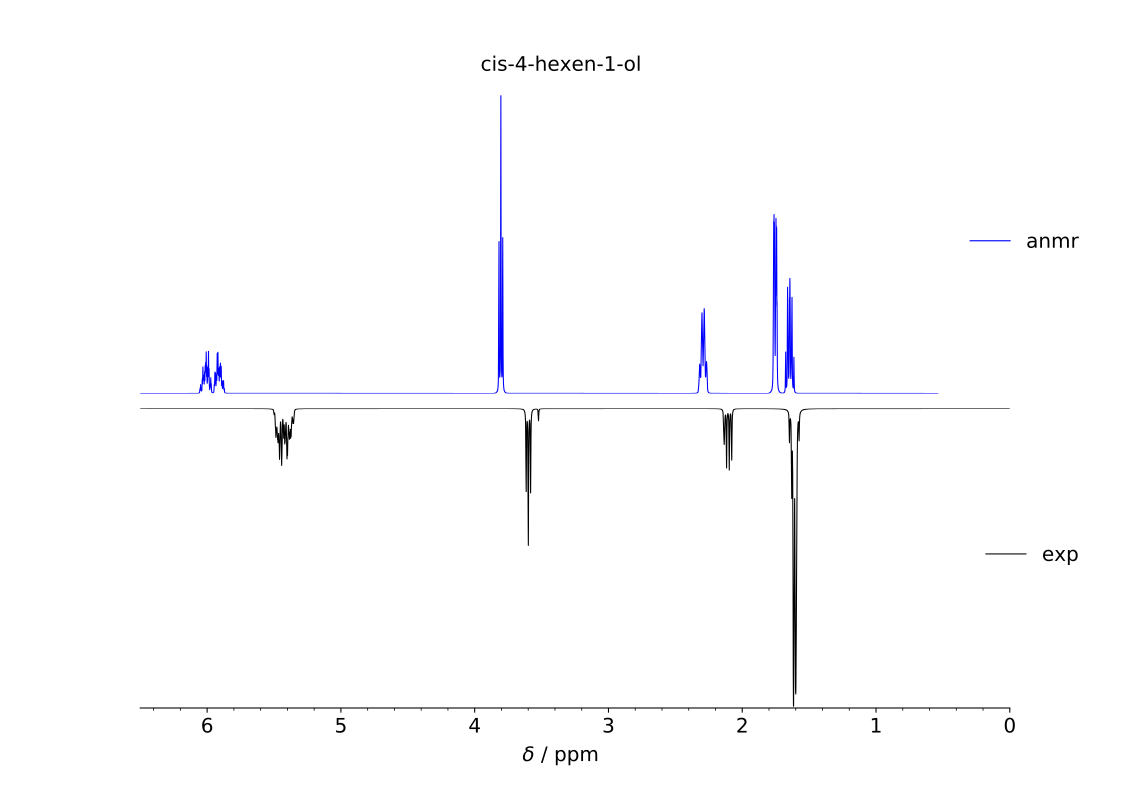
1H NMR spectrum of cis-4-hexen-1-ol in chloroform at 400 MHz, comparing calculated and experimental spectrum. Exp taken from [SDBSWeb : https://sdbs.db.aist.go.jp (National Institute of Advanced Industrial Science and Technology,16-10-2019) (SDBSNo. 11748)].
2-methyl-1-pentene
Example of calculating the 1H-NMR spectrum of 2-methyl-1-pentene in CHCl3 at 400 MHz using ORCA
$ cat coord
$coord
-5.1134989926 0.0445408597 0.0007215195 C
-2.3988260553 0.1202192416 0.9598504570 C
-2.0426150350 1.9467776447 1.8509773297 H
-0.4955528936 -0.3025973506 -1.1852527430 C
2.1853738985 -0.2583887206 -0.2367582425 C
3.4286190716 -2.3618737092 0.3005853656 C
2.5901373582 -4.2004809628 0.0485727882 H
5.3374488734 -2.3390838060 1.0097369787 H
3.3398174602 2.3079102171 0.0825121447 C
5.2234930962 2.1788391495 0.8913733279 H
2.1708137054 3.4751040066 1.3098219746 H
3.4682822356 3.2543034689 -1.7427970988 H
-0.7536049204 1.1708293724 -2.6081830586 H
-0.8901990516 -2.1258718566 -2.0673390015 H
-2.1284937554 -1.3401645088 2.3937954454 H
-6.4334377217 0.3509962700 1.5473865797 H
-5.4204111085 1.5054637513 -1.4143993805 H
-5.5276306722 -1.7796998127 -0.8540259276 H
$end
Input structure.
Start with your (at best already optimized) input structure and create the conformers
and rotamers for the CENSO and ANMR calculation.
$ crest coord -gfn2 -alpb chcl3 -T 4 -nmr > crest.out
In our case CREST found 9 conformers within an energy window of 6 kcal/mol.
We then create a new folder for the CENSO reranking and copy the necessary files:
$ mkdir censo
$ cp crest_conformers.xyz coord anmr_nucinfo anmr_rotamer censo/
$ cd censo/
CENSO requires only the file crest_conformers.xyz, but ANMR needs the last
three files. In the new folder we create the file flags.dat and adapt it to our
choosing. Here we want to calculate everything with ORCA using r2SCAN-3c and SMD for
geometry optimization and PBE0/def2-TZVP for the shielding calculation. B97-D/def2-SV(P)
(keyword “b97-d3” in ORCA) is used for the prescreening in part0 to save computation time.
$ censo --input crest_conformers.xyz -func0 b97-d3 -solvent chcl3 -smgsolv1 smd -sm2 smd
--smgsolv2 smd --prog orca -part4 on -prog4J orca -prog4S orca -funcJ pbe0
-funcS pbe0 -basisJ def2-TZVP -basisS def2-TZVP -cactive off > censo.out
______________________________________________________________
| |
| |
| CENSO - Commandline ENSO |
| v 1.1.2 |
| energetic sorting of CREST Conformer Rotamer Ensembles |
| University of Bonn, MCTC |
| Feb 2021 |
| based on ENSO version 2.0.1 |
| F. Bohle and S. Grimme |
| |
|______________________________________________________________|
Please cite:
S. Grimme, F. Bohle, A. Hansen, P. Pracht, S. Spicher, and M. Stahn
J. Phys. Chem. A 2021, 125, 19, 4039-4054.
DOI: https://doi.org/10.1021/acs.jpca.1c00971
This program is distributed in the hope that it will be useful,
but WITHOUT ANY WARRANTY; without even the implied warranty of
MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE.
----------------------------------------------------------------------------------------------------
PARAMETERS
----------------------------------------------------------------------------------------------------
program call: censo --input crest_conformers.xyz --restart -func0 b97-d3 -solvent chcl3 -smgsolv1 smd -sm2 smd --smgsolv2 smd --prog orca -part4 on -prog4J orca -prog4S orca -funcJ pbe0 -funcS pbe0 -basisJ def2-TZVP -basisS def2-TZVP -cactive off
The configuration file .censorc is read from /home/gorges/.censorc.
Reading conformer rotamer ensemble from: /tmp1/gorges/3881124.majestix.thch.uni-bonn.de/crest_conformers.xyz.
Reading file: censo_solvents.json
--------------------------------------------------
CRE SORTING SETTINGS
--------------------------------------------------
number of atoms in system: 18
number of considered conformers: 9
number of all conformers from input: 9
charge: 0
unpaired: 0
solvent: chcl3
temperature: 298.15
evaluate at different temperatures: on
temperature range: 273.15, 278.15, 283.15, 288.15, ...
calculate mRRHO contribution: on
consider symmetry for mRRHO contribution: on
cautious checking for error and failed calculations: on
checking the DFT-ensemble using CREST: off
maxthreads: 7
omp: 4
automatically balance maxthreads and omp: off
--------------------------------------------------
CRE CHEAP-PRESCREENING - PART0
--------------------------------------------------
part0: on
starting number of considered conformers: 9
program for part0: orca
functional for fast single-point: b97-d3
basis set for fast single-point: def2-SV(P)
threshold g_thr(0) for sorting in part0: 4.0
Solvent model used with xTB: alpb
short-notation:
b97-d3/def2-SV(P) // GFNn-xTB (Input geometry)
--------------------------------------------------
CRE PRESCREENING - PART1
--------------------------------------------------
part1: on
program for part1: orca
functional for initial evaluation: r2scan-3c
basis set for initial evaluation: def2-mTZVPP
calculate mRRHO contribution: on
program for mRRHO contribution: xtb
GFN version for mRRHO and/or GBSA_Gsolv: gfn2
Apply constraint to input geometry during mRRHO calculation: on
solvent model applied with xTB: alpb
evaluate at different temperatures: off
threshold g_thr(1) and G_thr(1) for sorting in part1: 3.5
solvent model for Gsolv contribution of part1: smd
short-notation:
r2scan-3c + SMD[chcl3] + GmRRHO(GFN2[alpb]-bhess) // GFNn-xTB (Input geometry)
--------------------------------------------------
CRE OPTIMIZATION - PART2
--------------------------------------------------
part2: on
program: orca
functional for part2: r2scan-3c
basis set for part2: def2-mTZVPP
using xTB-optimizer for optimization: on
using the new ensemble optimizer: on
optimize all conformers below this G_thr(opt,2) threshold: 2.5
spearmanthr: 0.940
optimization level in part2: lax
solvent model applied in the optimization: smd
solvent model for Gsolv contribution: smd
evaluate at different temperatures: on
Boltzmann sum threshold G_thr(2) for sorting in part2: 99.0
calculate mRRHO contribution: on
program for mRRHO contribution: xtb
GFN version for mRRHO and/or GBSA_Gsolv: gfn2
Apply constraint to input geometry during mRRHO calculation: on
solvent model applied with xTB: alpb
short-notation:
r2scan-3c + SMD[chcl3] + GmRRHO(GFN2[alpb]-bhess) // r2scan-3c[SMD]
--------------------------------------------------
NMR MODE SETTINGS
--------------------------------------------------
part4: on
calculate couplings (J): on
program for coupling calculations: orca
solvation model for coupling calculations: smd
functional for coupling calculation: pbe0
basis set for coupling calculation: def2-TZVP
calculate shieldings (S): on
program for shielding calculations: orca
solvation model for shielding calculations: smd
functional for shielding calculation: pbe0
basis set for shielding calculation: def2-TZVP
Calculating proton spectrum: on
reference for 1H: TMS
resonance frequency: 300.0
END of parameters
------------------------------------------------------------
PATHS of external QM programs
------------------------------------------------------------
The following program paths are used:
ORCA: /tmp1/orca_5_0_1_linux_x86-64_openmpi411
ORCA Version: 5.01
xTB: /home/abt-grimme/AK-bin/xtb
TURBOMOLE: /home/abt-grimme/TURBOMOLE.7.5//bin/em64t-unknown-linux-gnu_smp
Using cefine from /tmp/_MEI7bRtDs/cefine
PARNODES for TM or COSMO-RS calculation was set to 4
----------------------------------------------------------------------------------------------------
Processing data from previous run (enso.json)
----------------------------------------------------------------------------------------------------
Reading file: enso.json
Backing up enso.json to enso.json.1.
WARNING: setting basis_j was changed from def2-TZVPP to def2-TZVP!
WARNING: setting basis_s was changed from def2-TZVPP to def2-TZVP!
----------------------------------------------------------------------------------------------------
CRE CHEAP-PRESCREENING - PART0
----------------------------------------------------------------------------------------------------
program: orca
functional for part0: b97-d3
basis set for part0: def2-SV(P)
threshold g_thr(0): 4.0
starting number of considered conformers: 9
temperature: 298.15
Calculating efficient gas-phase single-point energies:
The efficient gas-phase single-point was calculated before for:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9
No conformers are considered additionally.
The efficient gas-phase single-point was successful for CONF1/r2scan-3c: E(DFT) = -235.43078946 Gsolv = -0.00918526
The efficient gas-phase single-point was successful for CONF2/r2scan-3c: E(DFT) = -235.43138285 Gsolv = -0.00887423
The efficient gas-phase single-point was successful for CONF3/r2scan-3c: E(DFT) = -235.43074850 Gsolv = -0.00917430
The efficient gas-phase single-point was successful for CONF4/r2scan-3c: E(DFT) = -235.42926214 Gsolv = -0.00889014
The efficient gas-phase single-point was successful for CONF5/r2scan-3c: E(DFT) = -235.42857572 Gsolv = -0.00921777
The efficient gas-phase single-point was successful for CONF6/r2scan-3c: E(DFT) = -235.42719486 Gsolv = -0.00896335
The efficient gas-phase single-point was successful for CONF7/r2scan-3c: E(DFT) = -235.42719983 Gsolv = -0.00896225
The efficient gas-phase single-point was successful for CONF8/r2scan-3c: E(DFT) = -235.42721003 Gsolv = -0.00896513
The efficient gas-phase single-point was successful for CONF9/r2scan-3c: E(DFT) = -235.42727703 Gsolv = -0.00896355
----------------------------------------------------------------------------------------------------
Removing high lying conformers by improved energy description
----------------------------------------------------------------------------------------------------
CONF# E [Eh] ΔE [kcal/mol] E [Eh] Gsolv [Eh] gtot ΔE(DFT) ΔGsolv Δgtot
GFN2-xTB GFN2-xTB b97-d3/def2-SV(P) alpb [Eh] [kcal/mol] [kcal/mol] [kcal/mol]
[alpb] [alpb] [gfn2]
CONF1 -18.9505968 0.00 -235.4307895 -0.0091853 -235.4399747 0.37 -0.20 0.18
CONF2 -18.9503605 0.15 -235.4313829 -0.0088742 -235.4402571 0.00 0.00 0.00 <------
CONF3 -18.9495194 0.68 -235.4307485 -0.0091743 -235.4399228 0.40 -0.19 0.21
CONF4 -18.9494414 0.73 -235.4292621 -0.0088901 -235.4381523 1.33 -0.01 1.32
CONF5 -18.9487831 1.14 -235.4285757 -0.0092178 -235.4377935 1.76 -0.22 1.55
CONF6 -18.9474716 1.96 -235.4271949 -0.0089633 -235.4361582 2.63 -0.06 2.57
CONF7 -18.9474509 1.97 -235.4271998 -0.0089622 -235.4361621 2.62 -0.06 2.57
CONF8 -18.9474396 1.98 -235.4272100 -0.0089651 -235.4361752 2.62 -0.06 2.56
CONF9 -18.9474298 1.99 -235.4272770 -0.0089635 -235.4362406 2.58 -0.06 2.52
----------------------------------------------------------------------------------------------------
Number of conformers observed within the following Δg windows:
Δg [kcal/mol] #CONF sum(Boltzmann_weights)
---------------------------------------------
0 - 0.5 3 0.91
0 - 1.0 3 0.91
0 - 1.5 4 0.95
0 - 2.0 5 0.98
0 - 2.5 5 0.98
0 - 3.0 9 1.00
---------------------------------------------
All relative (free) energies are below the initial g_thr(0) threshold of 4.0 kcal/mol.
All conformers are considered further.
Calculating Boltzmann averaged (free) energy of ensemble on input geometries (not DFT optimized)!
temperature /K: avE(T) /a.u. avG(T) /a.u.
----------------------------------------------------------------------------------------------------
298.15 -235.4308067 -235.4398576 <<==part0==
----------------------------------------------------------------------------------------------------
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>END of Part0<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
Ran part0 in 0.0364 seconds
----------------------------------------------------------------------------------------------------
CRE PRESCREENING - PART1
----------------------------------------------------------------------------------------------------
program: orca
functional for part1 and 2: r2scan-3c
basis set for part1 and 2: def2-mTZVPP
Solvent: chcl3
solvent model for Gsolv contribution: smd
threshold g_thr(1) and G_thr(1): 3.5
starting number of considered conformers: 9
calculate mRRHO contribution: on
program for mRRHO contribution: xtb
GFN version for mRRHO and/or GBSA_Gsolv: gfn2
Apply constraint to input geometry during mRRHO calculation: on
temperature: 298.15
Calculating single-point energies and solvation contribution (G_solv):
The prescreening_single-point was calculated before for:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9
No conformers are considered additionally.
prescreening_single-point calculation was successful for CONF1/r2scan-3c: -235.77440421
prescreening_single-point calculation was successful for CONF2/r2scan-3c: -235.77484560
prescreening_single-point calculation was successful for CONF3/r2scan-3c: -235.77497197
prescreening_single-point calculation was successful for CONF4/r2scan-3c: -235.77248195
prescreening_single-point calculation was successful for CONF5/r2scan-3c: -235.77264379
prescreening_single-point calculation was successful for CONF6/r2scan-3c: -235.77028863
prescreening_single-point calculation was successful for CONF7/r2scan-3c: -235.77038514
prescreening_single-point calculation was successful for CONF8/r2scan-3c: -235.77049552
prescreening_single-point calculation was successful for CONF9/r2scan-3c: -235.77064046
--------------------------------------------------
Removing high lying conformers
--------------------------------------------------
CONF# E(GFNn-xTB) ΔE(GFNn-xTB) E [Eh] Gsolv [Eh] gtot Δgtot
[a.u.] [kcal/mol] r2scan-3c incl. in E [Eh] [kcal/mol]
[SMD]
CONF1 -18.9492145 0.00 -235.7744042 0.0000000 -235.7744042 0.36
CONF2 -18.9489783 0.15 -235.7748456 0.0000000 -235.7748456 0.08
CONF3 -18.9481371 0.68 -235.7749720 0.0000000 -235.7749720 0.00 <------
CONF4 -18.9480601 0.72 -235.7724820 0.0000000 -235.7724820 1.56
CONF5 -18.9474005 1.14 -235.7726438 0.0000000 -235.7726438 1.46
CONF6 -18.9460883 1.96 -235.7702886 0.0000000 -235.7702886 2.94
CONF7 -18.9460685 1.97 -235.7703851 0.0000000 -235.7703851 2.88
CONF8 -18.9460572 1.98 -235.7704955 0.0000000 -235.7704955 2.81
CONF9 -18.9460478 1.99 -235.7706405 0.0000000 -235.7706405 2.72
All relative (free) energies are below the g_thr(1) threshold of 3.5 kcal/mol.
All conformers are considered further.
--------------------------------------------------
Calculating prescreening G_mRRHO with implicit solvation!
The prescreening G_mRRHO calculation was performed before for:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9
No conformers are considered additionally.
The prescreening G_mRRHO run @ c1 was successful for CONF1/rrho_part1: 0.13022286 S_rot(sym)= 0.0000000 using= 0.1302229
The prescreening G_mRRHO run @ c1 was successful for CONF2/rrho_part1: 0.13068166 S_rot(sym)= 0.0000000 using= 0.1306817
The prescreening G_mRRHO run @ cs was successful for CONF3/rrho_part1: 0.13042984 S_rot(sym)= 0.0000000 using= 0.1304298
The prescreening G_mRRHO run @ c1 was successful for CONF4/rrho_part1: 0.13100576 S_rot(sym)= 0.0000000 using= 0.1310058
The prescreening G_mRRHO run @ c1 was successful for CONF5/rrho_part1: 0.13054308 S_rot(sym)= 0.0000000 using= 0.1305431
The prescreening G_mRRHO run @ c1 was successful for CONF6/rrho_part1: 0.13055033 S_rot(sym)= 0.0000000 using= 0.1305503
The prescreening G_mRRHO run @ c1 was successful for CONF7/rrho_part1: 0.13056206 S_rot(sym)= 0.0000000 using= 0.1305621
The prescreening G_mRRHO run @ c1 was successful for CONF8/rrho_part1: 0.13036878 S_rot(sym)= 0.0000000 using= 0.1303688
The prescreening G_mRRHO run @ c1 was successful for CONF9/rrho_part1: 0.13054636 S_rot(sym)= 0.0000000 using= 0.1305464
--------------------------------------------------
* Gibbs free energies of part1 *
--------------------------------------------------
CONF# G(GFNn-xTB) ΔG(GFNn-xTB) E [Eh] Gsolv [Eh] GmRRHO [Eh] Gtot ΔGtot
[a.u.] [kcal/mol] r2scan-3c incl. in E GFN2 [Eh] [kcal/mol]
[SMD] [alpb]-bhess
CONF1 -18.8189917 0.00 -235.7744042 0.0000000 0.1302229 -235.6441813 0.23
CONF2 -18.8182966 0.44 -235.7748456 0.0000000 0.1306817 -235.6441639 0.24
CONF3 -18.8177073 0.81 -235.7749720 0.0000000 0.1304298 -235.6445421 0.00 <------
CONF4 -18.8170544 1.22 -235.7724820 0.0000000 0.1310058 -235.6414762 1.92
CONF5 -18.8168574 1.34 -235.7726438 0.0000000 0.1305431 -235.6421007 1.53
CONF6 -18.8155380 2.17 -235.7702886 0.0000000 0.1305503 -235.6397383 3.01
CONF7 -18.8155065 2.19 -235.7703851 0.0000000 0.1305621 -235.6398231 2.96
CONF8 -18.8156884 2.07 -235.7704955 0.0000000 0.1303688 -235.6401267 2.77
CONF9 -18.8155014 2.19 -235.7706405 0.0000000 0.1305464 -235.6400941 2.79
Number of conformers observed within the following ΔG windows:
ΔG [kcal/mol] #CONF sum(Boltzmann_weights)
---------------------------------------------
0 - 0.5 3 0.94
0 - 1.0 3 0.94
0 - 1.5 3 0.94
0 - 2.0 5 0.99
0 - 2.5 5 0.99
0 - 3.0 8 1.00
0 - 3.5 9 1.00
---------------------------------------------
Additional global 'fuzzy-threshold' based on the standard deviation of (G_mRRHO):
Std_dev(G_mRRHO) = 0.137 kcal/mol
Fuzzythreshold = 0.089 kcal/mol
Final sorting threshold G_thr(1) = 3.500 + 0.089 = 3.589 kcal/mol
Spearman correlation coefficient between (E + Solv) and (E + Solv + mRRHO) = 0.967
All relative (free) energies are below the initial G_thr(1) threshold of 3.5 kcal/mol.
All conformers are considered further.
Calculating Boltzmann averaged free energy of ensemble on input geometries (not DFT optimized)!
temperature /K: avE(T) /a.u. avGmRRHO(T) /a.u. avGsolv(T) /a.u. avG(T) /a.u.
----------------------------------------------------------------------------------------------------
298.15 -235.7746177 0.1304541 0.0000000 -235.6441637 <<==part1==
----------------------------------------------------------------------------------------------------
Backing up enso_ensemble_part1.xyz to enso_ensemble_part1.xyz.1.
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>END of Part1<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
Ran part1 in 0.0395 seconds
----------------------------------------------------------------------------------------------------
CRE OPTIMIZATION - PART2
----------------------------------------------------------------------------------------------------
program: orca
functional for part2: r2scan-3c
basis set for part2: def2-mTZVPP
using the xTB-optimizer for optimization: on
using the new ensemble optimizer: on
optimize all conformers below this G_thr(opt,2) threshold: 2.5
Spearman threshold: 0.940
number of optimization iterations: 8
radsize: 10
optimization level in part2: lax
solvent: chcl3
solvent model applied in the optimization: smd
solvent model for Gsolv contribution: smd
temperature: 298.15
evalulate at different temperatures: on
temperature range: 273.15, 278.15, 283.15, 288.15, ...
Boltzmann sum threshold G_thr(2) for sorting in part2: 99.0
calculate mRRHO contribution: on
program for mRRHO contribution: xtb
GFN version for mRRHO and/or GBSA_Gsolv: gfn2
Apply constraint to input geometry during mRRHO calculation: on
Optimizing geometries at DFT level with implicit solvation!
The optimization was performed before for:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9
No conformers are considered additionally.
Converged optimization for CONF1 after 6 cycles: -235.7754628
Converged optimization for CONF2 after 18 cycles: -235.7760704
Converged optimization for CONF3 after 3 cycles: -235.7759365
Converged optimization for CONF4 after 4 cycles: -235.7734469
Converged optimization for CONF5 after 18 cycles: -235.7758872
Converged optimization for CONF6 after 10 cycles: -235.7717864
Converged optimization for CONF7 after 9 cycles: -235.7718397
Converged optimization for CONF8 after 9 cycles: -235.7719344
Converged optimization for CONF9 after 9 cycles: -235.7720109
Calculating single-point energies and solvation contribution (G_solv)!
The low level gsolv calculation was calculated before for:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9
No conformers are considered additionally.
lowlevel single-point calculation was successful for CONF1/r2scan-3c: -235.7754631
lowlevel single-point calculation was successful for CONF2/r2scan-3c: -235.7760747
lowlevel single-point calculation was successful for CONF3/r2scan-3c: -235.7759375
lowlevel single-point calculation was successful for CONF4/r2scan-3c: -235.7734570
lowlevel single-point calculation was successful for CONF5/r2scan-3c: -235.7758872
lowlevel single-point calculation was successful for CONF6/r2scan-3c: -235.7717965
lowlevel single-point calculation was successful for CONF7/r2scan-3c: -235.7718474
lowlevel single-point calculation was successful for CONF8/r2scan-3c: -235.7719428
lowlevel single-point calculation was successful for CONF9/r2scan-3c: -235.7720165
Calculating lowlevel G_mRRHO with implicit solvation on DFT geometry!
The G_mRRHO calculation was performed before for:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9
No conformers are considered additionally.
The lowlevel G_mRRHO calculation @ c1 was successful for CONF1/rrho_part2: 0.13021490 S_rot(sym)= 0.0000000 using= 0.1302149
The lowlevel G_mRRHO calculation @ c1 was successful for CONF2/rrho_part2: 0.13074064 S_rot(sym)= 0.0000000 using= 0.1307406
The lowlevel G_mRRHO calculation @ cs was successful for CONF3/rrho_part2: 0.13042770 S_rot(sym)= 0.0000000 using= 0.1304277
The lowlevel G_mRRHO calculation @ c1 was successful for CONF4/rrho_part2: 0.13101639 S_rot(sym)= 0.0000000 using= 0.1310164
The lowlevel G_mRRHO calculation @ cs was successful for CONF5/rrho_part2: 0.13045448 S_rot(sym)= 0.0000000 using= 0.1304545
The lowlevel G_mRRHO calculation @ c1 was successful for CONF6/rrho_part2: 0.13093651 S_rot(sym)= 0.0000000 using= 0.1309365
The lowlevel G_mRRHO calculation @ c1 was successful for CONF7/rrho_part2: 0.13059532 S_rot(sym)= 0.0000000 using= 0.1305953
The lowlevel G_mRRHO calculation @ c1 was successful for CONF8/rrho_part2: 0.13076173 S_rot(sym)= 0.0000000 using= 0.1307617
The lowlevel G_mRRHO calculation @ c1 was successful for CONF9/rrho_part2: 0.13079724 S_rot(sym)= 0.0000000 using= 0.1307972
--------------------------------------------------
* Gibbs free energies of part2 *
--------------------------------------------------
CONF# E(GFNn-xTB) ΔE(GFNn-xTB) E [Eh] Gsolv [Eh] GmRRHO [Eh] Gtot ΔGtot Boltzmannweight
[a.u.] [kcal/mol] r2scan-3c incl. in E GFN2 [Eh] [kcal/mol] % at 298.15 K
[SMD] [alpb]-bhess
CONF1 -18.9492145 0.00 -235.7754631 0.0000000 0.1302149 -235.6452482 0.16 21.13
CONF2 -18.9489783 0.15 -235.7760747 0.0000000 0.1307406 -235.6453341 0.11 23.14
CONF3 -18.9481371 0.68 -235.7759375 0.0000000 0.1304277 -235.6455098 0.00 27.87 <------
CONF4 -18.9480601 0.72 -235.7734570 0.0000000 0.1310164 -235.6424406 1.93 1.08
CONF5 -18.9474005 1.14 -235.7758872 0.0000000 0.1304545 -235.6454328 0.05 25.69
CONF6 -18.9460883 1.96 -235.7717965 0.0000000 0.1309365 -235.6408600 2.92 0.20
CONF7 -18.9460685 1.97 -235.7718474 0.0000000 0.1305953 -235.6412521 2.67 0.31
CONF8 -18.9460572 1.98 -235.7719428 0.0000000 0.1307617 -235.6411810 2.72 0.28
CONF9 -18.9460478 1.99 -235.7720165 0.0000000 0.1307972 -235.6412192 2.69 0.30
Number of conformers observed within the following ΔG windows:
ΔG [kcal/mol] #CONF sum(Boltzmann_weights)
---------------------------------------------
0 - 0.5 4 0.98
0 - 1.0 4 0.98
0 - 1.5 4 0.98
0 - 2.0 5 0.99
0 - 2.5 5 0.99
0 - 3.0 9 1.00
---------------------------------------------
Calculating Boltzmann averaged free energy of ensemble!
temperature /K: avE(T) /a.u. avGmRRHO(T) /a.u. avGsolv(T) /a.u. avG(T) /a.u.
----------------------------------------------------------------------------------------------------
273.15 -235.7758086 0.1338393 0.0000000 -235.6419693
278.15 -235.7758041 0.1331736 0.0000000 -235.6426304
283.15 -235.7757995 0.1325044 0.0000000 -235.6432951
288.15 -235.7757950 0.1318308 0.0000000 -235.6439642
293.15 -235.7757902 0.1311531 0.0000000 -235.6446371
298.15 -235.7757854 0.1304720 0.0000000 -235.6453134 <<==part2==
303.15 -235.7757805 0.1297867 0.0000000 -235.6459938
308.15 -235.7757756 0.1290976 0.0000000 -235.6466779
313.15 -235.7757705 0.1284042 0.0000000 -235.6473663
318.15 -235.7757654 0.1277071 0.0000000 -235.6480583
323.15 -235.7757602 0.1270067 0.0000000 -235.6487536
328.15 -235.7757549 0.1263019 0.0000000 -235.6494530
333.15 -235.7757496 0.1255933 0.0000000 -235.6501563
338.15 -235.7757443 0.1248806 0.0000000 -235.6508637
343.15 -235.7757388 0.1241640 0.0000000 -235.6515748
348.15 -235.7757332 0.1234444 0.0000000 -235.6522888
353.15 -235.7757277 0.1227201 0.0000000 -235.6530076
358.15 -235.7757221 0.1219923 0.0000000 -235.6537299
363.15 -235.7757164 0.1212605 0.0000000 -235.6544559
368.15 -235.7757107 0.1205247 0.0000000 -235.6551860
373.15 -235.7757050 0.1197853 0.0000000 -235.6559197
----------------------------------------------------------------------------------------------------
--------------------------------------------------
Conformers considered further
--------------------------------------------------
Conformers that are below the Boltzmann threshold G_thr(2) of 99.0%:
CONF3, CONF5, CONF2, CONF1, CONF4
Backing up enso_ensemble_part2.xyz to enso_ensemble_part2.xyz.1.
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>END of Part2<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
Ran part2 in 0.0373 seconds
----------------------------------------------------------------------------------------------------
NMR MODE - PART4
----------------------------------------------------------------------------------------------------
calculate coupling constants: on
prog4J - program for coupling constant calculation: orca
funcJ - functional for coupling constant calculation: pbe0
basisJ - basis for coupling constant calculation: def2-TZVP
sm4J - solvent model for the coupling calculation: smd
calculate shielding constants σ: on
prog4S - program for shielding constant calculation: orca
funcS - functional for shielding constant calculation: pbe0
basisS - basis for shielding constant calculation: def2-TZVP
sm4S - solvent model for the shielding calculation: smd
Calculating proton spectrum: on
reference for 1H: TMS
spectrometer frequency: 300.0
Considering the following 5 conformers:
CONF1, CONF2, CONF3, CONF4, CONF5
--------------------------------------------------
* Gibbs free energies used in part4 *
--------------------------------------------------
CONF# E [Eh] Gsolv [Eh] GmRRHO [Eh] Gtot ΔGtot Boltzmannweight
r2scan-3c incl. in E GFN2 [Eh] [kcal/mol] % at 298.15 K
[SMD] [alpb]-bhess
CONF1 -235.7754631 0.0000000 0.1302149 -235.6452482 0.16 21.36
CONF2 -235.7760747 0.0000000 0.1307406 -235.6453341 0.11 23.39
CONF3 -235.7759375 0.0000000 0.1304277 -235.6455098 0.00 28.18 <------
CONF4 -235.7734570 0.0000000 0.1310164 -235.6424406 1.93 1.09
CONF5 -235.7758872 0.0000000 0.1304545 -235.6454328 0.05 25.97
Conformers that are below the Boltzmann-thr of 99.0:
CONF3, CONF5, CONF2, CONF1, CONF4
Constructed folders!
Performing coupling constant calculations:
Starting 5 coupling constants calculations
Running coupling calculation in CONF3/NMR
Running coupling calculation in CONF5/NMR
Running coupling calculation in CONF2/NMR
Running coupling calculation in CONF1/NMR
Running coupling calculation in CONF4/NMR
Tasks completed!
Coupling constant calculation was successful for CONF1/NMR
Coupling constant calculation was successful for CONF2/NMR
Coupling constant calculation was successful for CONF3/NMR
Coupling constant calculation was successful for CONF4/NMR
Coupling constant calculation was successful for CONF5/NMR
Performing shielding constant calculations:
Starting 5 shielding constants calculations
Running shielding calculation in CONF1/NMR
Running shielding calculation in CONF2/NMR
Running shielding calculation in CONF3/NMR
Running shielding calculation in CONF4/NMR
Running shielding calculation in CONF5/NMR
Tasks completed!
Shielding constant calculation was successful for CONF1/NMR
Shielding constant calculation was successful for CONF2/NMR
Shielding constant calculation was successful for CONF3/NMR
Shielding constant calculation was successful for CONF4/NMR
Shielding constant calculation was successful for CONF5/NMR
Generating file anmr_enso for processing with the ANMR program.
Writing .anmrrc!
ERROR: The reference absolute shielding constant for element h could not be found!
You have to edit the file .anmrrc by hand!
INFORMATION: The KeyError is: 'r2scan-3c'
Generating plain nmrprop.dat files for each populated conformer.
These files contain all calculated shielding and coupling constants.
The files can be read by ANMR using the keyword '-plain'.
Tasks completed!
Averaged shielding constants:
# in coord element σ(sigma) SD(σ based on SD Gsolv) SD(σ by 0.4 kcal/mol) shift σ_ref
---------------------------------------------------------------------------------------------------------
3 h 30.08 0.000000 0.011277 -30.08 0.000
7 h 26.60 0.000000 0.015161 -26.60 0.000
8 h 26.53 0.000000 0.006733 -26.53 0.000
10 h 29.76 0.000000 0.013050 -29.76 0.000
11 h 29.76 0.000000 0.013050 -29.76 0.000
12 h 29.76 0.000000 0.013050 -29.76 0.000
13 h 29.50 0.000000 0.033389 -29.50 0.000
14 h 29.50 0.000000 0.033389 -29.50 0.000
15 h 30.08 0.000000 0.011277 -30.08 0.000
16 h 30.71 0.000000 0.033284 -30.71 0.000
17 h 30.71 0.000000 0.033284 -30.71 0.000
18 h 30.71 0.000000 0.033284 -30.71 0.000
---------------------------------------------------------------------------------------------------------
# in coord element σ(sigma) min(σ)* CONFX max(σ)* CONFX Δ(max-min)
---------------------------------------------------------------------------------------------------------
3 h 30.08 30.01 CONF4 30.14 CONF1 0.13
7 h 26.60 26.52 CONF4 26.65 CONF5 0.13
8 h 26.53 26.32 CONF4 26.57 CONF1 0.24
10 h 29.76 29.70 CONF4 29.84 CONF2 0.13
11 h 29.76 29.70 CONF4 29.84 CONF2 0.13
12 h 29.76 29.70 CONF4 29.84 CONF2 0.13
13 h 29.50 29.32 CONF2 29.59 CONF5 0.26
14 h 29.50 29.32 CONF2 29.59 CONF5 0.26
15 h 30.08 30.01 CONF4 30.14 CONF1 0.13
16 h 30.71 30.64 CONF3 30.89 CONF2 0.25
17 h 30.71 30.64 CONF3 30.89 CONF2 0.25
18 h 30.71 30.64 CONF3 30.89 CONF2 0.25
---------------------------------------------------------------------------------------------------------
* min(σ) and max(σ) are averaged over the chemical equivalent atoms, but not Boltzmann weighted.
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>END of Part4<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
Ran part4 in 96.9462 seconds
Part : #conf time time (including restarts)
-----------------------------------------------------------------------
Input : 9 - -
Part0_all : 9 0.04 s 9.05 s
Part1_initial_sort : 9 0.00 s 21.65 s
Part1_all : 9 0.04 s 23.84 s
Part2_opt : 9 0.00 s 238.99 s
Part2_all : 5 0.04 s 250.17 s
Part4 : 5 96.95 s 252.84 s
-----------------------------------------------------------------------
All parts : - 97.06 s 535.90 s
CENSO all done!
$CENSO global configuration file: .censorc
$VERSION:1.1.2
ORCA: /home/$USER/orca_5_0_1_linux_x86-64_openmpi411
ORCA version: 5.0.1
GFN-xTB: /home/$USER/bin/xtb
CREST: /home/$USER/bin/crest
mpshift: /home/$USER/TURBOMOLE.7.5/bin/em64t-unknown-linux-gnu/mpshift
escf: /home/$USER/TURBOMOLE.7.5/bin/em64t-unknown-linux-gnu/escf
#COSMO-RS
ctd = BP_TZVP_C30_1601.ctd cdir = "/home/$USER/COSMOthermX16/COSMOtherm/CTDATA-FILES" ldir = "/home/$USER/COSMOthermX16/COSMOtherm/CTDATA-FILES"
$ENDPROGRAMS
$CRE SORTING SETTINGS:
$GENERAL SETTINGS:
nconf: all # ['all', 'number e.g. 10 up to all conformers']
charge: 0 # ['number e.g. 0']
unpaired: 0 # ['number e.g. 0']
solvent: gas # ['gas', 'acetone', 'acetonitrile', 'aniline', 'benzaldehyde', 'benzene', 'ccl4', '...']
prog_rrho: xtb # ['xtb']
temperature: 298.15 # ['temperature in K e.g. 298.15']
trange: [273.15, 378.15, 5] # ['temperature range [start, end, step]']
multitemp: on # ['on', 'off']
evaluate_rrho: on # ['on', 'off']
consider_sym: on # ['on', 'off']
bhess: on # ['on', 'off']
imagthr: automatic # ['automatic or e.g., -100 # in cm-1']
sthr: automatic # ['automatic or e.g., 50 # in cm-1']
scale: automatic # ['automatic or e.g., 1.0 ']
rmsdbias: off # ['on', 'off']
sm_rrho: alpb # ['alpb', 'gbsa']
progress: off # possibilities
check: on # ['on', 'off']
prog: tm # ['tm', 'orca']
func: r2scan-3c # ['b3-lyp', 'b3lyp', 'b3lyp-3c', 'b3lyp-d3', 'b3lyp-d3(0)', 'b3lyp-d4', 'b3lyp-nl', '...']
basis: automatic # ['automatic', 'def2-TZVP', 'def2-mSVP', 'def2-mSVP', 'def2-mSVP', 'def2-mSVP', '...']
maxthreads: 7 # ['number of threads e.g. 2']
omp: 4 # ['number cores per thread e.g. 4']
balance: off # possibilities
cosmorsparam: automatic # ['automatic', '12-fine', '12-normal', '13-fine', '13-normal', '14-fine', '...']
$PART0 - CHEAP-PRESCREENING - SETTINGS:
part0: on # ['on', 'off']
func0: b97-d # ['b3-lyp', 'b3lyp', 'b3lyp-3c', 'b3lyp-d3', 'b3lyp-d3(0)', 'b3lyp-d4', '...']
basis0: def2-SV(P) # ['automatic', 'def2-SV(P)', 'def2-TZVP', 'def2-mSVP', 'def2-mSVP', 'def2-mSVP', '...']
part0_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
part0_threshold: 4.0 # ['number e.g. 4.0']
$PART1 - PRESCREENING - SETTINGS:
# func and basis is set under GENERAL SETTINGS
part1: on # ['on', 'off']
smgsolv1: cosmors # ['alpb_gsolv', 'cosmo', 'cosmors', 'cosmors-fine', 'cpcm', 'dcosmors', '...']
part1_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
part1_threshold: 3.5 # ['number e.g. 5.0']
$PART2 - OPTIMIZATION - SETTINGS:
# func and basis is set under GENERAL SETTINGS
part2: on # ['on', 'off']
opt_limit: 2.5 # ['number e.g. 4.0']
sm2: default # ['cosmo', 'cpcm', 'dcosmors', 'default', 'smd']
smgsolv2: cosmors # ['alpb_gsolv', 'cosmo', 'cosmors', 'cosmors-fine', 'cpcm', 'dcosmors', '...']
part2_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
ancopt: on # ['on']
hlow: 0.01 # ['lowest force constant in ANC generation, e.g. 0.01']
opt_spearman: on # ['on', 'off']
part2_threshold: 99 # ['Boltzmann sum threshold in %. e.g. 95 (between 1 and 100)']
optlevel2: automatic # ['crude', 'sloppy', 'loose', 'lax', 'normal', 'tight', 'vtight', 'extreme', '...']
optcycles: 8 # ['number e.g. 5 or 10']
spearmanthr: -4.0 # ['value between -1 and 1, if outside set automatically']
radsize: 10 # ['number e.g. 8 or 10']
crestcheck: off # ['on', 'off']
$PART3 - REFINEMENT - SETTINGS:
part3: off # ['on', 'off']
prog3: prog # ['tm', 'orca', 'prog']
func3: pw6b95 # ['b3-lyp', 'b3lyp', 'b3lyp-3c', 'b3lyp-d3', 'b3lyp-d3(0)', 'b3lyp-d4', 'b3lyp-nl', '...']
basis3: def2-TZVPD # ['DZ', 'QZV', 'QZVP', 'QZVPP', 'SV(P)', 'SVP', 'TZVP', 'TZVPP', 'aug-cc-pV5Z', '...']
smgsolv3: cosmors # ['alpb_gsolv', 'cosmo', 'cosmors', 'cosmors-fine', 'cpcm', 'dcosmors', '...']
part3_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
part3_threshold: 99 # ['Boltzmann sum threshold in %. e.g. 95 (between 1 and 100)']
$NMR PROPERTY SETTINGS:
$PART4 SETTINGS:
part4: off # ['on', 'off']
couplings: on # ['on', 'off']
progJ: prog # ['tm', 'orca', 'adf', 'prog']
funcJ: pbe0 # ['b3-lyp', 'b3lyp', 'b3lyp-3c', 'b3lyp-d3', 'b3lyp-d3(0)', 'b3lyp-d4', 'b3lyp-nl', '...']
basisJ: def2-TZVP # ['DZ', 'QZV', 'QZVP', 'QZVPP', 'SV(P)', 'SVP', 'TZVP', 'TZVPP', 'aug-cc-pV5Z', '...']
sm4J: default # ['cosmo', 'cpcm', 'dcosmors', 'smd']
shieldings: on # ['on', 'off']
progS: prog # ['tm', 'orca', 'adf', 'prog']
funcS: pbe0 # ['b3-lyp', 'b3lyp', 'b3lyp-3c', 'b3lyp-d3', 'b3lyp-d3(0)', 'b3lyp-d4', 'b3lyp-nl', '...']
basisS: def2-TZVP # ['DZ', 'QZV', 'QZVP', 'QZVPP', 'SV(P)', 'SVP', 'TZVP', 'TZVPP', 'aug-cc-pV5Z', '...']
sm4S: default # ['cosmo', 'cpcm', 'dcosmors', 'smd']
reference_1H: TMS # ['TMS']
reference_13C: TMS # ['TMS']
reference_19F: CFCl3 # ['CFCl3']
reference_29Si: TMS # ['TMS']
reference_31P: TMP # ['TMP', 'PH3']
1H_active: on # ['on', 'off']
13C_active: on # ['on', 'off']
19F_active: off # ['on', 'off']
29Si_active: off # ['on', 'off']
31P_active: off # ['on', 'off']
resonance_frequency: 300.0 # ['MHz number of your experimental spectrometer setup']
$OPTICAL ROTATION PROPERTY SETTINGS:
$PART5 SETTINGS:
optical_rotation: off # ['on', 'off']
funcOR: pbe # ['functional for opt_rot e.g. pbe']
funcOR_SCF: r2scan-3c # ['functional for SCF in opt_rot e.g. r2scan-3c']
basisOR: def2-SVPD # ['basis set for opt_rot e.g. def2-SVPD']
frequency_optical_rot: [589.0] # ['list of freq in nm to evaluate opt rot at e.g. [589, 700]']
$END CENSORC
In our case CENSO printed an error-message that the reference absolute shielding constant at the level of
theory chosen is missing for hydrogen.
ERROR: The reference absolute shielding constant for element h could not be found!
You have to edit the file .anmrrc by hand!
To calculate it, the same calculation as for 2-methyl-1-pentene has to be performed for TMS in a new directory:
$ mkdir tms
$ cd tms
$ cat coord
$coord
2.05833045453195 -2.05833045453195 2.05833045453195 c
3.27901073396930 -3.27901073396930 0.93023223253204 h
3.27901073396930 -0.93023223253204 3.27901073396930 h
0.93023223253204 -3.27901073396930 3.27901073396930 h
-0.00000000000000 0.00000000000000 0.00000000000000 si
-2.05833045453195 2.05833045453195 2.05833045453195 c
-3.27901073396930 3.27901073396930 0.93023223253204 h
-0.93023223253204 3.27901073396930 3.27901073396930 h
-3.27901073396930 0.93023223253204 3.27901073396930 h
2.05833045453195 2.05833045453195 -2.05833045453195 c
0.93023223253204 3.27901073396930 -3.27901073396930 h
3.27901073396930 0.93023223253204 -3.27901073396930 h
3.27901073396930 3.27901073396930 -0.93023223253204 h
-2.05833045453195 -2.05833045453195 -2.05833045453195 c
-3.27901073396930 -3.27901073396930 -0.93023223253204 h
-3.27901073396930 -0.93023223253204 -3.27901073396930 h
-0.93023223253204 -3.27901073396930 -3.27901073396930 h
$end
$ crest coord -gfn2 -alpb chcl3 -T 4 -nmr > crest.out
$ mkdir censo
$ cp crest_conformers.xyz coord anmr_nucinfo anmr_rotamer censo/
$ cd censo/
$ censo --input crest_conformers.xyz -func0 b97-d3 -solvent chcl3 -smgsolv1 smd -sm2 smd
--smgsolv2 smd --prog orca -part4 on -prog4J orca -prog4S orca -funcJ pbe0
-funcS pbe0 -basisJ def2-TZVP -basisS def2-TZVP -cactive off > censo.out
______________________________________________________________
| |
| |
| CENSO - Commandline ENSO |
| v 1.1.2 |
| energetic sorting of CREST Conformer Rotamer Ensembles |
| University of Bonn, MCTC |
| Feb 2021 |
| based on ENSO version 2.0.1 |
| F. Bohle and S. Grimme |
| |
|______________________________________________________________|
Please cite:
S. Grimme, F. Bohle, A. Hansen, P. Pracht, S. Spicher, and M. Stahn
J. Phys. Chem. A 2021, 125, 19, 4039-4054.
DOI: https://doi.org/10.1021/acs.jpca.1c00971
This program is distributed in the hope that it will be useful,
but WITHOUT ANY WARRANTY; without even the implied warranty of
MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE.
----------------------------------------------------------------------------------------------------
PARAMETERS
----------------------------------------------------------------------------------------------------
program call: censo --input crest_conformers.xyz -func0 b97-d3 -solvent chcl3 -smgsolv1 smd -sm2 smd --smgsolv2 smd --prog orca -part4 on -prog4J orca -prog4S orca -funcJ pbe0 -funcS pbe0 -basisJ def2-TZVP -basisS def2-TZVP
The configuration file .censorc is read from /home/gorges/.censorc.
Reading conformer rotamer ensemble from: /tmp1/gorges/3881229.majestix.thch.uni-bonn.de/crest_conformers.xyz.
Reading file: censo_solvents.json
--------------------------------------------------
CRE SORTING SETTINGS
--------------------------------------------------
number of atoms in system: 17
number of considered conformers: 2
number of all conformers from input: 2
charge: 0
unpaired: 0
solvent: chcl3
temperature: 298.15
evaluate at different temperatures: on
temperature range: 273.15, 278.15, 283.15, 288.15, ...
calculate mRRHO contribution: on
consider symmetry for mRRHO contribution: on
cautious checking for error and failed calculations: on
checking the DFT-ensemble using CREST: off
maxthreads: 7
omp: 4
automatically balance maxthreads and omp: off
--------------------------------------------------
CRE CHEAP-PRESCREENING - PART0
--------------------------------------------------
part0: on
starting number of considered conformers: 2
program for part0: orca
functional for fast single-point: b97-d3
basis set for fast single-point: def2-SV(P)
threshold g_thr(0) for sorting in part0: 4.0
Solvent model used with xTB: alpb
short-notation:
b97-d3/def2-SV(P) // GFNn-xTB (Input geometry)
--------------------------------------------------
CRE PRESCREENING - PART1
--------------------------------------------------
part1: on
program for part1: orca
functional for initial evaluation: r2scan-3c
basis set for initial evaluation: def2-mTZVPP
calculate mRRHO contribution: on
program for mRRHO contribution: xtb
GFN version for mRRHO and/or GBSA_Gsolv: gfn2
Apply constraint to input geometry during mRRHO calculation: on
solvent model applied with xTB: alpb
evaluate at different temperatures: off
threshold g_thr(1) and G_thr(1) for sorting in part1: 3.5
solvent model for Gsolv contribution of part1: smd
short-notation:
r2scan-3c + SMD[chcl3] + GmRRHO(GFN2[alpb]-bhess) // GFNn-xTB (Input geometry)
--------------------------------------------------
CRE OPTIMIZATION - PART2
--------------------------------------------------
part2: on
program: orca
functional for part2: r2scan-3c
basis set for part2: def2-mTZVPP
using xTB-optimizer for optimization: on
using the new ensemble optimizer: on
optimize all conformers below this G_thr(opt,2) threshold: 2.5
spearmanthr: 0.941
optimization level in part2: lax
solvent model applied in the optimization: smd
solvent model for Gsolv contribution: smd
evaluate at different temperatures: on
Boltzmann sum threshold G_thr(2) for sorting in part2: 99.0
calculate mRRHO contribution: on
program for mRRHO contribution: xtb
GFN version for mRRHO and/or GBSA_Gsolv: gfn2
Apply constraint to input geometry during mRRHO calculation: on
solvent model applied with xTB: alpb
short-notation:
r2scan-3c + SMD[chcl3] + GmRRHO(GFN2[alpb]-bhess) // r2scan-3c[SMD]
--------------------------------------------------
NMR MODE SETTINGS
--------------------------------------------------
part4: on
calculate couplings (J): on
program for coupling calculations: orca
solvation model for coupling calculations: smd
functional for coupling calculation: pbe0
basis set for coupling calculation: def2-TZVP
calculate shieldings (S): on
program for shielding calculations: orca
solvation model for shielding calculations: smd
functional for shielding calculation: pbe0
basis set for shielding calculation: def2-TZVP
Calculating proton spectrum: on
reference for 1H: TMS
resonance frequency: 300.0
END of parameters
------------------------------------------------------------
PATHS of external QM programs
------------------------------------------------------------
The following program paths are used:
ORCA: /tmp1/orca_5_0_1_linux_x86-64_openmpi411
ORCA Version: 5.01
xTB: /home/abt-grimme/AK-bin/xtb
TURBOMOLE: /home/abt-grimme/TURBOMOLE.7.5//bin/em64t-unknown-linux-gnu_smp
Using cefine from /tmp/_MEIaCcz3S/cefine
PARNODES for TM or COSMO-RS calculation was set to 4
----------------------------------------------------------------------------------------------------
Processing data from previous run (enso.json)
----------------------------------------------------------------------------------------------------
INFORMATION: No restart information exists and is created during this run!
----------------------------------------------------------------------------------------------------
CRE CHEAP-PRESCREENING - PART0
----------------------------------------------------------------------------------------------------
program: orca
functional for part0: b97-d3
basis set for part0: def2-SV(P)
threshold g_thr(0): 4.0
starting number of considered conformers: 2
temperature: 298.15
Calculating efficient gas-phase single-point energies:
The efficient gas-phase single-point is calculated for:
CONF1, CONF2
Constructed folders!
Starting 2 ALPB-Gsolv calculations
Running single-point in CONF1/part0_sp
Running single-point in CONF2/part0_sp
Running ALPB_GSOLV calculation in 3881229.majestix.thch.uni-bonn.de/CONF2/part0_sp
Running ALPB_GSOLV calculation in 3881229.majestix.thch.uni-bonn.de/CONF1/part0_sp
Tasks completed!
The efficient gas-phase single-point was successful for CONF1/part0_sp: E(DFT) = -448.78335711 Gsolv = -0.00964593
The efficient gas-phase single-point was successful for CONF2/part0_sp: E(DFT) = -448.78106942 Gsolv = -0.00949351
----------------------------------------------------------------------------------------------------
Removing high lying conformers by improved energy description
----------------------------------------------------------------------------------------------------
CONF# E [Eh] ΔE [kcal/mol] E [Eh] Gsolv [Eh] gtot ΔE(DFT) ΔGsolv Δgtot
GFN2-xTB GFN2-xTB b97-d3/def2-SV(P) alpb [Eh] [kcal/mol] [kcal/mol] [kcal/mol]
[alpb] [alpb] [gfn2]
CONF1 -16.3966231 0.00 -448.7833571 -0.0096459 -448.7930030 0.00 0.00 0.00 <------
CONF2 -16.3954819 0.72 -448.7810694 -0.0094935 -448.7905629 1.44 0.10 1.53
----------------------------------------------------------------------------------------------------
Number of conformers observed within the following Δg windows:
Δg [kcal/mol] #CONF sum(Boltzmann_weights)
---------------------------------------------
0 - 0.5 1 0.93
0 - 1.0 1 0.93
0 - 1.5 1 0.93
0 - 2.0 2 1.00
---------------------------------------------
All relative (free) energies are below the initial g_thr(0) threshold of 4.0 kcal/mol.
All conformers are considered further.
Calculating Boltzmann averaged (free) energy of ensemble on input geometries (not DFT optimized)!
temperature /K: avE(T) /a.u. avG(T) /a.u.
----------------------------------------------------------------------------------------------------
298.15 -448.7831966 -448.7928319 <<==part0==
----------------------------------------------------------------------------------------------------
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>END of Part0<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
Ran part0 in 4.4449 seconds
----------------------------------------------------------------------------------------------------
CRE PRESCREENING - PART1
----------------------------------------------------------------------------------------------------
program: orca
functional for part1 and 2: r2scan-3c
basis set for part1 and 2: def2-mTZVPP
Solvent: chcl3
solvent model for Gsolv contribution: smd
threshold g_thr(1) and G_thr(1): 3.5
starting number of considered conformers: 2
calculate mRRHO contribution: on
program for mRRHO contribution: xtb
GFN version for mRRHO and/or GBSA_Gsolv: gfn2
Apply constraint to input geometry during mRRHO calculation: on
temperature: 298.15
Calculating single-point energies and solvation contribution (G_solv):
The prescreening_single-point is calculated for:
CONF1, CONF2
Constructed folders!
Running single-point in CONF1/r2scan-3c
Running single-point in CONF2/r2scan-3c
Tasks completed!
prescreening_single-point calculation was successful for CONF1/r2scan-3c: -449.11349203
prescreening_single-point calculation was successful for CONF2/r2scan-3c: -449.11143980
--------------------------------------------------
Removing high lying conformers
--------------------------------------------------
CONF# E(GFNn-xTB) ΔE(GFNn-xTB) E [Eh] Gsolv [Eh] gtot Δgtot
[a.u.] [kcal/mol] r2scan-3c incl. in E [Eh] [kcal/mol]
[SMD]
CONF1 -16.3952414 0.00 -449.1134920 0.0000000 -449.1134920 0.00 <------
CONF2 -16.3940994 0.72 -449.1114398 0.0000000 -449.1114398 1.29
All relative (free) energies are below the g_thr(1) threshold of 3.5 kcal/mol.
All conformers are considered further.
--------------------------------------------------
Calculating prescreening G_mRRHO with implicit solvation!
The prescreening G_mRRHO calculation is now performed for:
CONF1, CONF2
Constructed folders!
Starting 2 G_RRHO calculations.
Running GFN2-xTB mRRHO in CONF1/rrho_part1
Running GFN2-xTB mRRHO in CONF2/rrho_part1
WARNING: found 1 significant imaginary frequencies in CONF2/rrho_part1
Tasks completed!
The prescreening G_mRRHO run @ td was successful for CONF1/rrho_part1: 0.11573915 S_rot(sym)= 0.0023462 using= 0.1157391
The prescreening G_mRRHO run @ c3v was successful for CONF2/rrho_part1: 0.11540064 S_rot(sym)= 0.0010373 using= 0.1154006
--------------------------------------------------
* Gibbs free energies of part1 *
--------------------------------------------------
CONF# G(GFNn-xTB) ΔG(GFNn-xTB) E [Eh] Gsolv [Eh] GmRRHO [Eh] Gtot ΔGtot
[a.u.] [kcal/mol] r2scan-3c incl. in E GFN2 [Eh] [kcal/mol]
[SMD] [alpb]-bhess
CONF1 -16.2795022 0.00 -449.1134920 0.0000000 0.1157391 -448.9977529 0.00 <------
CONF2 -16.2786988 0.50 -449.1114398 0.0000000 0.1154006 -448.9960392 1.08
Number of conformers observed within the following ΔG windows:
ΔG [kcal/mol] #CONF sum(Boltzmann_weights)
---------------------------------------------
0 - 0.5 1 0.86
0 - 1.0 1 0.86
0 - 1.5 2 1.00
---------------------------------------------
Additional global 'fuzzy-threshold' based on the standard deviation of (G_mRRHO):
Std_dev(G_mRRHO) = 0.150 kcal/mol
Fuzzythreshold = 0.107 kcal/mol
Final sorting threshold G_thr(1) = 3.500 + 0.107 = 3.607 kcal/mol
Spearman correlation coefficient between (E + Solv) and (E + Solv + mRRHO) = 1.000
All relative (free) energies are below the initial G_thr(1) threshold of 3.5 kcal/mol.
All conformers are considered further.
Calculating Boltzmann averaged free energy of ensemble on input geometries (not DFT optimized)!
temperature /K: avE(T) /a.u. avGmRRHO(T) /a.u. avGsolv(T) /a.u. avG(T) /a.u.
----------------------------------------------------------------------------------------------------
298.15 -449.1132047 0.1156917 0.0000000 -448.9975129 <<==part1==
----------------------------------------------------------------------------------------------------
Calculating unbiased GFNn-xTB energy
Constructed folders!
Starting 2 xTB - single-point calculations.
gfn2-xTB energy for CONF1/GFN_unbiased = -16.3966231
gfn2-xTB energy for CONF2/GFN_unbiased = -16.3954819
Tasks completed!
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>END of Part1<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
Ran part1 in 9.1081 seconds
----------------------------------------------------------------------------------------------------
CRE OPTIMIZATION - PART2
----------------------------------------------------------------------------------------------------
program: orca
functional for part2: r2scan-3c
basis set for part2: def2-mTZVPP
using the xTB-optimizer for optimization: on
using the new ensemble optimizer: on
optimize all conformers below this G_thr(opt,2) threshold: 2.5
Spearman threshold: 0.941
number of optimization iterations: 8
radsize: 10
optimization level in part2: lax
solvent: chcl3
solvent model applied in the optimization: smd
solvent model for Gsolv contribution: smd
temperature: 298.15
evalulate at different temperatures: on
temperature range: 273.15, 278.15, 283.15, 288.15, ...
Boltzmann sum threshold G_thr(2) for sorting in part2: 99.0
calculate mRRHO contribution: on
program for mRRHO contribution: xtb
GFN version for mRRHO and/or GBSA_Gsolv: gfn2
Apply constraint to input geometry during mRRHO calculation: on
Optimizing geometries at DFT level with implicit solvation!
The optimization is calculated for:
CONF1, CONF2
Constructed folders!
Preparing 2 calculations.
Tasks completed!
************************Starting optimizations************************
Starting threshold is set to 2.5 + 60.0 % = 4.0 kcal/mol
Lower limit is set to G_thr(opt,2) = 2.5 kcal/mol
*******************************CYCLE 1********************************
Starting 2 optimizations.
Running optimization in CONF1/r2scan-3c
Running optimization in CONF2/r2scan-3c
Tasks completed!
Geometry optimization converged for: CONF1 within 3 cycles
Geometry optimization converged for: CONF2 within 3 cycles
Constructed folders!
Starting 2 G_RRHO calculations.
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Tasks completed!
The G_mRRHO calculation on crudely optimized DFT geometry @ td was successful for CONF1/r2scan-3c/rrho_crude: 0.1157450 S_rot(sym)= 0.0023462 using= 0.1157450
The G_mRRHO calculation on crudely optimized DFT geometry @ c3v was successful for CONF2/r2scan-3c/rrho_crude: 0.1146578 S_rot(sym)= 0.0010373 using= 0.1146578
***********************Finished optimizations!************************
Timings:
Cycle: [s] #nconfs Spearman coeff.
1 34.85 2
sum: 34.85
CONVERGED optimizations for the following remaining conformers:
Converged optimization for CONF1 after 3 cycles: -449.1155065
Converged optimization for CONF2 after 3 cycles: -449.1133322
Calculating single-point energies and solvation contribution (G_solv)!
CONF1, CONF2
Running single-point in CONF1/r2scan-3c
Running single-point in CONF2/r2scan-3c
Tasks completed!
lowlevel single-point calculation was successful for CONF1/r2scan-3c: -449.11550627
lowlevel single-point calculation was successful for CONF2/r2scan-3c: -449.11333157
Calculating lowlevel G_mRRHO with implicit solvation on DFT geometry!
The lowlevel G_mRRHO calculation is now performed for:
CONF1, CONF2
Constructed folders!
Starting 2 G_RRHO calculations.
Running GFN2-xTB mRRHO in CONF1/rrho_part2
Running GFN2-xTB mRRHO in CONF2/rrho_part2
Tasks completed!
The lowlevel G_mRRHO calculation @ td was successful for CONF1/rrho_part2: 0.11574502 S_rot(sym)= 0.0023462 using= 0.1157450
The lowlevel G_mRRHO calculation @ c3v was successful for CONF2/rrho_part2: 0.11465779 S_rot(sym)= 0.0010373 using= 0.1146578
--------------------------------------------------
* Gibbs free energies of part2 *
--------------------------------------------------
CONF# E(GFNn-xTB) ΔE(GFNn-xTB) E [Eh] Gsolv [Eh] GmRRHO [Eh] Gtot ΔGtot Boltzmannweight
[a.u.] [kcal/mol] r2scan-3c incl. in E GFN2 [Eh] [kcal/mol] % at 298.15 K
[SMD] [alpb]-bhess
CONF1 -16.3952414 0.00 -449.1155063 0.0000000 0.1157450 -448.9997612 0.00 75.98 <------
CONF2 -16.3940994 0.72 -449.1133316 0.0000000 0.1146578 -448.9986738 0.68 24.02
Number of conformers observed within the following ΔG windows:
ΔG [kcal/mol] #CONF sum(Boltzmann_weights)
---------------------------------------------
0 - 0.5 1 0.76
0 - 1.0 2 1.00
---------------------------------------------
Calculating Boltzmann averaged free energy of ensemble!
temperature /K: avE(T) /a.u. avGmRRHO(T) /a.u. avGsolv(T) /a.u. avG(T) /a.u.
----------------------------------------------------------------------------------------------------
273.15 -449.1150651 0.1190010 0.0000000 -448.9960641
278.15 -449.1150486 0.1183075 0.0000000 -448.9967412
283.15 -449.1150323 0.1176087 0.0000000 -448.9974236
288.15 -449.1150158 0.1169054 0.0000000 -448.9981104
293.15 -449.1149998 0.1161973 0.0000000 -448.9988025
298.15 -449.1149840 0.1154839 0.0000000 -448.9995001 <<==part2==
303.15 -449.1149681 0.1147661 0.0000000 -449.0002020
308.15 -449.1149528 0.1140434 0.0000000 -449.0009094
313.15 -449.1149376 0.1133166 0.0000000 -449.0016210
318.15 -449.1149227 0.1125847 0.0000000 -449.0023381
323.15 -449.1149077 0.1118481 0.0000000 -449.0030596
328.15 -449.1148932 0.1111069 0.0000000 -449.0037863
333.15 -449.1148790 0.1103615 0.0000000 -449.0045175
338.15 -449.1148646 0.1096115 0.0000000 -449.0052531
343.15 -449.1148509 0.1088570 0.0000000 -449.0059939
348.15 -449.1148374 0.1080973 0.0000000 -449.0067401
353.15 -449.1148237 0.1073340 0.0000000 -449.0074897
358.15 -449.1148107 0.1065661 0.0000000 -449.0082446
363.15 -449.1147975 0.1057936 0.0000000 -449.0090039
368.15 -449.1147853 0.1050171 0.0000000 -449.0097682
373.15 -449.1147730 0.1042360 0.0000000 -449.0105370
----------------------------------------------------------------------------------------------------
--------------------------------------------------
Conformers considered further
--------------------------------------------------
Conformers that are below the Boltzmann threshold G_thr(2) of 99.0%:
CONF1, CONF2
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>END of Part2<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
Ran part2 in 39.8941 seconds
----------------------------------------------------------------------------------------------------
NMR MODE - PART4
----------------------------------------------------------------------------------------------------
calculate coupling constants: on
prog4J - program for coupling constant calculation: orca
funcJ - functional for coupling constant calculation: pbe0
basisJ - basis for coupling constant calculation: def2-TZVP
sm4J - solvent model for the coupling calculation: smd
calculate shielding constants σ: on
prog4S - program for shielding constant calculation: orca
funcS - functional for shielding constant calculation: pbe0
basisS - basis for shielding constant calculation: def2-TZVP
sm4S - solvent model for the shielding calculation: smd
Calculating proton spectrum: on
reference for 1H: TMS
spectrometer frequency: 300.0
Considering the following 2 conformers:
CONF1, CONF2
--------------------------------------------------
* Gibbs free energies used in part4 *
--------------------------------------------------
CONF# E [Eh] Gsolv [Eh] GmRRHO [Eh] Gtot ΔGtot Boltzmannweight
r2scan-3c incl. in E GFN2 [Eh] [kcal/mol] % at 298.15 K
[SMD] [alpb]-bhess
CONF1 -449.1155063 0.0000000 0.1157450 -448.9997612 0.00 75.98 <------
CONF2 -449.1133316 0.0000000 0.1146578 -448.9986738 0.68 24.02
Conformers that are below the Boltzmann-thr of 99.0:
CONF1, CONF2
Constructed folders!
Performing coupling constant calculations:
Starting 2 coupling constants calculations
Running coupling calculation in CONF1/NMR
Running coupling calculation in CONF2/NMR
Tasks completed!
Coupling constant calculation was successful for CONF1/NMR
Coupling constant calculation was successful for CONF2/NMR
Performing shielding constant calculations:
Starting 2 shielding constants calculations
Running shielding calculation in CONF1/NMR
Running shielding calculation in CONF2/NMR
Tasks completed!
Shielding constant calculation was successful for CONF1/NMR
Shielding constant calculation was successful for CONF2/NMR
Generating file anmr_enso for processing with the ANMR program.
Writing .anmrrc!
ERROR: The reference absolute shielding constant for element h could not be found!
You have to edit the file .anmrrc by hand!
INFORMATION: The KeyError is: 'r2scan-3c'
Generating plain nmrprop.dat files for each populated conformer.
These files contain all calculated shielding and coupling constants.
The files can be read by ANMR using the keyword '-plain'.
Tasks completed!
Averaged shielding constants:
# in coord element σ(sigma) SD(σ based on SD Gsolv) SD(σ by 0.4 kcal/mol) shift σ_ref
---------------------------------------------------------------------------------------------------------
2 h 31.59 0.000000 0.002263 -31.59 0.000
3 h 31.59 0.000000 0.002263 -31.59 0.000
4 h 31.59 0.000000 0.002263 -31.59 0.000
7 h 31.59 0.000000 0.002263 -31.59 0.000
8 h 31.59 0.000000 0.002263 -31.59 0.000
9 h 31.59 0.000000 0.002263 -31.59 0.000
11 h 31.59 0.000000 0.002263 -31.59 0.000
12 h 31.59 0.000000 0.002263 -31.59 0.000
13 h 31.59 0.000000 0.002263 -31.59 0.000
15 h 31.59 0.000000 0.002263 -31.59 0.000
16 h 31.59 0.000000 0.002263 -31.59 0.000
17 h 31.59 0.000000 0.002263 -31.59 0.000
---------------------------------------------------------------------------------------------------------
# in coord element σ(sigma) min(σ)* CONFX max(σ)* CONFX Δ(max-min)
---------------------------------------------------------------------------------------------------------
2 h 31.59 31.58 CONF2 31.60 CONF1 0.01
3 h 31.59 31.58 CONF2 31.60 CONF1 0.01
4 h 31.59 31.58 CONF2 31.60 CONF1 0.01
7 h 31.59 31.58 CONF2 31.60 CONF1 0.01
8 h 31.59 31.58 CONF2 31.60 CONF1 0.01
9 h 31.59 31.58 CONF2 31.60 CONF1 0.01
11 h 31.59 31.58 CONF2 31.60 CONF1 0.01
12 h 31.59 31.58 CONF2 31.60 CONF1 0.01
13 h 31.59 31.58 CONF2 31.60 CONF1 0.01
15 h 31.59 31.58 CONF2 31.60 CONF1 0.01
16 h 31.59 31.58 CONF2 31.60 CONF1 0.01
17 h 31.59 31.58 CONF2 31.60 CONF1 0.01
---------------------------------------------------------------------------------------------------------
* min(σ) and max(σ) are averaged over the chemical equivalent atoms, but not Boltzmann weighted.
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>END of Part4<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
Ran part4 in 66.5507 seconds
Part : #conf time time (including restarts)
-----------------------------------------------------------------------
Input : 2 - -
Part0_all : 2 4.44 s 4.44 s
Part1_initial_sort : 2 8.03 s 8.03 s
Part1_all : 2 9.11 s 9.11 s
Part2_opt : 2 34.85 s 34.85 s
Part2_all : 2 39.89 s 39.89 s
Part4 : 2 66.55 s 66.55 s
-----------------------------------------------------------------------
All parts : - 120.00 s 120.00 s
CENSO all done!
The calculated shift has now to be inserted into the .anmrrc file of the NMR-calculation for the respective molecule:
$ cat .anmrrc
7 8 XH acid atoms
ENSO qm= ORCA mf= 300.0 lw= 1.0 J= on S= on T= 298.15
TMS[chcl3] pbe0[SMD]/def2-TZVP//r2scan-3c[SMD]/def2-mTZVPP
1 31.59 0.0 1
Now all information is present and ANMR can be called to calculate the full
NMR spectrum at 400 MHz:
$ anmr --mf 400 --plain > anmr.out 2> error.anmr &
+--------------------------------------+
| A N M R |
| S. Grimme |
| Universitaet Bonn, MCTC |
| 1989-2019 |
| version 3.5.1 |
| Sat Feb 9 06:41:57 CET 2019 |
+--------------------------------------+
Based on a TurboPascal program written
in 1989 which was translated to F90 in
2005 and re-activated in 2017.
Please cite work employing this code as:
ANMR Ver. 3.5: An automatic, QC based
coupled NMR spectra simulation program.
S. Grimme, Universitaet Bonn, 2019
S. Grimme, C. Bannwarth, S. Dohm, A. Hansen
J. Pisarek, P. Pracht, J. Seibert, F. Neese
Angew. Chem. Int. Ed. 2017, 56, 14763-14769.
DOI:10.1002/anie.201708266
=============================
# OMP threads = 4
=============================
reading <.anmrrc> for standard data
1 31.3600000000000 0.000000000000000E+000 1
1H resonance frequency (-mf <real>) : 400.00
line width (-lw <real>) : 1.00
number of all conformers : 5
temperature in K : 298.15
remove J couplings to OH groups : T
maximum spin system size in a fragment :14
fragmentation type (0=none,1=at,2=mol) : 2
chemical shift scalings a,b : 1.00 0.00
spin-spin coupling scal factor : 1.07
plot offset : 0.00
Active nuclei :H
reading from anmr_enso
conformational energies: 5
0.000 -0.384 -0.298 1.259 -0.266
conformational RRHO energies:
0.000 0.330 0.134 0.503 0.150
conformational Gsolv free energies:
0.000 0.000 0.000 0.000 0.000
conformational free energies:
0.000 -0.054 -0.164 1.762 -0.116
gi per conformer:
1.00 1.00 1.00 1.00 1.00
conformational populations calculated by anmr:
0.214 0.234 0.282 0.011 0.260
conformational populations calculated by enso:
These are used:
0.214 0.234 0.282 0.011 0.260
1( 1) 21.4
2( 2) 23.4
3( 3) 28.2
4( 4) 1.1
5( 5) 26.0
ensemble average free energy (Eh) : -235.621795
ensemble entropy (cal/mol K) : 2.833
ensemble total free energy (kcal/mol) : -235.622196
reading rotamer data from <anmr_rotamer>
number of unique conformers from CREST 9
number of unique conformers from enso 5
conformer 1 averaging over 18 rotamers
conformer 2 averaging over 18 rotamers
conformer 3 averaging over 9 rotamers
conformer 4 averaging over 18 rotamers
conformer 5 averaging over 3 rotamers
conformer 6 averaging over 2 rotamers
conformer 6 not in anmr_enso, skipping
conformer 7 averaging over 3 rotamers
conformer 7 not in anmr_enso, skipping
conformer 8 averaging over 2 rotamers
conformer 8 not in anmr_enso, skipping
conformer 9 averaging over 7 rotamers
conformer 9 not in anmr_enso, skipping
average over 66 in CRE
====================================================
reading J/sigma data for conformer 1
====================================================
reading from CONF1/NMR/nmrprop.dat
reading from CONF1/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 3 1 1.429
2 7 1 5.027
3 8 1 5.030
4 10 3 1.857
5 13 1 2.060
6 14 1 2.060
7 15 1 1.429
8 16 3 0.939
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 0.09252 0.00000
3 0.13509 2.69100 0.00000
4 0.05900 -1.32779 -1.75345 0.00000
5 3.89576 -0.90653 -0.11142 -0.18086 0.00000
6 11.33430 -0.94647 -0.11658 -0.26594 -11.23900 0.00000
7 -12.11500 0.09348 0.13391 0.05167 11.34670 3.86524
8 6.59230 0.01232 0.04920 0.01920 -0.28147 -0.28276
7 8
7 0.00000
8 6.31838 0.00000
writing spin system data for this confomer to tmpanmr.1
====================================================
reading J/sigma data for conformer 2
====================================================
reading from CONF2/NMR/nmrprop.dat
reading from CONF2/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 3 1 1.535
2 7 1 5.059
3 8 1 5.087
4 10 3 1.763
5 13 1 2.257
6 14 1 2.257
7 15 1 1.535
8 16 3 0.697
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 0.10988 0.00000
3 0.22973 2.89300 0.00000
4 0.02594 -1.27612 -1.68436 0.00000
5 7.86085 -1.23898 -0.79352 -0.41233 0.00000
6 3.58185 -1.36002 -0.86348 -0.52688 -12.63300 0.00000
7 -13.12500 0.11312 0.26527 0.02106 3.58715 7.61915
8 6.61576 0.02486 0.01738 0.01347 -0.11732 -0.06556
7 8
7 0.00000
8 7.04988 0.00000
writing spin system data for this confomer to tmpanmr.2
====================================================
reading J/sigma data for conformer 3
====================================================
reading from CONF3/NMR/nmrprop.dat
reading from CONF3/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 3 1 1.528
2 7 1 4.955
3 8 1 5.062
4 10 3 1.858
5 13 1 2.008
6 14 1 2.008
7 15 1 1.528
8 16 3 0.952
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -0.10764 0.00000
3 0.05673 2.03300 0.00000
4 0.07595 -1.27211 -1.68368 0.00000
5 3.43542 -2.48836 -2.32770 -1.08276 0.00000
6 11.07761 -2.48764 -2.32730 -1.08248 -18.04000 0.00000
7 -11.83800 -0.10636 0.05927 0.07650 11.07839 3.43658
8 6.77197 0.05314 0.06095 0.02536 -0.27921 -0.27614
7 8
7 0.00000
8 6.63456 0.00000
writing spin system data for this confomer to tmpanmr.3
====================================================
reading J/sigma data for conformer 4
====================================================
reading from CONF4/NMR/nmrprop.dat
reading from CONF4/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 3 1 1.568
2 7 1 5.080
3 8 1 5.275
4 10 3 1.891
5 13 1 2.069
6 14 1 2.069
7 15 1 1.568
8 16 3 0.834
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -0.07767 0.00000
3 -0.01136 2.19300 0.00000
4 0.07832 -1.57617 -1.91938 0.00000
5 7.43188 -0.97458 -0.19076 -0.40014 0.00000
6 3.50479 -0.86742 -0.08124 -0.29695 -12.25400 0.00000
7 -12.31900 -0.07433 0.00036 0.09255 3.52021 7.66912
8 6.68971 -0.01191 -0.01468 0.00885 -0.01423 -0.08317
7 8
7 0.00000
8 6.38153 0.00000
writing spin system data for this confomer to tmpanmr.4
====================================================
reading J/sigma data for conformer 5
====================================================
reading from CONF5/NMR/nmrprop.dat
reading from CONF5/NMR/nmrprop.dat
# in coord file # nucs delta(ppm)
1 3 1 1.509
2 7 1 4.949
3 8 1 5.068
4 10 3 1.850
5 13 1 1.991
6 14 1 1.991
7 15 1 1.509
8 16 3 0.931
MATRIX PRINTED: rotamer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -0.08970 0.00000
3 0.06455 2.08300 0.00000
4 0.08418 -1.30420 -1.74112 0.00000
5 3.43661 -2.49568 -2.31700 -1.10064 0.00000
6 11.06903 -2.47732 -2.30300 -1.10694 -18.09000 0.00000
7 -11.95000 -0.08630 0.06545 0.08495 11.06497 3.43539
8 6.59808 0.05759 0.06609 0.02891 -0.26317 -0.26839
7 8
7 0.00000
8 6.34345 0.00000
writing spin system data for this confomer to tmpanmr.5
=====================================================
average over conformers
=====================================================
conformer 1 (normalized) population : 0.214
conformer 2 (normalized) population : 0.234
conformer 3 (normalized) population : 0.282
conformer 4 (normalized) population : 0.011
conformer 5 (normalized) population : 0.260
# in coord file # nucs delta(ppm)
1 3 1 1.504 +/- 0.02
2 7 1 4.995 +/- 0.02
3 8 1 5.065 +/- 0.01
4 10 3 1.834 +/- 0.02
5 13 1 2.074 +/- 0.05
6 14 1 2.074 +/- 0.05
7 15 1 1.504 +/- 0.02
8 16 3 0.883 +/- 0.05
MATRIX PRINTED: conformer average J (Hz) matrix
1 2 3 4 5 6
1 0.00000
2 -0.00900 0.00000
3 0.11520 2.38947 0.00000
4 0.06280 -1.29659 -1.71625 0.00000
5 4.61283 -1.84360 -1.46929 -0.73047 0.00000
6 9.29421 -1.87428 -1.48183 -0.77586 -15.27225 0.00000
7 -12.23257 -0.00678 0.12439 0.06062 9.29747 4.55239
8 6.65101 0.03825 0.04878 0.02202 -0.23477 -0.22419
7 8
7 0.00000
8 6.58584 0.00000
====================================================
building sub-systems
====================================================
J neighbors of spin 1 : 3 5 6 7 8
J neighbors of spin 2 : 3 4 5 6
J neighbors of spin 3 : 1 2 4 5 6 7
J neighbors of spin 4 : 2 3 5 6
J neighbors of spin 5 : 1 2 3 4 6 7 8
J neighbors of spin 6 : 1 2 3 4 5 7 8
J neighbors of spin 7 : 1 3 5 6 8
J neighbors of spin 8 : 1 5 6 7
1:14 max/av. J neglected : 0.000 0.000
2:13 max/av. J neglected : 0.000 0.000
3:12 max/av. J neglected : 0.000 0.000
4:11 max/av. J neglected : 0.000 0.000
5:10 max/av. J neglected : 0.000 0.000
6: 9 max/av. J neglected : 0.000 0.000
7: 8 max/av. J neglected : 0.000 0.000
8: 7 max/av. J neglected : 0.063 0.031
9: 6 max/av. J neglected : 0.115 0.059
selected scheme 7
system largest neglected JAB nuclei (dummy atoms are negative)
1 0.000 1 2 3 4 5 6 7 8
=== FRAGMENTED SYSTEM ===
====================================================
solving (J/sigma) averaged spin Hamiltonian
====================================================
spinsystem 1 with 8 spins
1024 product functions 13 Mt blocks, largest is 196
1( 1) 11( 30) 2( 8) 3( 30) 8( 176) 12( 8) 13( 1) 5( 127) 4( 72) 6( 176) 9( 127) 7( 196) 10( 72)
first maxtrix multiply, sparsity in % 99.561 ...
second maxtrix multiply, sparsity in % 95.728 ...
512 product functions 11 Mt blocks, largest is 112
10( 8) 4( 64) 1( 1) 2( 8) 11( 1) 3( 29) 7( 98) 5( 98) 8( 64) 6( 112) 9( 29)
first maxtrix multiply, sparsity in % 99.170 ...
second maxtrix multiply, sparsity in % 93.246 ...
512 product functions 11 Mt blocks, largest is 112
10( 8) 1( 1) 2( 8) 11( 1) 3( 29) 7( 98) 4( 64) 5( 98) 8( 64) 9( 29) 6( 112)
first maxtrix multiply, sparsity in % 99.170 ...
second maxtrix multiply, sparsity in % 93.449 ...
256 product functions 9 Mt blocks, largest is 70
1( 1) 6( 56) 2( 8) 3( 28) 4( 56) 8( 8) 9( 1) 7( 28) 5( 70)
first maxtrix multiply, sparsity in % 98.438 ...
second maxtrix multiply, sparsity in % 89.922 ...
done.
11888 non-zero transitions.
spectrum on <anmr.dat>
Range (delta in ppm) -0.148694152832031 6.07862304687500
Range (delta in Hz) -59.4776611328125 2431.44921875000
Min/max Int. ) 4.255446638228635E-003
computing spectrum ...
done.
writing output file ...
done.
All done.
After “ANMR” is finished computing, the file anrm.dat is written and it contains the spectrum (intensity vs shift) the user can plot:
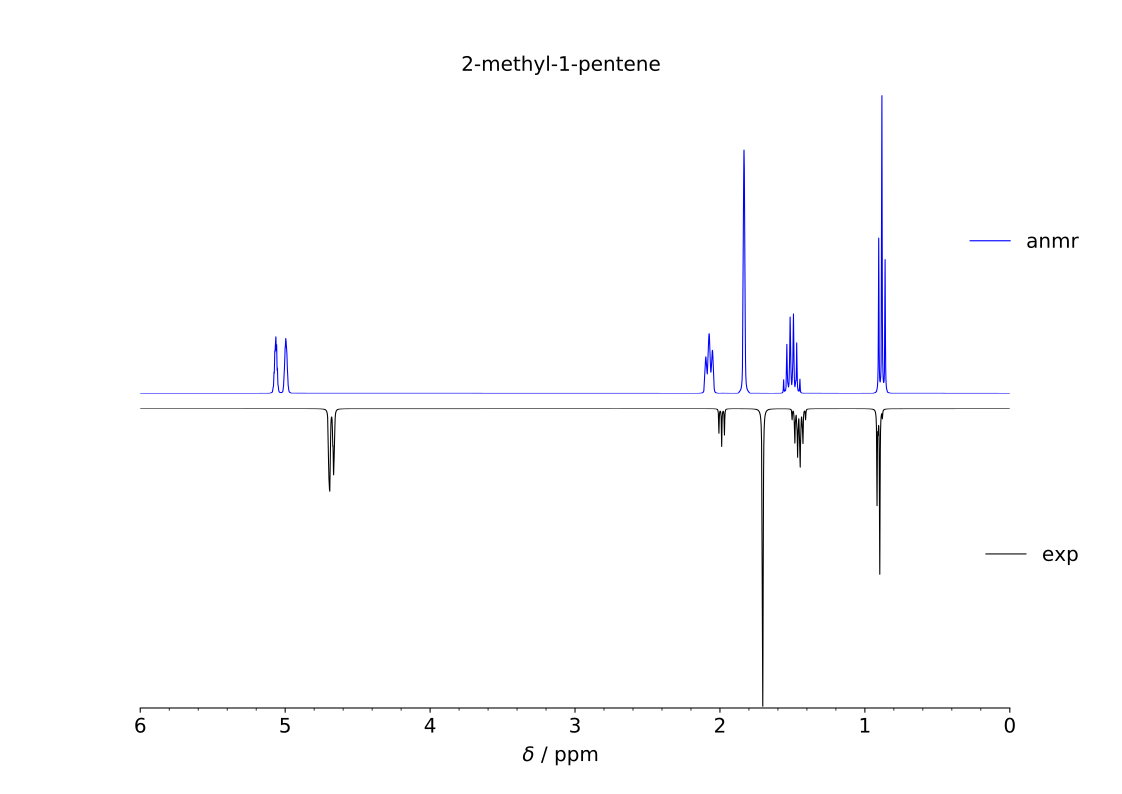
1H NMR spectrum of 2-methyl-1-pentene in chloroform at 400 MHz, comparing calculated and experimental spectrum. Exp taken from [SDBSWeb: https://sdsbs.db.aist.go.jp (National Institute of Advanced Industria Science and Technology, 16-10-2019) (SDBSNo. 225)].
Calculation of optical rotation
Example of calculating the optical rotation (OR) of \({\alpha}\)-D-glucopyranose and \({\beta}\)-D-glucopyranose
$ cat coord
$coord
4.767617729643 1.419722956160 -0.935928523270 c
2.060519933151 1.018564106099 0.018508999801 c
1.117123538906 -1.672829858373 -0.503205491294 c
-1.630185557104 -1.907024231918 0.381798382833 c
-3.231934402283 0.114098396821 -0.946387162026 c
-2.054506607483 2.733521342735 -0.476945496486 c
6.573917814231 0.299757760377 0.644171554348 o
2.701764145723 -3.389883248762 0.757273415150 o
-2.474976317765 -4.364547825619 -0.194465063937 o
-5.756659411064 0.028968027273 -0.147075665723 o
-2.392843407897 3.285715609737 2.075065264520 o
0.494646986154 2.818420632247 -1.217058181223 o
5.166309029039 3.444411414995 -0.955696975717 h
4.917082594970 0.697482954388 -2.875513316086 h
2.028271057226 1.334217639095 2.072518794364 h
1.166232043304 -2.021210902373 -2.556485813657 h
-1.706933193804 -1.591394158358 2.438772245996 h
-3.243398855045 -0.269083275045 -2.979576642617 h
-2.986117144565 4.141194310390 -1.687833317894 h
6.101393643219 -1.455981517989 0.903224373247 h
1.931942418123 -5.050271483321 0.669979783712 h
-4.301727918165 -4.398236354676 -0.073956936283 h
-5.867865848582 0.801456550117 1.513775016901 h
-1.904503272873 5.015209896774 2.427076955272 h
$end
> cat coord
$coord
-4.593283219880 -2.038743186735 -0.636452967953 c
-2.036936962862 -1.118771241606 0.381072259227 c
-1.393672207872 1.545169424514 -0.573991926259 c
1.230803011074 2.315828381590 0.381771457073 c
3.170942908620 0.300973102283 -0.355491229536 c
2.255573338318 -2.307742232726 0.541438449566 c
-6.640392582629 -0.916964528483 0.611579609638 o
-3.264742854030 3.226185184337 0.267849459933 o
1.809278772107 4.682981692940 -0.676188805997 o
5.487273113550 0.953020687813 0.761673737265 o
3.923898544696 -4.099604118098 -0.357934563930 o
-0.172496743908 -2.860970129555 -0.403247323969 o
-4.737864133466 -4.071051670096 -0.309006279484 h
-4.675563768934 -1.688074041579 -2.679528559296 h
-2.130211757399 -1.068689023410 2.461433765691 h
-1.332118968415 1.513661477252 -2.654856311664 h
1.188111054696 2.454048409034 2.459177008664 h
3.338386206165 0.235397437902 -2.427108484216 h
2.195785280720 -2.308246164055 2.641197391967 h
-6.417624332157 0.905517603874 0.564577678187 h
-2.725410970230 4.932451514073 -0.127888169080 h
3.533151721221 5.114003866682 -0.238203868775 h
6.718898090783 -0.348078354690 0.386760489292 h
3.188626269201 -5.770181712835 -0.192504534303 h
$end
input structure of \({\alpha}\)-D-glucopyranose (left) and \({\beta}\)-D-glucopyranose (right)
Start with an input structure (this one is taken from Pubchem) and create the conformers for the CENSO calculation.
For the computation of the OR, it is important to get an (ideally) complete conformer ensemble.
Therefore, the crest_combi bash script is used. It automatically starts several CREST
runs at GFN-FF and GFN2-xTB theory levels complemented by searching on artificial PES (scaled disp and charge)
to overcome possible method deficiencies to efficiently scan a large part of
the PES of the respective molecule. To use it, download the script from the release page https://github.com/grimme-lab/CRENSO/releases
and make it executable (chmod u+x crest_combi).
Hint
To save computational costs, the crest_combi script reduces the number of generated structures to
the most representative structures via the clustering algorithm of CREST.
The -or command sets the maximum of generated structures to 1000 instead of the default value of 500
to prevent that relevant conformers are sorted out in this step.
$ crest_combi coord -solvent h2o -T 14 -or > crest.out
In our case CREST found 54 conformers within an energy window of 8 kcal/mol (default value of crest_combi).
The final ensemble is optimized at GFN2-xTB level with the ALPB implicit solvation model.
We then create a new folder for the CENSO reranking and copy the necessary files:
$ mkdir censo
$ cp crest_combi.xyz censo/
$ cd censo/
To prevent sorting out relevant conformers, we increase the energy thresholds of part 0 and part 1 to 5.0 and 3.0 kcal/mol, respectively. By default, the PBE kernel is employed on the r2SCAN orbitals for the calculation of the optical rotation.
Warning
Since optical rotation is currently not implemented in the ORCA code, OR calculations are only possible with TURBOMOLE.
Hint
Since the generated structures of crest_combi and hence the final ensemble for the OR calculation
are non-deterministic the computed OR values of different runs can differ. Therefore, it is recommended
to take the average of at least three independent runs of crest_combi + CENSO.
$ censo --input crest_combi.xyz --solvent h2o --OR on -thrpart0 5.0 -thrpart1 3.0 > censo.out
______________________________________________________________
| |
| |
| CENSO - Commandline ENSO |
| v 1.1.2 |
| energetic sorting of CREST Conformer Rotamer Ensembles |
| University of Bonn, MCTC |
| Feb 2021 |
| based on ENSO version 2.0.1 |
| F. Bohle and S. Grimme |
| |
|______________________________________________________________|
Please cite:
S. Grimme, F. Bohle, A. Hansen, P. Pracht, S. Spicher, and M. Stahn
J. Phys. Chem. A 2021, 125, 19, 4039-4054.
DOI: https://doi.org/10.1021/acs.jpca.1c00971
This program is distributed in the hope that it will be useful,
but WITHOUT ANY WARRANTY; without even the implied warranty of
MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE.
----------------------------------------------------------------------------------------------------
PARAMETERS
----------------------------------------------------------------------------------------------------
program call: censo --input crest_combi.xyz --solvent h2o --OR on -thrpart0 5.0 -thrpart1 3.0
The configuration file .censorc is read from /home/gorges/.censorc.
Reading conformer rotamer ensemble from: /tmp1/gorges/3762289.majestix.thch.uni-bonn.de/crest_combi.xyz.
Reading file: censo_solvents.json
--------------------------------------------------
CRE SORTING SETTINGS
--------------------------------------------------
number of atoms in system: 24
number of considered conformers: 58
number of all conformers from input: 58
charge: 0
unpaired: 0
solvent: h2o
temperature: 298.15
evaluate at different temperatures: on
temperature range: 273.15, 278.15, 283.15, 288.15, ...
calculate mRRHO contribution: on
consider symmetry for mRRHO contribution: on
cautious checking for error and failed calculations: on
checking the DFT-ensemble using CREST: off
maxthreads: 7
omp: 4
automatically balance maxthreads and omp: off
--------------------------------------------------
CRE CHEAP-PRESCREENING - PART0
--------------------------------------------------
part0: on
starting number of considered conformers: 58
program for part0: tm
functional for fast single-point: b97-d3(0)
basis set for fast single-point: def2-SV(P)
threshold g_thr(0) for sorting in part0: 5.0
Solvent model used with xTB: alpb
short-notation:
b97-d3(0)/def2-SV(P) // GFNn-xTB (Input geometry)
--------------------------------------------------
CRE PRESCREENING - PART1
--------------------------------------------------
part1: on
program for part1: tm
functional for initial evaluation: r2scan-3c
basis set for initial evaluation: def2-mTZVPP
calculate mRRHO contribution: on
program for mRRHO contribution: xtb
GFN version for mRRHO and/or GBSA_Gsolv: gfn2
Apply constraint to input geometry during mRRHO calculation: on
solvent model applied with xTB: alpb
evaluate at different temperatures: off
threshold g_thr(1) and G_thr(1) for sorting in part1: 3.0
solvent model for Gsolv contribution of part1: cosmors
short-notation:
r2scan-3c + COSMORS[h2o] + GmRRHO(GFN2[alpb]-bhess) // GFNn-xTB (Input geometry)
--------------------------------------------------
CRE OPTIMIZATION - PART2
--------------------------------------------------
part2: on
program: tm
functional for part2: r2scan-3c
basis set for part2: def2-mTZVPP
using xTB-optimizer for optimization: on
using the new ensemble optimizer: on
optimize all conformers below this G_thr(opt,2) threshold: 2.5
spearmanthr: 0.936
optimization level in part2: lax
solvent model applied in the optimization: dcosmors
solvent model for Gsolv contribution: cosmors
evaluate at different temperatures: on
Boltzmann sum threshold G_thr(2) for sorting in part2: 99.0
calculate mRRHO contribution: on
program for mRRHO contribution: xtb
GFN version for mRRHO and/or GBSA_Gsolv: gfn2
Apply constraint to input geometry during mRRHO calculation: on
solvent model applied with xTB: alpb
short-notation:
r2scan-3c + COSMORS[h2o] + GmRRHO(GFN2[alpb]-bhess) // r2scan-3c[DCOSMORS]
--------------------------------------------------
OPTICAL ROTATION MODE - PART5
--------------------------------------------------
part5: on
frequency in [nm]: 589.0
functional for SCF: r2scan-3c
functional for optical rotation: pbe-d4
basis set for optical rotation: def2-SVPD
Boltzmann sum threshold employed: 99.0
END of parameters
------------------------------------------------------------
PATHS of external QM programs
------------------------------------------------------------
The following program paths are used:
xTB: /home/abt-grimme/AK-bin/xtb
TURBOMOLE: /home/abt-grimme/TURBOMOLE.7.5//bin/em64t-unknown-linux-gnu_smp
Setup of COSMO-RS:
ctd = BP_TZVP_C30_1601.ctd
cdir = "/software/cluster/COSMOthermX16/COSMOtherm/CTDATA-FILES"
ldir = "/software/cluster/COSMOthermX16/COSMOtherm/CTDATA-FILES"
Using /software/cluster/COSMOthermX16/COSMOtherm/DATABASE-COSMO/BP-TZVP-COSMO
as path to the COSMO-RS NORMAL DATABASE.
Using cefine from /tmp/_MEI9YTTJR/cefine
PARNODES for TM or COSMO-RS calculation was set to 4
Using COSMOtherm from /software/cluster/COSMOthermX16/COSMOtherm/BIN-LINUX/cosmotherm
----------------------------------------------------------------------------------------------------
Processing data from previous run (enso.json)
----------------------------------------------------------------------------------------------------
INFORMATION: No restart information exists and is created during this run!
----------------------------------------------------------------------------------------------------
CRE CHEAP-PRESCREENING - PART0
----------------------------------------------------------------------------------------------------
program: tm
functional for part0: b97-d3(0)
basis set for part0: def2-SV(P)
threshold g_thr(0): 5.0
starting number of considered conformers: 58
temperature: 298.15
Calculating efficient gas-phase single-point energies:
The efficient gas-phase single-point is calculated for:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF10, CONF11
CONF12, CONF13, CONF14, CONF15, CONF16, CONF17, CONF18, CONF19, CONF20, CONF21, CONF22
CONF23, CONF24, CONF25, CONF26, CONF27, CONF28, CONF29, CONF30, CONF31, CONF32, CONF33
CONF34, CONF35, CONF36, CONF37, CONF38, CONF39, CONF40, CONF41, CONF42, CONF43, CONF44
CONF45, CONF46, CONF47, CONF48, CONF49, CONF50, CONF51, CONF52, CONF53, CONF54, CONF55
CONF56, CONF57, CONF58
Constructed folders!
Starting 58 ALPB-Gsolv calculations
Running single-point in CONF1/part0_sp
Running single-point in CONF2/part0_sp
Running single-point in CONF4/part0_sp
Running single-point in CONF3/part0_sp
Running single-point in CONF6/part0_sp
Running single-point in CONF5/part0_sp
Running single-point in CONF7/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF1/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF2/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF3/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF6/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF7/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF5/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF4/part0_sp
Running single-point in CONF8/part0_sp
Running single-point in CONF9/part0_sp
Running single-point in CONF10/part0_sp
Running single-point in CONF11/part0_sp
Running single-point in CONF12/part0_sp
Running single-point in CONF13/part0_sp
Running single-point in CONF14/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF8/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF9/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF10/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF11/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF12/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF13/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF14/part0_sp
Running single-point in CONF15/part0_sp
Running single-point in CONF16/part0_sp
Running single-point in CONF17/part0_sp
Running single-point in CONF18/part0_sp
Running single-point in CONF19/part0_sp
Running single-point in CONF20/part0_sp
Running single-point in CONF21/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF15/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF16/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF17/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF19/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF18/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF20/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF21/part0_sp
Running single-point in CONF22/part0_sp
Running single-point in CONF23/part0_sp
Running single-point in CONF24/part0_sp
Running single-point in CONF25/part0_sp
Running single-point in CONF26/part0_sp
Running single-point in CONF27/part0_sp
Running single-point in CONF28/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF22/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF23/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF24/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF26/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF25/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF27/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF28/part0_sp
Running single-point in CONF29/part0_sp
Running single-point in CONF30/part0_sp
Running single-point in CONF31/part0_sp
Running single-point in CONF32/part0_sp
Running single-point in CONF33/part0_sp
Running single-point in CONF34/part0_sp
Running single-point in CONF35/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF29/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF30/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF31/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF32/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF35/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF33/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF34/part0_sp
Running single-point in CONF36/part0_sp
Running single-point in CONF37/part0_sp
Running single-point in CONF38/part0_sp
Running single-point in CONF39/part0_sp
Running single-point in CONF42/part0_sp
Running single-point in CONF40/part0_sp
Running single-point in CONF41/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF36/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF37/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF38/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF39/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF42/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF40/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF41/part0_sp
Running single-point in CONF43/part0_sp
Running single-point in CONF44/part0_sp
Running single-point in CONF45/part0_sp
Running single-point in CONF46/part0_sp
Running single-point in CONF47/part0_sp
Running single-point in CONF48/part0_sp
Running single-point in CONF49/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF43/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF44/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF45/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF46/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF47/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF49/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF48/part0_sp
Running single-point in CONF50/part0_sp
Running single-point in CONF51/part0_sp
Running single-point in CONF52/part0_sp
Running single-point in CONF53/part0_sp
Running single-point in CONF55/part0_sp
Running single-point in CONF54/part0_sp
Running single-point in CONF56/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF50/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF51/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF52/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF54/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF53/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF55/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF56/part0_sp
Running single-point in CONF57/part0_sp
Running single-point in CONF58/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF57/part0_sp
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF58/part0_sp
Tasks completed!
The efficient gas-phase single-point was successful for CONF1/part0_sp: E(DFT) = -686.03824317 Gsolv = -0.02621038
The efficient gas-phase single-point was successful for CONF2/part0_sp: E(DFT) = -686.03792683 Gsolv = -0.02797994
The efficient gas-phase single-point was successful for CONF3/part0_sp: E(DFT) = -686.03775935 Gsolv = -0.02787613
The efficient gas-phase single-point was successful for CONF4/part0_sp: E(DFT) = -686.03338798 Gsolv = -0.02822958
The efficient gas-phase single-point was successful for CONF5/part0_sp: E(DFT) = -686.03432035 Gsolv = -0.02809768
The efficient gas-phase single-point was successful for CONF6/part0_sp: E(DFT) = -686.03323088 Gsolv = -0.02905745
The efficient gas-phase single-point was successful for CONF7/part0_sp: E(DFT) = -686.03231779 Gsolv = -0.02954455
The efficient gas-phase single-point was successful for CONF8/part0_sp: E(DFT) = -686.03607417 Gsolv = -0.02710004
The efficient gas-phase single-point was successful for CONF9/part0_sp: E(DFT) = -686.03304877 Gsolv = -0.02886089
The efficient gas-phase single-point was successful for CONF10/part0_sp: E(DFT) = -686.03485843 Gsolv = -0.02716784
The efficient gas-phase single-point was successful for CONF11/part0_sp: E(DFT) = -686.03194476 Gsolv = -0.02930216
The efficient gas-phase single-point was successful for CONF12/part0_sp: E(DFT) = -686.03253190 Gsolv = -0.03030240
The efficient gas-phase single-point was successful for CONF13/part0_sp: E(DFT) = -686.03236871 Gsolv = -0.02937302
The efficient gas-phase single-point was successful for CONF14/part0_sp: E(DFT) = -686.03287967 Gsolv = -0.02896135
The efficient gas-phase single-point was successful for CONF15/part0_sp: E(DFT) = -686.03337031 Gsolv = -0.02927880
The efficient gas-phase single-point was successful for CONF16/part0_sp: E(DFT) = -686.03523328 Gsolv = -0.02711044
The efficient gas-phase single-point was successful for CONF17/part0_sp: E(DFT) = -686.03050697 Gsolv = -0.02947881
The efficient gas-phase single-point was successful for CONF18/part0_sp: E(DFT) = -686.03014192 Gsolv = -0.02787104
The efficient gas-phase single-point was successful for CONF19/part0_sp: E(DFT) = -686.03226408 Gsolv = -0.02751159
The efficient gas-phase single-point was successful for CONF20/part0_sp: E(DFT) = -686.02988165 Gsolv = -0.02743968
The efficient gas-phase single-point was successful for CONF21/part0_sp: E(DFT) = -686.02887551 Gsolv = -0.03087333
The efficient gas-phase single-point was successful for CONF22/part0_sp: E(DFT) = -686.02855171 Gsolv = -0.02765567
The efficient gas-phase single-point was successful for CONF23/part0_sp: E(DFT) = -686.02817357 Gsolv = -0.02811716
The efficient gas-phase single-point was successful for CONF24/part0_sp: E(DFT) = -686.03519426 Gsolv = -0.02747200
The efficient gas-phase single-point was successful for CONF25/part0_sp: E(DFT) = -686.03096537 Gsolv = -0.03059177
The efficient gas-phase single-point was successful for CONF26/part0_sp: E(DFT) = -686.03353987 Gsolv = -0.02792358
The efficient gas-phase single-point was successful for CONF27/part0_sp: E(DFT) = -686.03280188 Gsolv = -0.02738026
The efficient gas-phase single-point was successful for CONF28/part0_sp: E(DFT) = -686.03415690 Gsolv = -0.02595701
The efficient gas-phase single-point was successful for CONF29/part0_sp: E(DFT) = -686.03302739 Gsolv = -0.02936445
The efficient gas-phase single-point was successful for CONF30/part0_sp: E(DFT) = -686.03328415 Gsolv = -0.02519214
The efficient gas-phase single-point was successful for CONF31/part0_sp: E(DFT) = -686.02988503 Gsolv = -0.03116630
The efficient gas-phase single-point was successful for CONF32/part0_sp: E(DFT) = -686.02767535 Gsolv = -0.02940219
The efficient gas-phase single-point was successful for CONF33/part0_sp: E(DFT) = -686.03078606 Gsolv = -0.02456167
The efficient gas-phase single-point was successful for CONF34/part0_sp: E(DFT) = -686.03068178 Gsolv = -0.02493813
The efficient gas-phase single-point was successful for CONF35/part0_sp: E(DFT) = -686.03063368 Gsolv = -0.02487044
The efficient gas-phase single-point was successful for CONF36/part0_sp: E(DFT) = -686.03074458 Gsolv = -0.02696015
The efficient gas-phase single-point was successful for CONF37/part0_sp: E(DFT) = -686.02807003 Gsolv = -0.02811961
The efficient gas-phase single-point was successful for CONF38/part0_sp: E(DFT) = -686.02631311 Gsolv = -0.02681294
The efficient gas-phase single-point was successful for CONF39/part0_sp: E(DFT) = -686.03188975 Gsolv = -0.02586421
The efficient gas-phase single-point was successful for CONF40/part0_sp: E(DFT) = -686.02628039 Gsolv = -0.03059854
The efficient gas-phase single-point was successful for CONF41/part0_sp: E(DFT) = -686.02600626 Gsolv = -0.02955414
The efficient gas-phase single-point was successful for CONF42/part0_sp: E(DFT) = -686.03061205 Gsolv = -0.02864620
The efficient gas-phase single-point was successful for CONF43/part0_sp: E(DFT) = -686.02831616 Gsolv = -0.02970543
The efficient gas-phase single-point was successful for CONF44/part0_sp: E(DFT) = -686.02583139 Gsolv = -0.02925069
The efficient gas-phase single-point was successful for CONF45/part0_sp: E(DFT) = -686.02568848 Gsolv = -0.02779597
The efficient gas-phase single-point was successful for CONF46/part0_sp: E(DFT) = -686.03205976 Gsolv = -0.02748900
The efficient gas-phase single-point was successful for CONF47/part0_sp: E(DFT) = -686.02939924 Gsolv = -0.02829244
The efficient gas-phase single-point was successful for CONF48/part0_sp: E(DFT) = -686.02868080 Gsolv = -0.02703056
The efficient gas-phase single-point was successful for CONF49/part0_sp: E(DFT) = -686.02740071 Gsolv = -0.02802221
The efficient gas-phase single-point was successful for CONF50/part0_sp: E(DFT) = -686.02915868 Gsolv = -0.02690862
The efficient gas-phase single-point was successful for CONF51/part0_sp: E(DFT) = -686.02844438 Gsolv = -0.02974364
The efficient gas-phase single-point was successful for CONF52/part0_sp: E(DFT) = -686.02643283 Gsolv = -0.02847615
The efficient gas-phase single-point was successful for CONF53/part0_sp: E(DFT) = -686.02323161 Gsolv = -0.03060674
The efficient gas-phase single-point was successful for CONF54/part0_sp: E(DFT) = -686.02874379 Gsolv = -0.02656045
The efficient gas-phase single-point was successful for CONF55/part0_sp: E(DFT) = -686.02712584 Gsolv = -0.02748822
The efficient gas-phase single-point was successful for CONF56/part0_sp: E(DFT) = -686.02612533 Gsolv = -0.02734670
The efficient gas-phase single-point was successful for CONF57/part0_sp: E(DFT) = -686.02703634 Gsolv = -0.02785506
The efficient gas-phase single-point was successful for CONF58/part0_sp: E(DFT) = -686.02655304 Gsolv = -0.02741292
----------------------------------------------------------------------------------------------------
Removing high lying conformers by improved energy description
----------------------------------------------------------------------------------------------------
CONF# E [Eh] ΔE [kcal/mol] E [Eh] Gsolv [Eh] gtot ΔE(DFT) ΔGsolv Δgtot
GFN2-xTB GFN2-xTB b97-d3(0)/def2-SV(P) alpb [Eh] [kcal/mol] [kcal/mol] [kcal/mol]
[alpb] [alpb] [gfn2]
CONF1 -43.3944150 0.00 -686.0382432 -0.0262104 -686.0644536 -0.20 1.11 0.91
CONF2 -43.3941370 0.17 -686.0379268 -0.0279799 -686.0659068 0.00 0.00 0.00 <------
CONF3 -43.3938279 0.37 -686.0377594 -0.0278761 -686.0656355 0.11 0.07 0.17
CONF4 -43.3924509 1.23 -686.0333880 -0.0282296 -686.0616176 2.85 -0.16 2.69
CONF5 -43.3922430 1.36 -686.0343203 -0.0280977 -686.0624180 2.26 -0.07 2.19
CONF6 -43.3908696 2.22 -686.0332309 -0.0290574 -686.0622883 2.95 -0.68 2.27
CONF7 -43.3907915 2.27 -686.0323178 -0.0295445 -686.0618623 3.52 -0.98 2.54
CONF8 -43.3906693 2.35 -686.0360742 -0.0271000 -686.0631742 1.16 0.55 1.71
CONF9 -43.3906140 2.39 -686.0330488 -0.0288609 -686.0619097 3.06 -0.55 2.51
CONF10 -43.3905816 2.41 -686.0348584 -0.0271678 -686.0620263 1.93 0.51 2.44
CONF11 -43.3903938 2.52 -686.0319448 -0.0293022 -686.0612469 3.75 -0.83 2.92
CONF12 -43.3903248 2.57 -686.0325319 -0.0303024 -686.0628343 3.39 -1.46 1.93
CONF13 -43.3902228 2.63 -686.0323687 -0.0293730 -686.0617417 3.49 -0.87 2.61
CONF14 -43.3901776 2.66 -686.0328797 -0.0289614 -686.0618410 3.17 -0.62 2.55
CONF15 -43.3899521 2.80 -686.0333703 -0.0292788 -686.0626491 2.86 -0.82 2.04
CONF16 -43.3898750 2.85 -686.0352333 -0.0271104 -686.0623437 1.69 0.55 2.24
CONF17 -43.3896242 3.01 -686.0305070 -0.0294788 -686.0599858 4.66 -0.94 3.72
CONF18 -43.3893461 3.18 -686.0301419 -0.0278710 -686.0580130 4.89 0.07 4.95
CONF19 -43.3892644 3.23 -686.0322641 -0.0275116 -686.0597757 3.55 0.29 3.85
CONF20 -43.3890861 3.34 -686.0298817 -0.0274397 -686.0573213 5.05 0.34 5.39
CONF21 -43.3889876 3.41 -686.0288755 -0.0308733 -686.0597488 5.68 -1.82 3.86
CONF22 -43.3889257 3.44 -686.0285517 -0.0276557 -686.0562074 5.88 0.20 6.09
CONF23 -43.3888683 3.48 -686.0281736 -0.0281172 -686.0562907 6.12 -0.09 6.03
CONF24 -43.3888012 3.52 -686.0351943 -0.0274720 -686.0626663 1.71 0.32 2.03
CONF25 -43.3887821 3.53 -686.0309654 -0.0305918 -686.0615571 4.37 -1.64 2.73
CONF26 -43.3887129 3.58 -686.0335399 -0.0279236 -686.0614635 2.75 0.04 2.79
CONF27 -43.3886231 3.63 -686.0328019 -0.0273803 -686.0601821 3.22 0.38 3.59
CONF28 -43.3885781 3.66 -686.0341569 -0.0259570 -686.0601139 2.37 1.27 3.64
CONF29 -43.3883958 3.78 -686.0330274 -0.0293644 -686.0623918 3.07 -0.87 2.21
CONF30 -43.3881924 3.90 -686.0332841 -0.0251921 -686.0584763 2.91 1.75 4.66
CONF31 -43.3877136 4.21 -686.0298850 -0.0311663 -686.0610513 5.05 -2.00 3.05
CONF32 -43.3877133 4.21 -686.0276754 -0.0294022 -686.0570775 6.43 -0.89 5.54
CONF33 -43.3876468 4.25 -686.0307861 -0.0245617 -686.0553477 4.48 2.14 6.63
CONF34 -43.3874315 4.38 -686.0306818 -0.0249381 -686.0556199 4.55 1.91 6.46
CONF35 -43.3874202 4.39 -686.0306337 -0.0248704 -686.0555041 4.58 1.95 6.53
CONF36 -43.3872487 4.50 -686.0307446 -0.0269602 -686.0577047 4.51 0.64 5.15
CONF37 -43.3872377 4.50 -686.0280700 -0.0281196 -686.0561896 6.19 -0.09 6.10
CONF38 -43.3870696 4.61 -686.0263131 -0.0268129 -686.0531260 7.29 0.73 8.02
CONF39 -43.3869909 4.66 -686.0318898 -0.0258642 -686.0577540 3.79 1.33 5.12
CONF40 -43.3869456 4.69 -686.0262804 -0.0305985 -686.0568789 7.31 -1.64 5.67
CONF41 -43.3868778 4.73 -686.0260063 -0.0295541 -686.0555604 7.48 -0.99 6.49
CONF42 -43.3868589 4.74 -686.0306120 -0.0286462 -686.0592582 4.59 -0.42 4.17
CONF43 -43.3868177 4.77 -686.0283162 -0.0297054 -686.0580216 6.03 -1.08 4.95
CONF44 -43.3866458 4.88 -686.0258314 -0.0292507 -686.0550821 7.59 -0.80 6.79
CONF45 -43.3864981 4.97 -686.0256885 -0.0277960 -686.0534845 7.68 0.12 7.80
CONF46 -43.3864925 4.97 -686.0320598 -0.0274890 -686.0595488 3.68 0.31 3.99
CONF47 -43.3864737 4.98 -686.0293992 -0.0282924 -686.0576917 5.35 -0.20 5.16
CONF48 -43.3856106 5.52 -686.0286808 -0.0270306 -686.0557114 5.80 0.60 6.40
CONF49 -43.3853412 5.69 -686.0274007 -0.0280222 -686.0554229 6.61 -0.03 6.58
CONF50 -43.3845739 6.18 -686.0291587 -0.0269086 -686.0560673 5.50 0.67 6.17
CONF51 -43.3844952 6.22 -686.0284444 -0.0297436 -686.0581880 5.95 -1.11 4.84
CONF52 -43.3844717 6.24 -686.0264328 -0.0284761 -686.0549090 7.21 -0.31 6.90
CONF53 -43.3837119 6.72 -686.0232316 -0.0306067 -686.0538384 9.22 -1.65 7.57
CONF54 -43.3833306 6.96 -686.0287438 -0.0265605 -686.0553042 5.76 0.89 6.65
CONF55 -43.3832271 7.02 -686.0271258 -0.0274882 -686.0546141 6.78 0.31 7.09
CONF56 -43.3827789 7.30 -686.0261253 -0.0273467 -686.0534720 7.41 0.40 7.80
CONF57 -43.3825259 7.46 -686.0270363 -0.0278551 -686.0548914 6.83 0.08 6.91
CONF58 -43.3819213 7.84 -686.0265530 -0.0274129 -686.0539660 7.14 0.36 7.49
----------------------------------------------------------------------------------------------------
Number of conformers observed within the following Δg windows:
Δg [kcal/mol] #CONF sum(Boltzmann_weights)
---------------------------------------------
0 - 0.5 2 0.75
0 - 1.0 3 0.84
0 - 1.5 3 0.84
0 - 2.0 5 0.88
0 - 2.5 12 0.95
0 - 3.0 20 0.99
0 - 3.5 21 0.99
0 - 4.0 27 1.00
0 - 4.5 28 1.00
0 - 5.0 32 1.00
0 - 5.5 36 1.00
---------------------------------------------
--------------------------------------------------
Conformers considered further
--------------------------------------------------
These conformers are below the 5.000 kcal/mol g_thr(0) threshold.
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF10, CONF11
CONF12, CONF13, CONF14, CONF15, CONF16, CONF17, CONF18, CONF19, CONF21, CONF24, CONF25
CONF26, CONF27, CONF28, CONF29, CONF30, CONF31, CONF42, CONF43, CONF46, CONF51
Calculating Boltzmann averaged (free) energy of ensemble on input geometries (not DFT optimized)!
temperature /K: avE(T) /a.u. avG(T) /a.u.
----------------------------------------------------------------------------------------------------
298.15 -686.0372298 -686.0651055 <<==part0==
----------------------------------------------------------------------------------------------------
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>END of Part0<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
Ran part0 in 42.1222 seconds
----------------------------------------------------------------------------------------------------
CRE PRESCREENING - PART1
----------------------------------------------------------------------------------------------------
program: tm
functional for part1 and 2: r2scan-3c
basis set for part1 and 2: def2-mTZVPP
Solvent: h2o
solvent model for Gsolv contribution: cosmors
threshold g_thr(1) and G_thr(1): 3.0
starting number of considered conformers: 32
calculate mRRHO contribution: on
program for mRRHO contribution: xtb
GFN version for mRRHO and/or GBSA_Gsolv: gfn2
Apply constraint to input geometry during mRRHO calculation: on
temperature: 298.15
Calculating single-point energies and solvation contribution (G_solv):
The prescreening COSMO-RS is calculated for:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF10, CONF11
CONF12, CONF13, CONF14, CONF15, CONF16, CONF17, CONF18, CONF19, CONF21, CONF24, CONF25
CONF26, CONF27, CONF28, CONF29, CONF30, CONF31, CONF42, CONF43, CONF46, CONF51
Constructed folders!
Constructed folders!
Starting 32 COSMO-RS-Gsolv calculations.
Running COSMO-RS calculation in CONF1/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF2/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF3/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF4/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF5/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF6/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF7/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF8/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF9/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF10/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF11/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF12/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF13/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF14/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF15/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF16/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF17/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF18/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF19/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF21/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF24/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF25/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF26/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF27/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF28/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF29/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF30/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF31/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF42/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF43/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF46/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF51/r2scan-3c/COSMO
Tasks completed!
prescreening COSMO-RS calculation was successful for CONF1/r2scan-3c/COSMO: -0.02095183
prescreening COSMO-RS calculation was successful for CONF2/r2scan-3c/COSMO: -0.02304427
prescreening COSMO-RS calculation was successful for CONF3/r2scan-3c/COSMO: -0.02264314
prescreening COSMO-RS calculation was successful for CONF4/r2scan-3c/COSMO: -0.02689097
prescreening COSMO-RS calculation was successful for CONF5/r2scan-3c/COSMO: -0.02626860
prescreening COSMO-RS calculation was successful for CONF6/r2scan-3c/COSMO: -0.02575685
prescreening COSMO-RS calculation was successful for CONF7/r2scan-3c/COSMO: -0.02777475
prescreening COSMO-RS calculation was successful for CONF8/r2scan-3c/COSMO: -0.02187017
prescreening COSMO-RS calculation was successful for CONF9/r2scan-3c/COSMO: -0.02745793
prescreening COSMO-RS calculation was successful for CONF10/r2scan-3c/COSMO: -0.02259685
prescreening COSMO-RS calculation was successful for CONF11/r2scan-3c/COSMO: -0.02944421
prescreening COSMO-RS calculation was successful for CONF12/r2scan-3c/COSMO: -0.02727746
prescreening COSMO-RS calculation was successful for CONF13/r2scan-3c/COSMO: -0.02712694
prescreening COSMO-RS calculation was successful for CONF14/r2scan-3c/COSMO: -0.02762407
prescreening COSMO-RS calculation was successful for CONF15/r2scan-3c/COSMO: -0.02606066
prescreening COSMO-RS calculation was successful for CONF16/r2scan-3c/COSMO: -0.02208103
prescreening COSMO-RS calculation was successful for CONF17/r2scan-3c/COSMO: -0.02844436
prescreening COSMO-RS calculation was successful for CONF18/r2scan-3c/COSMO: -0.02677215
prescreening COSMO-RS calculation was successful for CONF19/r2scan-3c/COSMO: -0.02810842
prescreening COSMO-RS calculation was successful for CONF21/r2scan-3c/COSMO: -0.02975232
prescreening COSMO-RS calculation was successful for CONF24/r2scan-3c/COSMO: -0.02241619
prescreening COSMO-RS calculation was successful for CONF25/r2scan-3c/COSMO: -0.02891334
prescreening COSMO-RS calculation was successful for CONF26/r2scan-3c/COSMO: -0.02417223
prescreening COSMO-RS calculation was successful for CONF27/r2scan-3c/COSMO: -0.02467468
prescreening COSMO-RS calculation was successful for CONF28/r2scan-3c/COSMO: -0.02194300
prescreening COSMO-RS calculation was successful for CONF29/r2scan-3c/COSMO: -0.02505501
prescreening COSMO-RS calculation was successful for CONF30/r2scan-3c/COSMO: -0.02194166
prescreening COSMO-RS calculation was successful for CONF31/r2scan-3c/COSMO: -0.02975930
prescreening COSMO-RS calculation was successful for CONF42/r2scan-3c/COSMO: -0.02025841
prescreening COSMO-RS calculation was successful for CONF43/r2scan-3c/COSMO: -0.03036999
prescreening COSMO-RS calculation was successful for CONF46/r2scan-3c/COSMO: -0.02521187
prescreening COSMO-RS calculation was successful for CONF51/r2scan-3c/COSMO: -0.02913591
--------------------------------------------------
Removing high lying conformers
--------------------------------------------------
CONF# E(GFNn-xTB) ΔE(GFNn-xTB) E [Eh] Gsolv [Eh] gtot Δgtot
[a.u.] [kcal/mol] r2scan-3c COSMO-RS-normal [Eh] [kcal/mol]
[r2scan-3c]
CONF1 -43.3962305 0.00 -687.0946362 -0.0209518 -687.1155880 1.75
CONF2 -43.3959511 0.18 -687.0948696 -0.0230443 -687.1179139 0.29
CONF3 -43.3956417 0.37 -687.0951918 -0.0226431 -687.1178349 0.34
CONF4 -43.3942646 1.23 -687.0909200 -0.0268910 -687.1178110 0.36
CONF5 -43.3940567 1.36 -687.0915134 -0.0262686 -687.1177820 0.37
CONF6 -43.3926837 2.23 -687.0899435 -0.0257568 -687.1157003 1.68
CONF7 -43.3926044 2.28 -687.0890024 -0.0277747 -687.1167771 1.00
CONF8 -43.3924830 2.35 -687.0926336 -0.0218702 -687.1145038 2.43
CONF9 -43.3924281 2.39 -687.0897247 -0.0274579 -687.1171827 0.75
CONF10 -43.3923958 2.41 -687.0910490 -0.0225968 -687.1136459 2.97
CONF11 -43.3922081 2.52 -687.0889338 -0.0294442 -687.1183780 0.00 <------
CONF12 -43.3921384 2.57 -687.0900446 -0.0272775 -687.1173221 0.66
CONF13 -43.3920387 2.63 -687.0897804 -0.0271269 -687.1169073 0.92
CONF14 -43.3919912 2.66 -687.0902769 -0.0276241 -687.1179009 0.30
CONF15 -43.3917659 2.80 -687.0913013 -0.0260607 -687.1173619 0.64
CONF16 -43.3916896 2.85 -687.0917146 -0.0220810 -687.1137956 2.88
CONF17 -43.3914378 3.01 -687.0884564 -0.0284444 -687.1169008 0.93
CONF18 -43.3911600 3.18 -687.0876507 -0.0267722 -687.1144229 2.48
CONF19 -43.3910782 3.23 -687.0884940 -0.0281084 -687.1166024 1.11
CONF21 -43.3908014 3.41 -687.0873013 -0.0297523 -687.1170536 0.83
CONF24 -43.3906145 3.52 -687.0914313 -0.0224162 -687.1138475 2.84
CONF25 -43.3905959 3.54 -687.0880110 -0.0289133 -687.1169243 0.91
CONF26 -43.3905265 3.58 -687.0903285 -0.0241722 -687.1145008 2.43
CONF27 -43.3904368 3.64 -687.0890720 -0.0246747 -687.1137467 2.91
CONF28 -43.3903921 3.66 -687.0899359 -0.0219430 -687.1118789 4.08
CONF29 -43.3902113 3.78 -687.0904874 -0.0250550 -687.1155424 1.78
CONF30 -43.3900065 3.91 -687.0900450 -0.0219417 -687.1119867 4.01
CONF31 -43.3895276 4.21 -687.0880936 -0.0297593 -687.1178529 0.33
CONF42 -43.3886754 4.74 -687.0892550 -0.0202584 -687.1095134 5.56
CONF43 -43.3886315 4.77 -687.0865504 -0.0303700 -687.1169204 0.91
CONF46 -43.3883061 4.97 -687.0892486 -0.0252119 -687.1144605 2.46
CONF51 -43.3863091 6.23 -687.0854275 -0.0291359 -687.1145634 2.39
--------------------------------------------------
Conformers considered further
--------------------------------------------------
Below the g_thr(1) threshold of 3.0 kcal/mol.
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF10, CONF11
CONF12, CONF13, CONF14, CONF15, CONF16, CONF17, CONF18, CONF19, CONF21, CONF24, CONF25
CONF26, CONF27, CONF29, CONF31, CONF43, CONF46, CONF51
--------------------------------------------------
Calculating prescreening G_mRRHO with implicit solvation!
The prescreening G_mRRHO calculation is now performed for:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF10, CONF11
CONF12, CONF13, CONF14, CONF15, CONF16, CONF17, CONF18, CONF19, CONF21, CONF24, CONF25
CONF26, CONF27, CONF29, CONF31, CONF43, CONF46, CONF51
Constructed folders!
Starting 29 G_RRHO calculations.
Running GFN2-xTB mRRHO in CONF1/rrho_part1
Running GFN2-xTB mRRHO in CONF2/rrho_part1
Running GFN2-xTB mRRHO in CONF3/rrho_part1
Running GFN2-xTB mRRHO in CONF4/rrho_part1
Running GFN2-xTB mRRHO in CONF5/rrho_part1
Running GFN2-xTB mRRHO in CONF6/rrho_part1
Running GFN2-xTB mRRHO in CONF7/rrho_part1
Running GFN2-xTB mRRHO in CONF8/rrho_part1
Running GFN2-xTB mRRHO in CONF9/rrho_part1
Running GFN2-xTB mRRHO in CONF10/rrho_part1
Running GFN2-xTB mRRHO in CONF11/rrho_part1
Running GFN2-xTB mRRHO in CONF12/rrho_part1
Running GFN2-xTB mRRHO in CONF13/rrho_part1
Running GFN2-xTB mRRHO in CONF14/rrho_part1
Running GFN2-xTB mRRHO in CONF15/rrho_part1
Running GFN2-xTB mRRHO in CONF16/rrho_part1
Running GFN2-xTB mRRHO in CONF17/rrho_part1
Running GFN2-xTB mRRHO in CONF18/rrho_part1
Running GFN2-xTB mRRHO in CONF19/rrho_part1
Running GFN2-xTB mRRHO in CONF21/rrho_part1
Running GFN2-xTB mRRHO in CONF24/rrho_part1
Running GFN2-xTB mRRHO in CONF25/rrho_part1
Running GFN2-xTB mRRHO in CONF26/rrho_part1
Running GFN2-xTB mRRHO in CONF27/rrho_part1
Running GFN2-xTB mRRHO in CONF29/rrho_part1
Running GFN2-xTB mRRHO in CONF31/rrho_part1
Running GFN2-xTB mRRHO in CONF43/rrho_part1
Running GFN2-xTB mRRHO in CONF46/rrho_part1
Running GFN2-xTB mRRHO in CONF51/rrho_part1
Tasks completed!
The prescreening G_mRRHO run @ c1 was successful for CONF1/rrho_part1: 0.14940136 S_rot(sym)= 0.0000000 using= 0.1494014
The prescreening G_mRRHO run @ c1 was successful for CONF2/rrho_part1: 0.14899103 S_rot(sym)= 0.0000000 using= 0.1489910
The prescreening G_mRRHO run @ c1 was successful for CONF3/rrho_part1: 0.14915314 S_rot(sym)= 0.0000000 using= 0.1491531
The prescreening G_mRRHO run @ c1 was successful for CONF4/rrho_part1: 0.14858223 S_rot(sym)= 0.0000000 using= 0.1485822
The prescreening G_mRRHO run @ c1 was successful for CONF5/rrho_part1: 0.14875593 S_rot(sym)= 0.0000000 using= 0.1487559
The prescreening G_mRRHO run @ c1 was successful for CONF6/rrho_part1: 0.14909219 S_rot(sym)= 0.0000000 using= 0.1490922
The prescreening G_mRRHO run @ c1 was successful for CONF7/rrho_part1: 0.14870022 S_rot(sym)= 0.0000000 using= 0.1487002
The prescreening G_mRRHO run @ c1 was successful for CONF8/rrho_part1: 0.14917772 S_rot(sym)= 0.0000000 using= 0.1491777
The prescreening G_mRRHO run @ c1 was successful for CONF9/rrho_part1: 0.14904462 S_rot(sym)= 0.0000000 using= 0.1490446
The prescreening G_mRRHO run @ c1 was successful for CONF10/rrho_part1: 0.14847948 S_rot(sym)= 0.0000000 using= 0.1484795
The prescreening G_mRRHO run @ c1 was successful for CONF11/rrho_part1: 0.14864395 S_rot(sym)= 0.0000000 using= 0.1486440
The prescreening G_mRRHO run @ c1 was successful for CONF12/rrho_part1: 0.14858265 S_rot(sym)= 0.0000000 using= 0.1485827
The prescreening G_mRRHO run @ c1 was successful for CONF13/rrho_part1: 0.14870203 S_rot(sym)= 0.0000000 using= 0.1487020
The prescreening G_mRRHO run @ c1 was successful for CONF14/rrho_part1: 0.14876961 S_rot(sym)= 0.0000000 using= 0.1487696
The prescreening G_mRRHO run @ c1 was successful for CONF15/rrho_part1: 0.14875142 S_rot(sym)= 0.0000000 using= 0.1487514
The prescreening G_mRRHO run @ c1 was successful for CONF16/rrho_part1: 0.14877029 S_rot(sym)= 0.0000000 using= 0.1487703
The prescreening G_mRRHO run @ c1 was successful for CONF17/rrho_part1: 0.14815133 S_rot(sym)= 0.0000000 using= 0.1481513
The prescreening G_mRRHO run @ c1 was successful for CONF18/rrho_part1: 0.14846220 S_rot(sym)= 0.0000000 using= 0.1484622
The prescreening G_mRRHO run @ c1 was successful for CONF19/rrho_part1: 0.14903724 S_rot(sym)= 0.0000000 using= 0.1490372
The prescreening G_mRRHO run @ c1 was successful for CONF21/rrho_part1: 0.14802739 S_rot(sym)= 0.0000000 using= 0.1480274
The prescreening G_mRRHO run @ c1 was successful for CONF24/rrho_part1: 0.14835112 S_rot(sym)= 0.0000000 using= 0.1483511
The prescreening G_mRRHO run @ c1 was successful for CONF25/rrho_part1: 0.14875826 S_rot(sym)= 0.0000000 using= 0.1487583
The prescreening G_mRRHO run @ c1 was successful for CONF26/rrho_part1: 0.14813671 S_rot(sym)= 0.0000000 using= 0.1481367
The prescreening G_mRRHO run @ c1 was successful for CONF27/rrho_part1: 0.14838150 S_rot(sym)= 0.0000000 using= 0.1483815
The prescreening G_mRRHO run @ c1 was successful for CONF29/rrho_part1: 0.14814453 S_rot(sym)= 0.0000000 using= 0.1481445
The prescreening G_mRRHO run @ c1 was successful for CONF31/rrho_part1: 0.14813613 S_rot(sym)= 0.0000000 using= 0.1481361
The prescreening G_mRRHO run @ c1 was successful for CONF43/rrho_part1: 0.14813090 S_rot(sym)= 0.0000000 using= 0.1481309
The prescreening G_mRRHO run @ c1 was successful for CONF46/rrho_part1: 0.14807024 S_rot(sym)= 0.0000000 using= 0.1480702
The prescreening G_mRRHO run @ c1 was successful for CONF51/rrho_part1: 0.14838107 S_rot(sym)= 0.0000000 using= 0.1483811
--------------------------------------------------
* Gibbs free energies of part1 *
--------------------------------------------------
CONF# G(GFNn-xTB) ΔG(GFNn-xTB) E [Eh] Gsolv [Eh] GmRRHO [Eh] Gtot ΔGtot
[a.u.] [kcal/mol] r2scan-3c COSMO-RS-normal GFN2 [Eh] [kcal/mol]
[r2scan-3c] [alpb]-bhess
CONF1 -43.2468292 0.08 -687.0946362 -0.0209518 0.1494014 -686.9661867 2.23
CONF2 -43.2469600 0.00 -687.0948696 -0.0230443 0.1489910 -686.9689229 0.51
CONF3 -43.2464886 0.30 -687.0951918 -0.0226431 0.1491531 -686.9686818 0.66
CONF4 -43.2456823 0.80 -687.0909200 -0.0268910 0.1485822 -686.9692288 0.32
CONF5 -43.2453008 1.04 -687.0915134 -0.0262686 0.1487559 -686.9690260 0.44
CONF6 -43.2435915 2.11 -687.0899435 -0.0257568 0.1490922 -686.9666081 1.96
CONF7 -43.2439042 1.92 -687.0890024 -0.0277747 0.1487002 -686.9680769 1.04
CONF8 -43.2433052 2.29 -687.0926336 -0.0218702 0.1491777 -686.9653261 2.77
CONF9 -43.2433835 2.24 -687.0897247 -0.0274579 0.1490446 -686.9681381 1.00
CONF10 -43.2439163 1.91 -687.0910490 -0.0225968 0.1484795 -686.9651664 2.87
CONF11 -43.2435641 2.13 -687.0889338 -0.0294442 0.1486440 -686.9697341 0.00 <------
CONF12 -43.2435558 2.14 -687.0900446 -0.0272775 0.1485827 -686.9687394 0.62
CONF13 -43.2433367 2.27 -687.0897804 -0.0271269 0.1487020 -686.9682053 0.96
CONF14 -43.2432216 2.35 -687.0902769 -0.0276241 0.1487696 -686.9691313 0.38
CONF15 -43.2430145 2.48 -687.0913013 -0.0260607 0.1487514 -686.9686105 0.71
CONF16 -43.2429193 2.54 -687.0917146 -0.0220810 0.1487703 -686.9650253 2.95
CONF17 -43.2432865 2.31 -687.0884564 -0.0284444 0.1481513 -686.9687495 0.62
CONF18 -43.2426978 2.67 -687.0876507 -0.0267722 0.1484622 -686.9659607 2.37
CONF19 -43.2420410 3.09 -687.0884940 -0.0281084 0.1490372 -686.9675652 1.36
CONF21 -43.2427741 2.63 -687.0873013 -0.0297523 0.1480274 -686.9690262 0.44
CONF24 -43.2422634 2.95 -687.0914313 -0.0224162 0.1483511 -686.9654964 2.66
CONF25 -43.2418376 3.21 -687.0880110 -0.0289133 0.1487583 -686.9681660 0.98
CONF26 -43.2423898 2.87 -687.0903285 -0.0241722 0.1481367 -686.9663641 2.11
CONF27 -43.2420553 3.08 -687.0890720 -0.0246747 0.1483815 -686.9653652 2.74
CONF29 -43.2420668 3.07 -687.0904874 -0.0250550 0.1481445 -686.9673979 1.47
CONF31 -43.2413915 3.49 -687.0880936 -0.0297593 0.1481361 -686.9697168 0.01
CONF43 -43.2405006 4.05 -687.0865504 -0.0303700 0.1481309 -686.9687895 0.59
CONF46 -43.2402359 4.22 -687.0892486 -0.0252119 0.1480702 -686.9663903 2.10
CONF51 -43.2379281 5.67 -687.0854275 -0.0291359 0.1483811 -686.9661823 2.23
Number of conformers observed within the following ΔG windows:
ΔG [kcal/mol] #CONF sum(Boltzmann_weights)
---------------------------------------------
0 - 0.5 6 0.55
0 - 1.0 14 0.90
0 - 1.5 18 0.97
0 - 2.0 19 0.98
0 - 2.5 24 0.99
0 - 3.0 29 1.00
---------------------------------------------
Additional global 'fuzzy-threshold' based on the standard deviation of (G_mRRHO):
Std_dev(G_mRRHO) = 0.239 kcal/mol
Fuzzythreshold = 0.249 kcal/mol
Final sorting threshold G_thr(1) = 3.000 + 0.249 = 3.249 kcal/mol
Spearman correlation coefficient between (E + Solv) and (E + Solv + mRRHO) = 0.928
All relative (free) energies are below the initial G_thr(1) threshold of 3.0 kcal/mol.
All conformers are considered further.
Calculating Boltzmann averaged free energy of ensemble on input geometries (not DFT optimized)!
temperature /K: avE(T) /a.u. avGmRRHO(T) /a.u. avGsolv(T) /a.u. avG(T) /a.u.
----------------------------------------------------------------------------------------------------
298.15 -687.0898345 0.1485567 -0.0276741 -686.9689519 <<==part1==
----------------------------------------------------------------------------------------------------
Calculating unbiased GFNn-xTB energy
Constructed folders!
Starting 29 xTB - single-point calculations.
gfn2-xTB energy for CONF1/GFN_unbiased = -43.3944150
gfn2-xTB energy for CONF2/GFN_unbiased = -43.3941370
gfn2-xTB energy for CONF3/GFN_unbiased = -43.3938279
gfn2-xTB energy for CONF4/GFN_unbiased = -43.3924509
gfn2-xTB energy for CONF5/GFN_unbiased = -43.3922430
gfn2-xTB energy for CONF6/GFN_unbiased = -43.3908696
gfn2-xTB energy for CONF7/GFN_unbiased = -43.3907915
gfn2-xTB energy for CONF9/GFN_unbiased = -43.3906140
gfn2-xTB energy for CONF8/GFN_unbiased = -43.3906693
gfn2-xTB energy for CONF11/GFN_unbiased = -43.3903938
gfn2-xTB energy for CONF10/GFN_unbiased = -43.3905816
gfn2-xTB energy for CONF12/GFN_unbiased = -43.3903248
gfn2-xTB energy for CONF13/GFN_unbiased = -43.3902228
gfn2-xTB energy for CONF14/GFN_unbiased = -43.3901776
gfn2-xTB energy for CONF16/GFN_unbiased = -43.3898750
gfn2-xTB energy for CONF15/GFN_unbiased = -43.3899521
gfn2-xTB energy for CONF17/GFN_unbiased = -43.3896242
gfn2-xTB energy for CONF18/GFN_unbiased = -43.3893461
gfn2-xTB energy for CONF19/GFN_unbiased = -43.3892644
gfn2-xTB energy for CONF21/GFN_unbiased = -43.3889876
gfn2-xTB energy for CONF25/GFN_unbiased = -43.3887821
gfn2-xTB energy for CONF24/GFN_unbiased = -43.3888012
gfn2-xTB energy for CONF26/GFN_unbiased = -43.3887129
gfn2-xTB energy for CONF27/GFN_unbiased = -43.3886231
gfn2-xTB energy for CONF29/GFN_unbiased = -43.3883958
gfn2-xTB energy for CONF31/GFN_unbiased = -43.3877136
gfn2-xTB energy for CONF43/GFN_unbiased = -43.3868177
gfn2-xTB energy for CONF51/GFN_unbiased = -43.3844952
gfn2-xTB energy for CONF46/GFN_unbiased = -43.3864925
Tasks completed!
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>END of Part1<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
Ran part1 in 214.0779 seconds
----------------------------------------------------------------------------------------------------
CRE OPTIMIZATION - PART2
----------------------------------------------------------------------------------------------------
program: tm
functional for part2: r2scan-3c
basis set for part2: def2-mTZVPP
using the xTB-optimizer for optimization: on
using the new ensemble optimizer: on
optimize all conformers below this G_thr(opt,2) threshold: 2.5
Spearman threshold: 0.936
number of optimization iterations: 8
radsize: 10
optimization level in part2: lax
solvent: h2o
solvent model applied in the optimization: dcosmors
solvent model for Gsolv contribution: cosmors
temperature: 298.15
evalulate at different temperatures: on
temperature range: 273.15, 278.15, 283.15, 288.15, ...
Boltzmann sum threshold G_thr(2) for sorting in part2: 99.0
calculate mRRHO contribution: on
program for mRRHO contribution: xtb
GFN version for mRRHO and/or GBSA_Gsolv: gfn2
Apply constraint to input geometry during mRRHO calculation: on
Optimizing geometries at DFT level with implicit solvation!
The optimization is calculated for:
CONF1, CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF10, CONF11
CONF12, CONF13, CONF14, CONF15, CONF16, CONF17, CONF18, CONF19, CONF21, CONF24, CONF25
CONF26, CONF27, CONF29, CONF31, CONF43, CONF46, CONF51
Constructed folders!
Preparing 29 calculations.
Tasks completed!
************************Starting optimizations************************
Starting threshold is set to 2.5 + 60.0 % = 4.0 kcal/mol
Lower limit is set to G_thr(opt,2) = 2.5 kcal/mol
*******************************CYCLE 1********************************
Starting 29 optimizations.
Running optimization in CONF1/r2scan-3c
Running optimization in CONF2/r2scan-3c
Running optimization in CONF3/r2scan-3c
Running optimization in CONF4/r2scan-3c
Running optimization in CONF5/r2scan-3c
Running optimization in CONF6/r2scan-3c
Running optimization in CONF7/r2scan-3c
Running optimization in CONF8/r2scan-3c
Running optimization in CONF9/r2scan-3c
Running optimization in CONF10/r2scan-3c
Running optimization in CONF11/r2scan-3c
Running optimization in CONF12/r2scan-3c
Running optimization in CONF13/r2scan-3c
Running optimization in CONF14/r2scan-3c
Running optimization in CONF15/r2scan-3c
Running optimization in CONF16/r2scan-3c
Running optimization in CONF17/r2scan-3c
Running optimization in CONF18/r2scan-3c
Running optimization in CONF19/r2scan-3c
Running optimization in CONF21/r2scan-3c
Running optimization in CONF24/r2scan-3c
Running optimization in CONF25/r2scan-3c
Running optimization in CONF26/r2scan-3c
Running optimization in CONF27/r2scan-3c
Running optimization in CONF29/r2scan-3c
Running optimization in CONF31/r2scan-3c
Running optimization in CONF43/r2scan-3c
Running optimization in CONF46/r2scan-3c
Running optimization in CONF51/r2scan-3c
Tasks completed!
Geometry optimization converged for: CONF4 within 8 cycles
Constructed folders!
Starting 29 G_RRHO calculations.
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Running GFN2-xTB mRRHO in r2scan-3c/rrho_crude
Tasks completed!
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF1/r2scan-3c/rrho_crude: 0.1491729 S_rot(sym)= 0.0000000 using= 0.1491729
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF2/r2scan-3c/rrho_crude: 0.1489606 S_rot(sym)= 0.0000000 using= 0.1489606
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF3/r2scan-3c/rrho_crude: 0.1488268 S_rot(sym)= 0.0000000 using= 0.1488268
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF4/r2scan-3c/rrho_crude: 0.1484927 S_rot(sym)= 0.0000000 using= 0.1484927
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF5/r2scan-3c/rrho_crude: 0.1486893 S_rot(sym)= 0.0000000 using= 0.1486893
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF6/r2scan-3c/rrho_crude: 0.1487938 S_rot(sym)= 0.0000000 using= 0.1487938
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF7/r2scan-3c/rrho_crude: 0.1485369 S_rot(sym)= 0.0000000 using= 0.1485369
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF8/r2scan-3c/rrho_crude: 0.1483310 S_rot(sym)= 0.0000000 using= 0.1483310
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF9/r2scan-3c/rrho_crude: 0.1490465 S_rot(sym)= 0.0000000 using= 0.1490465
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF10/r2scan-3c/rrho_crude: 0.1483287 S_rot(sym)= 0.0000000 using= 0.1483287
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF11/r2scan-3c/rrho_crude: 0.1486458 S_rot(sym)= 0.0000000 using= 0.1486458
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF12/r2scan-3c/rrho_crude: 0.1483757 S_rot(sym)= 0.0000000 using= 0.1483757
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF13/r2scan-3c/rrho_crude: 0.1483694 S_rot(sym)= 0.0000000 using= 0.1483694
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF14/r2scan-3c/rrho_crude: 0.1485930 S_rot(sym)= 0.0000000 using= 0.1485930
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF15/r2scan-3c/rrho_crude: 0.1486309 S_rot(sym)= 0.0000000 using= 0.1486309
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF16/r2scan-3c/rrho_crude: 0.1480533 S_rot(sym)= 0.0000000 using= 0.1480533
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF17/r2scan-3c/rrho_crude: 0.1481168 S_rot(sym)= 0.0000000 using= 0.1481168
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF18/r2scan-3c/rrho_crude: 0.1483304 S_rot(sym)= 0.0000000 using= 0.1483304
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF19/r2scan-3c/rrho_crude: 0.1488035 S_rot(sym)= 0.0000000 using= 0.1488035
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF21/r2scan-3c/rrho_crude: 0.1479899 S_rot(sym)= 0.0000000 using= 0.1479899
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF24/r2scan-3c/rrho_crude: 0.1483497 S_rot(sym)= 0.0000000 using= 0.1483497
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF25/r2scan-3c/rrho_crude: 0.1486322 S_rot(sym)= 0.0000000 using= 0.1486322
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF26/r2scan-3c/rrho_crude: 0.1480434 S_rot(sym)= 0.0000000 using= 0.1480434
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF27/r2scan-3c/rrho_crude: 0.1481630 S_rot(sym)= 0.0000000 using= 0.1481630
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF29/r2scan-3c/rrho_crude: 0.1481791 S_rot(sym)= 0.0000000 using= 0.1481791
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF31/r2scan-3c/rrho_crude: 0.1478742 S_rot(sym)= 0.0000000 using= 0.1478742
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF43/r2scan-3c/rrho_crude: 0.1480666 S_rot(sym)= 0.0000000 using= 0.1480666
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF46/r2scan-3c/rrho_crude: 0.1480836 S_rot(sym)= 0.0000000 using= 0.1480836
The G_mRRHO calculation on crudely optimized DFT geometry @ c1 was successful for CONF51/r2scan-3c/rrho_crude: 0.1483798 S_rot(sym)= 0.0000000 using= 0.1483798
Max number of performed iterations: 8
Spearman rank evaluation is performed in the next cycle.
CONF1 initial ΔG = 3.87 kcal/mol and current ΔG = 3.12 kcal/mol. (not_converged)
CONF2 initial ΔG = 1.63 kcal/mol and current ΔG = 1.33 kcal/mol. (not_converged)
CONF3 initial ΔG = 1.54 kcal/mol and current ΔG = 1.01 kcal/mol. (not_converged)
CONF4 initial ΔG = 1.03 kcal/mol and current ΔG = 0.78 kcal/mol. (converged)
CONF5 initial ΔG = 1.11 kcal/mol and current ΔG = 0.64 kcal/mol. (not_converged)
CONF6 initial ΔG = 2.81 kcal/mol and current ΔG = 2.46 kcal/mol. (not_converged)
CONF7 initial ΔG = 1.23 kcal/mol and current ΔG = 0.56 kcal/mol. (not_converged)
CONF8 initial ΔG = 3.05 kcal/mol and current ΔG = 0.67 kcal/mol. (not_converged)
CONF9 initial ΔG = 2.37 kcal/mol and current ΔG = 2.41 kcal/mol. (not_converged)
CONF10 initial ΔG = 3.48 kcal/mol and current ΔG = 3.06 kcal/mol. (not_converged)
CONF11 initial ΔG = 0.56 kcal/mol and current ΔG = 0.58 kcal/mol. (not_converged)
CONF12 initial ΔG = 1.00 kcal/mol and current ΔG = 0.53 kcal/mol. (not_converged)
CONF13 initial ΔG = 1.23 kcal/mol and current ΔG = 0.26 kcal/mol. (not_converged)
CONF14 initial ΔG = 1.10 kcal/mol and current ΔG = 0.38 kcal/mol. (not_converged)
CONF15 initial ΔG = 0.99 kcal/mol and current ΔG = 0.49 kcal/mol. (not_converged)
CONF16 initial ΔG = 3.27 kcal/mol and current ΔG = 2.55 kcal/mol. (not_converged)
CONF17 initial ΔG = 1.31 kcal/mol and current ΔG = 0.74 kcal/mol. (not_converged)
CONF18 initial ΔG = 3.64 kcal/mol and current ΔG = 2.59 kcal/mol. (not_converged)
CONF19 initial ΔG = 2.20 kcal/mol and current ΔG = 2.11 kcal/mol. (not_converged)
CONF21 initial ΔG = 0.51 kcal/mol and current ΔG = 0.14 kcal/mol. (not_converged)
CONF24 initial ΔG = 3.62 kcal/mol and current ΔG = 3.31 kcal/mol. (not_converged)
CONF25 initial ΔG = 2.78 kcal/mol and current ΔG = 2.51 kcal/mol. (not_converged)
CONF26 initial ΔG = 3.30 kcal/mol and current ΔG = 2.98 kcal/mol. (not_converged)
CONF27 initial ΔG = 3.21 kcal/mol and current ΔG = 2.51 kcal/mol. (not_converged)
CONF29 initial ΔG = 2.73 kcal/mol and current ΔG = 2.67 kcal/mol. (not_converged)
CONF31 initial ΔG = 0.00 kcal/mol and current ΔG = 0.29 kcal/mol. (not_converged)
CONF43 initial ΔG = 0.57 kcal/mol and current ΔG = 0.00 kcal/mol. (not_converged)
CONF46 initial ΔG = 2.74 kcal/mol and current ΔG = 2.56 kcal/mol. (not_converged)
CONF51 initial ΔG = 3.44 kcal/mol and current ΔG = 3.72 kcal/mol. (not_converged)
CYCLE 1 performed in 2367.5719 seconds
*******************************CYCLE 2********************************
Starting 28 optimizations.
Running optimization in CONF1/r2scan-3c
Running optimization in CONF2/r2scan-3c
Running optimization in CONF3/r2scan-3c
Running optimization in CONF5/r2scan-3c
Running optimization in CONF6/r2scan-3c
Running optimization in CONF7/r2scan-3c
Running optimization in CONF8/r2scan-3c
Running optimization in CONF9/r2scan-3c
Running optimization in CONF10/r2scan-3c
Running optimization in CONF11/r2scan-3c
Running optimization in CONF12/r2scan-3c
Running optimization in CONF13/r2scan-3c
Running optimization in CONF14/r2scan-3c
Running optimization in CONF15/r2scan-3c
Running optimization in CONF16/r2scan-3c
Running optimization in CONF17/r2scan-3c
Running optimization in CONF18/r2scan-3c
Running optimization in CONF19/r2scan-3c
Running optimization in CONF21/r2scan-3c
Running optimization in CONF24/r2scan-3c
Running optimization in CONF25/r2scan-3c
Running optimization in CONF26/r2scan-3c
Running optimization in CONF27/r2scan-3c
Running optimization in CONF29/r2scan-3c
Running optimization in CONF31/r2scan-3c
Running optimization in CONF43/r2scan-3c
Running optimization in CONF46/r2scan-3c
Running optimization in CONF51/r2scan-3c
Tasks completed!
Geometry optimization converged for: CONF5 within 9 cycles
Geometry optimization converged for: CONF6 within 16 cycles
Geometry optimization converged for: CONF11 within 9 cycles
Geometry optimization converged for: CONF12 within 9 cycles
Geometry optimization converged for: CONF13 within 13 cycles
Geometry optimization converged for: CONF15 within 10 cycles
Geometry optimization converged for: CONF17 within 16 cycles
Geometry optimization converged for: CONF19 within 12 cycles
Geometry optimization converged for: CONF21 within 12 cycles
Geometry optimization converged for: CONF25 within 14 cycles
Geometry optimization converged for: CONF26 within 10 cycles
Geometry optimization converged for: CONF31 within 16 cycles
Geometry optimization converged for: CONF46 within 11 cycles
Geometry optimization converged for: CONF51 within 9 cycles
Max number of performed iterations: 16
CONF1 previous ΔG = 3.12 kcal/mol and current ΔG = 3.07 kcal/mol. (not_converged)
CONF2 previous ΔG = 1.33 kcal/mol and current ΔG = 1.52 kcal/mol. (not_converged)
CONF3 previous ΔG = 1.01 kcal/mol and current ΔG = 1.19 kcal/mol. (not_converged)
CONF4 previous ΔG = 0.78 kcal/mol and current ΔG = 1.05 kcal/mol. (converged)
CONF5 previous ΔG = 0.64 kcal/mol and current ΔG = 0.91 kcal/mol. (converged)
CONF6 previous ΔG = 2.46 kcal/mol and current ΔG = 2.46 kcal/mol. (converged)
CONF7 previous ΔG = 0.56 kcal/mol and current ΔG = 1.60 kcal/mol. (not_converged)
CONF8 previous ΔG = 0.67 kcal/mol and current ΔG = 0.70 kcal/mol. (not_converged)
CONF9 previous ΔG = 2.41 kcal/mol and current ΔG = 2.58 kcal/mol. (not_converged)
CONF10 previous ΔG = 3.06 kcal/mol and current ΔG = 2.99 kcal/mol. (not_converged)
CONF11 previous ΔG = 0.58 kcal/mol and current ΔG = 0.84 kcal/mol. (converged)
CONF12 previous ΔG = 0.53 kcal/mol and current ΔG = 0.80 kcal/mol. (converged)
CONF13 previous ΔG = 0.26 kcal/mol and current ΔG = 0.36 kcal/mol. (converged)
CONF14 previous ΔG = 0.38 kcal/mol and current ΔG = 0.53 kcal/mol. (not_converged)
CONF15 previous ΔG = 0.49 kcal/mol and current ΔG = 0.70 kcal/mol. (converged)
CONF16 previous ΔG = 2.55 kcal/mol and current ΔG = 2.48 kcal/mol. (not_converged)
CONF17 previous ΔG = 0.74 kcal/mol and current ΔG = 0.70 kcal/mol. (converged)
CONF18 previous ΔG = 2.59 kcal/mol and current ΔG = 2.33 kcal/mol. (not_converged)
CONF19 previous ΔG = 2.11 kcal/mol and current ΔG = 2.35 kcal/mol. (converged)
CONF21 previous ΔG = 0.14 kcal/mol and current ΔG = 0.35 kcal/mol. (converged)
CONF24 previous ΔG = 3.31 kcal/mol and current ΔG = 3.35 kcal/mol. (not_converged)
CONF25 previous ΔG = 2.51 kcal/mol and current ΔG = 2.80 kcal/mol. (converged)
CONF26 previous ΔG = 2.98 kcal/mol and current ΔG = 3.21 kcal/mol. (converged)
CONF27 previous ΔG = 2.51 kcal/mol and current ΔG = 2.53 kcal/mol. (not_converged)
CONF29 previous ΔG = 2.67 kcal/mol and current ΔG = 3.00 kcal/mol. (not_converged)
CONF31 previous ΔG = 0.29 kcal/mol and current ΔG = 0.42 kcal/mol. (converged)
CONF43 previous ΔG = 0.00 kcal/mol and current ΔG = 0.00 kcal/mol. (not_converged)
CONF46 previous ΔG = 2.56 kcal/mol and current ΔG = 2.82 kcal/mol. (converged)
CONF51 previous ΔG = 3.72 kcal/mol and current ΔG = 3.98 kcal/mol. (converged)
Spearman coeff. from 8 --> 11 = 0.9695
Spearman coeff. from 9 --> 12 = 0.9906
Spearman coeff. from 10 --> 13 = 0.9931
Spearman coeff. from 11 --> 14 = 0.9818
Evaluating Spearman coeff. from 12 --> 15 = 0.9897
Evaluating Spearman coeff. from 13 --> 16 = 0.9660
Final averaged Spearman correlation coefficient: 0.9778
PES is assumed to be parallel
Updated optimization threshold to: 3.50 kcal/mol
CYCLE 2 performed in 1577.6956 seconds
*******************************CYCLE 3********************************
Starting 14 optimizations.
Running optimization in CONF1/r2scan-3c
Running optimization in CONF2/r2scan-3c
Running optimization in CONF3/r2scan-3c
Running optimization in CONF7/r2scan-3c
Running optimization in CONF8/r2scan-3c
Running optimization in CONF9/r2scan-3c
Running optimization in CONF10/r2scan-3c
Running optimization in CONF14/r2scan-3c
Running optimization in CONF16/r2scan-3c
Running optimization in CONF18/r2scan-3c
Running optimization in CONF24/r2scan-3c
Running optimization in CONF27/r2scan-3c
Running optimization in CONF29/r2scan-3c
Running optimization in CONF43/r2scan-3c
Tasks completed!
Geometry optimization converged for: CONF2 within 20 cycles
Geometry optimization converged for: CONF3 within 21 cycles
Geometry optimization converged for: CONF9 within 19 cycles
Geometry optimization converged for: CONF10 within 18 cycles
Geometry optimization converged for: CONF14 within 18 cycles
Geometry optimization converged for: CONF16 within 17 cycles
Geometry optimization converged for: CONF18 within 18 cycles
Geometry optimization converged for: CONF43 within 23 cycles
Max number of performed iterations: 24
CONF1 previous ΔG = 3.07 kcal/mol and current ΔG = 3.02 kcal/mol. (not_converged)
CONF2 previous ΔG = 1.52 kcal/mol and current ΔG = 1.45 kcal/mol. (converged)
CONF3 previous ΔG = 1.19 kcal/mol and current ΔG = 1.19 kcal/mol. (converged)
CONF4 previous ΔG = 1.05 kcal/mol and current ΔG = 1.06 kcal/mol. (converged)
CONF5 previous ΔG = 0.91 kcal/mol and current ΔG = 0.92 kcal/mol. (converged)
CONF6 previous ΔG = 2.46 kcal/mol and current ΔG = 2.47 kcal/mol. (converged)
CONF7 previous ΔG = 1.60 kcal/mol and current ΔG = 0.71 kcal/mol. (not_converged)
CONF8 previous ΔG = 0.70 kcal/mol and current ΔG = 0.78 kcal/mol. (not_converged)
CONF9 previous ΔG = 2.58 kcal/mol and current ΔG = 2.59 kcal/mol. (converged)
CONF10 previous ΔG = 2.99 kcal/mol and current ΔG = 3.04 kcal/mol. (converged)
CONF11 previous ΔG = 0.84 kcal/mol and current ΔG = 0.85 kcal/mol. (converged)
CONF12 previous ΔG = 0.80 kcal/mol and current ΔG = 0.81 kcal/mol. (converged)
CONF13 previous ΔG = 0.36 kcal/mol and current ΔG = 0.37 kcal/mol. (converged)
CONF14 previous ΔG = 0.53 kcal/mol and current ΔG = 0.53 kcal/mol. (converged)
CONF15 previous ΔG = 0.70 kcal/mol and current ΔG = 0.71 kcal/mol. (converged)
CONF16 previous ΔG = 2.48 kcal/mol and current ΔG = 2.49 kcal/mol. (converged)
CONF17 previous ΔG = 0.70 kcal/mol and current ΔG = 0.71 kcal/mol. (converged)
CONF18 previous ΔG = 2.33 kcal/mol and current ΔG = 2.31 kcal/mol. (converged)
CONF19 previous ΔG = 2.35 kcal/mol and current ΔG = 2.37 kcal/mol. (converged)
CONF21 previous ΔG = 0.35 kcal/mol and current ΔG = 0.36 kcal/mol. (converged)
CONF24 previous ΔG = 3.35 kcal/mol and current ΔG = 3.29 kcal/mol. (not_converged)
CONF25 previous ΔG = 2.80 kcal/mol and current ΔG = 2.81 kcal/mol. (converged)
CONF26 previous ΔG = 3.21 kcal/mol and current ΔG = 3.22 kcal/mol. (converged)
CONF27 previous ΔG = 2.53 kcal/mol and current ΔG = 2.30 kcal/mol. (not_converged)
CONF29 previous ΔG = 3.00 kcal/mol and current ΔG = 2.94 kcal/mol. (not_converged)
CONF31 previous ΔG = 0.42 kcal/mol and current ΔG = 0.43 kcal/mol. (converged)
CONF43 previous ΔG = 0.00 kcal/mol and current ΔG = 0.00 kcal/mol. (converged)
CONF46 previous ΔG = 2.82 kcal/mol and current ΔG = 2.83 kcal/mol. (converged)
CONF51 previous ΔG = 3.98 kcal/mol and current ΔG = 4.00 kcal/mol. (converged)
Spearman coeff. from 16 --> 19 = 0.9916
Spearman coeff. from 17 --> 20 = 0.9975
Spearman coeff. from 18 --> 21 = 0.9143
Spearman coeff. from 19 --> 22 = 0.9980
Evaluating Spearman coeff. from 20 --> 23 = 0.9990
Evaluating Spearman coeff. from 21 --> 24 = 0.8739
Final averaged Spearman correlation coefficient: 0.9365
PES is assumed to be parallel
Updated optimization threshold to: 3.00 kcal/mol
CONF1 is above 3.0 kcal/mol and gradient norm (0.00159487087) is below 0.01.
CONF1 is removed because of the lowered threshold!
CONF1 is above threshold, dont optimize further and remove conformer.
CONF24 is above 3.0 kcal/mol and gradient norm (0.004389820046) is below 0.01.
CONF24 is removed because of the lowered threshold!
CONF24 is above threshold, dont optimize further and remove conformer.
CYCLE 3 performed in 736.4307 seconds
*******************************CYCLE 4********************************
Starting 4 optimizations.
Running optimization in CONF7/r2scan-3c
Running optimization in CONF8/r2scan-3c
Running optimization in CONF27/r2scan-3c
Running optimization in CONF29/r2scan-3c
Tasks completed!
Geometry optimization converged for: CONF7 within 25 cycles
Geometry optimization converged for: CONF29 within 28 cycles
Max number of performed iterations: 32
CONF2 previous ΔG = 1.45 kcal/mol and current ΔG = 1.45 kcal/mol. (converged)
CONF3 previous ΔG = 1.19 kcal/mol and current ΔG = 1.19 kcal/mol. (converged)
CONF4 previous ΔG = 1.06 kcal/mol and current ΔG = 1.06 kcal/mol. (converged)
CONF5 previous ΔG = 0.92 kcal/mol and current ΔG = 0.92 kcal/mol. (converged)
CONF6 previous ΔG = 2.47 kcal/mol and current ΔG = 2.47 kcal/mol. (converged)
CONF7 previous ΔG = 0.71 kcal/mol and current ΔG = 0.70 kcal/mol. (converged)
CONF8 previous ΔG = 0.78 kcal/mol and current ΔG = 0.76 kcal/mol. (not_converged)
CONF9 previous ΔG = 2.59 kcal/mol and current ΔG = 2.59 kcal/mol. (converged)
CONF10 previous ΔG = 3.04 kcal/mol and current ΔG = 3.04 kcal/mol. (converged)
CONF11 previous ΔG = 0.85 kcal/mol and current ΔG = 0.85 kcal/mol. (converged)
CONF12 previous ΔG = 0.81 kcal/mol and current ΔG = 0.81 kcal/mol. (converged)
CONF13 previous ΔG = 0.37 kcal/mol and current ΔG = 0.37 kcal/mol. (converged)
CONF14 previous ΔG = 0.53 kcal/mol and current ΔG = 0.53 kcal/mol. (converged)
CONF15 previous ΔG = 0.71 kcal/mol and current ΔG = 0.71 kcal/mol. (converged)
CONF16 previous ΔG = 2.49 kcal/mol and current ΔG = 2.49 kcal/mol. (converged)
CONF17 previous ΔG = 0.71 kcal/mol and current ΔG = 0.71 kcal/mol. (converged)
CONF18 previous ΔG = 2.31 kcal/mol and current ΔG = 2.31 kcal/mol. (converged)
CONF19 previous ΔG = 2.37 kcal/mol and current ΔG = 2.37 kcal/mol. (converged)
CONF21 previous ΔG = 0.36 kcal/mol and current ΔG = 0.36 kcal/mol. (converged)
CONF25 previous ΔG = 2.81 kcal/mol and current ΔG = 2.81 kcal/mol. (converged)
CONF26 previous ΔG = 3.22 kcal/mol and current ΔG = 3.22 kcal/mol. (converged)
CONF27 previous ΔG = 2.30 kcal/mol and current ΔG = 2.00 kcal/mol. (not_converged)
CONF29 previous ΔG = 2.94 kcal/mol and current ΔG = 2.88 kcal/mol. (converged)
CONF31 previous ΔG = 0.43 kcal/mol and current ΔG = 0.43 kcal/mol. (converged)
CONF43 previous ΔG = 0.00 kcal/mol and current ΔG = 0.00 kcal/mol. (converged)
CONF46 previous ΔG = 2.83 kcal/mol and current ΔG = 2.83 kcal/mol. (converged)
CONF51 previous ΔG = 4.00 kcal/mol and current ΔG = 4.00 kcal/mol. (converged)
Spearman coeff. from 24 --> 27 = 0.9591
Spearman coeff. from 25 --> 28 = 1.0000
Spearman coeff. from 26 --> 29 = 1.0000
Spearman coeff. from 27 --> 30 = 0.9994
Evaluating Spearman coeff. from 28 --> 31 = 1.0000
Evaluating Spearman coeff. from 29 --> 32 = 1.0000
Final averaged Spearman correlation coefficient: 1.0000
PES is assumed to be parallel
Updated optimization threshold to: 2.50 kcal/mol
CYCLE 4 performed in 502.0425 seconds
*******************************CYCLE 5********************************
Starting 2 optimizations.
Running optimization in CONF8/r2scan-3c
Running optimization in CONF27/r2scan-3c
Tasks completed!
Geometry optimization converged for: CONF8 within 34 cycles
Max number of performed iterations: 40
CONF2 previous ΔG = 1.45 kcal/mol and current ΔG = 1.45 kcal/mol. (converged)
CONF3 previous ΔG = 1.19 kcal/mol and current ΔG = 1.19 kcal/mol. (converged)
CONF4 previous ΔG = 1.06 kcal/mol and current ΔG = 1.06 kcal/mol. (converged)
CONF5 previous ΔG = 0.92 kcal/mol and current ΔG = 0.92 kcal/mol. (converged)
CONF6 previous ΔG = 2.47 kcal/mol and current ΔG = 2.47 kcal/mol. (converged)
CONF7 previous ΔG = 0.70 kcal/mol and current ΔG = 0.70 kcal/mol. (converged)
CONF8 previous ΔG = 0.76 kcal/mol and current ΔG = 0.72 kcal/mol. (converged)
CONF9 previous ΔG = 2.59 kcal/mol and current ΔG = 2.59 kcal/mol. (converged)
CONF10 previous ΔG = 3.04 kcal/mol and current ΔG = 3.04 kcal/mol. (converged)
CONF11 previous ΔG = 0.85 kcal/mol and current ΔG = 0.85 kcal/mol. (converged)
CONF12 previous ΔG = 0.81 kcal/mol and current ΔG = 0.81 kcal/mol. (converged)
CONF13 previous ΔG = 0.37 kcal/mol and current ΔG = 0.37 kcal/mol. (converged)
CONF14 previous ΔG = 0.53 kcal/mol and current ΔG = 0.53 kcal/mol. (converged)
CONF15 previous ΔG = 0.71 kcal/mol and current ΔG = 0.71 kcal/mol. (converged)
CONF16 previous ΔG = 2.49 kcal/mol and current ΔG = 2.49 kcal/mol. (converged)
CONF17 previous ΔG = 0.71 kcal/mol and current ΔG = 0.71 kcal/mol. (converged)
CONF18 previous ΔG = 2.31 kcal/mol and current ΔG = 2.31 kcal/mol. (converged)
CONF19 previous ΔG = 2.37 kcal/mol and current ΔG = 2.37 kcal/mol. (converged)
CONF21 previous ΔG = 0.36 kcal/mol and current ΔG = 0.36 kcal/mol. (converged)
CONF25 previous ΔG = 2.81 kcal/mol and current ΔG = 2.81 kcal/mol. (converged)
CONF26 previous ΔG = 3.22 kcal/mol and current ΔG = 3.22 kcal/mol. (converged)
CONF27 previous ΔG = 2.00 kcal/mol and current ΔG = 2.18 kcal/mol. (not_converged)
CONF29 previous ΔG = 2.88 kcal/mol and current ΔG = 2.88 kcal/mol. (converged)
CONF31 previous ΔG = 0.43 kcal/mol and current ΔG = 0.43 kcal/mol. (converged)
CONF43 previous ΔG = 0.00 kcal/mol and current ΔG = 0.00 kcal/mol. (converged)
CONF46 previous ΔG = 2.83 kcal/mol and current ΔG = 2.83 kcal/mol. (converged)
CONF51 previous ΔG = 4.00 kcal/mol and current ΔG = 4.00 kcal/mol. (converged)
Spearman coeff. from 32 --> 35 = 1.0000
Spearman coeff. from 33 --> 36 = 1.0000
Spearman coeff. from 34 --> 37 = 0.9597
Spearman coeff. from 35 --> 38 = 1.0000
Evaluating Spearman coeff. from 36 --> 39 = 1.0000
Evaluating Spearman coeff. from 37 --> 40 = 0.9597
Final averaged Spearman correlation coefficient: 0.9799
PES is assumed to be parallel
Current optimization threshold: 2.50 kcal/mol
CYCLE 5 performed in 403.7743 seconds
*******************************CYCLE 6********************************
Starting 1 optimizations.
Running optimization in CONF27/r2scan-3c
Tasks completed!
Max number of performed iterations: 48
CONF2 previous ΔG = 1.45 kcal/mol and current ΔG = 1.45 kcal/mol. (converged)
CONF3 previous ΔG = 1.19 kcal/mol and current ΔG = 1.19 kcal/mol. (converged)
CONF4 previous ΔG = 1.06 kcal/mol and current ΔG = 1.06 kcal/mol. (converged)
CONF5 previous ΔG = 0.92 kcal/mol and current ΔG = 0.92 kcal/mol. (converged)
CONF6 previous ΔG = 2.47 kcal/mol and current ΔG = 2.47 kcal/mol. (converged)
CONF7 previous ΔG = 0.70 kcal/mol and current ΔG = 0.70 kcal/mol. (converged)
CONF8 previous ΔG = 0.72 kcal/mol and current ΔG = 0.72 kcal/mol. (converged)
CONF9 previous ΔG = 2.59 kcal/mol and current ΔG = 2.59 kcal/mol. (converged)
CONF10 previous ΔG = 3.04 kcal/mol and current ΔG = 3.04 kcal/mol. (converged)
CONF11 previous ΔG = 0.85 kcal/mol and current ΔG = 0.85 kcal/mol. (converged)
CONF12 previous ΔG = 0.81 kcal/mol and current ΔG = 0.81 kcal/mol. (converged)
CONF13 previous ΔG = 0.37 kcal/mol and current ΔG = 0.37 kcal/mol. (converged)
CONF14 previous ΔG = 0.53 kcal/mol and current ΔG = 0.53 kcal/mol. (converged)
CONF15 previous ΔG = 0.71 kcal/mol and current ΔG = 0.71 kcal/mol. (converged)
CONF16 previous ΔG = 2.49 kcal/mol and current ΔG = 2.49 kcal/mol. (converged)
CONF17 previous ΔG = 0.71 kcal/mol and current ΔG = 0.71 kcal/mol. (converged)
CONF18 previous ΔG = 2.31 kcal/mol and current ΔG = 2.31 kcal/mol. (converged)
CONF19 previous ΔG = 2.37 kcal/mol and current ΔG = 2.37 kcal/mol. (converged)
CONF21 previous ΔG = 0.36 kcal/mol and current ΔG = 0.36 kcal/mol. (converged)
CONF25 previous ΔG = 2.81 kcal/mol and current ΔG = 2.81 kcal/mol. (converged)
CONF26 previous ΔG = 3.22 kcal/mol and current ΔG = 3.22 kcal/mol. (converged)
CONF27 previous ΔG = 2.18 kcal/mol and current ΔG = 0.29 kcal/mol. (not_converged)
CONF29 previous ΔG = 2.88 kcal/mol and current ΔG = 2.88 kcal/mol. (converged)
CONF31 previous ΔG = 0.43 kcal/mol and current ΔG = 0.43 kcal/mol. (converged)
CONF43 previous ΔG = 0.00 kcal/mol and current ΔG = 0.00 kcal/mol. (converged)
CONF46 previous ΔG = 2.83 kcal/mol and current ΔG = 2.83 kcal/mol. (converged)
CONF51 previous ΔG = 4.00 kcal/mol and current ΔG = 4.00 kcal/mol. (converged)
Spearman coeff. from 40 --> 43 = 0.9664
Spearman coeff. from 41 --> 44 = 0.9982
Spearman coeff. from 42 --> 45 = 0.9872
Spearman coeff. from 43 --> 46 = 0.9982
Evaluating Spearman coeff. from 44 --> 47 = 1.0000
Evaluating Spearman coeff. from 45 --> 48 = 0.9963
Final averaged Spearman correlation coefficient: 0.9982
PES is assumed to be parallel
Current optimization threshold: 2.50 kcal/mol
CYCLE 6 performed in 415.7367 seconds
*******************************CYCLE 7********************************
Starting 1 optimizations.
Running optimization in CONF27/r2scan-3c
Tasks completed!
Geometry optimization converged for: CONF27 within 56 cycles
***********************Finished optimizations!************************
Timings:
Cycle: [s] #nconfs Spearman coeff.
1 2367.57 29
2 1577.70 29 0.978
3 736.43 27 0.936
4 502.04 27 1.000
5 403.77 27 0.980
6 415.74 27 0.998
7 370.96 27
sum: 6374.21
CONVERGED optimizations for the following remaining conformers:
Converged optimization for CONF2 after 20 cycles: -687.1368441
Converged optimization for CONF3 after 21 cycles: -687.1371271
Converged optimization for CONF4 after 8 cycles: -687.1369975
Converged optimization for CONF5 after 9 cycles: -687.1374205
Converged optimization for CONF6 after 16 cycles: -687.1350498
Converged optimization for CONF7 after 25 cycles: -687.1376162
Converged optimization for CONF8 after 34 cycles: -687.1373828
Converged optimization for CONF9 after 19 cycles: -687.1351135
Converged optimization for CONF10 after 18 cycles: -687.1336843
Converged optimization for CONF11 after 9 cycles: -687.1374836
Converged optimization for CONF12 after 9 cycles: -687.1372828
Converged optimization for CONF13 after 13 cycles: -687.1379671
Converged optimization for CONF14 after 18 cycles: -687.1379442
Converged optimization for CONF15 after 10 cycles: -687.1376884
Converged optimization for CONF16 after 17 cycles: -687.1342764
Converged optimization for CONF17 after 16 cycles: -687.1371830
Converged optimization for CONF18 after 18 cycles: -687.1348346
Converged optimization for CONF19 after 12 cycles: -687.1352273
Converged optimization for CONF21 after 12 cycles: -687.1376088
Converged optimization for CONF25 after 14 cycles: -687.1343453
Converged optimization for CONF26 after 10 cycles: -687.1331080
Converged optimization for CONF27 after 56 cycles: -687.1382272
Converged optimization for CONF29 after 28 cycles: -687.1337889
Converged optimization for CONF31 after 16 cycles: -687.1373798
Converged optimization for CONF43 after 23 cycles: -687.1382594
Converged optimization for CONF46 after 11 cycles: -687.1337629
Converged optimization for CONF51 after 9 cycles: -687.1322056
Calculating single-point energies and solvation contribution (G_solv)!
The low level gsolv calculation is now calculated for:
Constructed folders!
CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF10, CONF11, CONF12
CONF13, CONF14, CONF15, CONF16, CONF17, CONF18, CONF19, CONF21, CONF25, CONF26, CONF27
CONF29, CONF31, CONF43, CONF46, CONF51
Starting 27 COSMO-RS-Gsolv calculations.
Running COSMO-RS calculation in CONF2/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF3/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF4/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF5/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF6/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF7/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF8/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF9/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF10/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF11/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF12/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF13/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF14/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF15/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF16/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF17/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF18/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF19/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF21/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF25/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF26/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF27/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF29/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF31/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF43/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF46/r2scan-3c/COSMO
Running COSMO-RS calculation in CONF51/r2scan-3c/COSMO
Tasks completed!
lowlevel COSMO-RS calculation was successful for CONF2/r2scan-3c/COSMO: -0.02793810
lowlevel COSMO-RS calculation was successful for CONF3/r2scan-3c/COSMO: -0.02766926
lowlevel COSMO-RS calculation was successful for CONF4/r2scan-3c/COSMO: -0.03087413
lowlevel COSMO-RS calculation was successful for CONF5/r2scan-3c/COSMO: -0.02996521
lowlevel COSMO-RS calculation was successful for CONF6/r2scan-3c/COSMO: -0.02959240
lowlevel COSMO-RS calculation was successful for CONF7/r2scan-3c/COSMO: -0.03296469
lowlevel COSMO-RS calculation was successful for CONF8/r2scan-3c/COSMO: -0.03163953
lowlevel COSMO-RS calculation was successful for CONF9/r2scan-3c/COSMO: -0.02993054
lowlevel COSMO-RS calculation was successful for CONF10/r2scan-3c/COSMO: -0.03054774
lowlevel COSMO-RS calculation was successful for CONF11/r2scan-3c/COSMO: -0.03227388
lowlevel COSMO-RS calculation was successful for CONF12/r2scan-3c/COSMO: -0.03092105
lowlevel COSMO-RS calculation was successful for CONF13/r2scan-3c/COSMO: -0.03268001
lowlevel COSMO-RS calculation was successful for CONF14/r2scan-3c/COSMO: -0.03307002
lowlevel COSMO-RS calculation was successful for CONF15/r2scan-3c/COSMO: -0.02972748
lowlevel COSMO-RS calculation was successful for CONF16/r2scan-3c/COSMO: -0.03003773
lowlevel COSMO-RS calculation was successful for CONF17/r2scan-3c/COSMO: -0.03390852
lowlevel COSMO-RS calculation was successful for CONF18/r2scan-3c/COSMO: -0.03380992
lowlevel COSMO-RS calculation was successful for CONF19/r2scan-3c/COSMO: -0.03141740
lowlevel COSMO-RS calculation was successful for CONF21/r2scan-3c/COSMO: -0.03347077
lowlevel COSMO-RS calculation was successful for CONF25/r2scan-3c/COSMO: -0.03201466
lowlevel COSMO-RS calculation was successful for CONF26/r2scan-3c/COSMO: -0.02865093
lowlevel COSMO-RS calculation was successful for CONF27/r2scan-3c/COSMO: -0.03772663
lowlevel COSMO-RS calculation was successful for CONF29/r2scan-3c/COSMO: -0.02963006
lowlevel COSMO-RS calculation was successful for CONF31/r2scan-3c/COSMO: -0.03197205
lowlevel COSMO-RS calculation was successful for CONF43/r2scan-3c/COSMO: -0.03650841
lowlevel COSMO-RS calculation was successful for CONF46/r2scan-3c/COSMO: -0.02882594
lowlevel COSMO-RS calculation was successful for CONF51/r2scan-3c/COSMO: -0.03099948
Calculating lowlevel G_mRRHO with implicit solvation on DFT geometry!
The lowlevel G_mRRHO calculation is now performed for:
CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF10, CONF11, CONF12
CONF13, CONF14, CONF15, CONF16, CONF17, CONF18, CONF19, CONF21, CONF25, CONF26, CONF27
CONF29, CONF31, CONF43, CONF46, CONF51
Constructed folders!
Starting 27 G_RRHO calculations.
Running GFN2-xTB mRRHO in CONF2/rrho_part2
Running GFN2-xTB mRRHO in CONF3/rrho_part2
Running GFN2-xTB mRRHO in CONF4/rrho_part2
Running GFN2-xTB mRRHO in CONF5/rrho_part2
Running GFN2-xTB mRRHO in CONF6/rrho_part2
Running GFN2-xTB mRRHO in CONF7/rrho_part2
Running GFN2-xTB mRRHO in CONF8/rrho_part2
Running GFN2-xTB mRRHO in CONF9/rrho_part2
Running GFN2-xTB mRRHO in CONF10/rrho_part2
Running GFN2-xTB mRRHO in CONF11/rrho_part2
Running GFN2-xTB mRRHO in CONF12/rrho_part2
Running GFN2-xTB mRRHO in CONF13/rrho_part2
Running GFN2-xTB mRRHO in CONF14/rrho_part2
Running GFN2-xTB mRRHO in CONF15/rrho_part2
Running GFN2-xTB mRRHO in CONF16/rrho_part2
Running GFN2-xTB mRRHO in CONF17/rrho_part2
Running GFN2-xTB mRRHO in CONF18/rrho_part2
Running GFN2-xTB mRRHO in CONF19/rrho_part2
Running GFN2-xTB mRRHO in CONF21/rrho_part2
Running GFN2-xTB mRRHO in CONF25/rrho_part2
Running GFN2-xTB mRRHO in CONF26/rrho_part2
Running GFN2-xTB mRRHO in CONF27/rrho_part2
Running GFN2-xTB mRRHO in CONF29/rrho_part2
Running GFN2-xTB mRRHO in CONF31/rrho_part2
Running GFN2-xTB mRRHO in CONF43/rrho_part2
Running GFN2-xTB mRRHO in CONF46/rrho_part2
Running GFN2-xTB mRRHO in CONF51/rrho_part2
Tasks completed!
The lowlevel G_mRRHO calculation @ c1 was successful for CONF2/rrho_part2: 0.14871023 S_rot(sym)= 0.0000000 using= 0.1487102
The lowlevel G_mRRHO calculation @ c1 was successful for CONF3/rrho_part2: 0.14874236 S_rot(sym)= 0.0000000 using= 0.1487424
The lowlevel G_mRRHO calculation @ c1 was successful for CONF4/rrho_part2: 0.14849266 S_rot(sym)= 0.0000000 using= 0.1484927
The lowlevel G_mRRHO calculation @ c1 was successful for CONF5/rrho_part2: 0.14869023 S_rot(sym)= 0.0000000 using= 0.1486902
The lowlevel G_mRRHO calculation @ c1 was successful for CONF6/rrho_part2: 0.14879878 S_rot(sym)= 0.0000000 using= 0.1487988
The lowlevel G_mRRHO calculation @ c1 was successful for CONF7/rrho_part2: 0.14832878 S_rot(sym)= 0.0000000 using= 0.1483288
The lowlevel G_mRRHO calculation @ c1 was successful for CONF8/rrho_part2: 0.14806304 S_rot(sym)= 0.0000000 using= 0.1480630
The lowlevel G_mRRHO calculation @ c1 was successful for CONF9/rrho_part2: 0.14904685 S_rot(sym)= 0.0000000 using= 0.1490469
The lowlevel G_mRRHO calculation @ c1 was successful for CONF10/rrho_part2: 0.14763468 S_rot(sym)= 0.0000000 using= 0.1476347
The lowlevel G_mRRHO calculation @ c1 was successful for CONF11/rrho_part2: 0.14864192 S_rot(sym)= 0.0000000 using= 0.1486419
The lowlevel G_mRRHO calculation @ c1 was successful for CONF12/rrho_part2: 0.14837107 S_rot(sym)= 0.0000000 using= 0.1483711
The lowlevel G_mRRHO calculation @ c1 was successful for CONF13/rrho_part2: 0.14820187 S_rot(sym)= 0.0000000 using= 0.1482019
The lowlevel G_mRRHO calculation @ c1 was successful for CONF14/rrho_part2: 0.14845505 S_rot(sym)= 0.0000000 using= 0.1484551
The lowlevel G_mRRHO calculation @ c1 was successful for CONF15/rrho_part2: 0.14860006 S_rot(sym)= 0.0000000 using= 0.1486001
The lowlevel G_mRRHO calculation @ c1 was successful for CONF16/rrho_part2: 0.14769607 S_rot(sym)= 0.0000000 using= 0.1476961
The lowlevel G_mRRHO calculation @ c1 was successful for CONF17/rrho_part2: 0.14789947 S_rot(sym)= 0.0000000 using= 0.1478995
The lowlevel G_mRRHO calculation @ c1 was successful for CONF18/rrho_part2: 0.14810269 S_rot(sym)= 0.0000000 using= 0.1481027
The lowlevel G_mRRHO calculation @ c1 was successful for CONF19/rrho_part2: 0.14879913 S_rot(sym)= 0.0000000 using= 0.1487991
The lowlevel G_mRRHO calculation @ c1 was successful for CONF21/rrho_part2: 0.14804882 S_rot(sym)= 0.0000000 using= 0.1480488
The lowlevel G_mRRHO calculation @ c1 was successful for CONF25/rrho_part2: 0.14861542 S_rot(sym)= 0.0000000 using= 0.1486154
The lowlevel G_mRRHO calculation @ c1 was successful for CONF26/rrho_part2: 0.14803643 S_rot(sym)= 0.0000000 using= 0.1480364
The lowlevel G_mRRHO calculation @ c1 was successful for CONF27/rrho_part2: 0.14783227 S_rot(sym)= 0.0000000 using= 0.1478323
The lowlevel G_mRRHO calculation @ c1 was successful for CONF29/rrho_part2: 0.14775858 S_rot(sym)= 0.0000000 using= 0.1477586
The lowlevel G_mRRHO calculation @ c1 was successful for CONF31/rrho_part2: 0.14812484 S_rot(sym)= 0.0000000 using= 0.1481248
The lowlevel G_mRRHO calculation @ c1 was successful for CONF43/rrho_part2: 0.14760766 S_rot(sym)= 0.0000000 using= 0.1476077
The lowlevel G_mRRHO calculation @ c1 was successful for CONF46/rrho_part2: 0.14807609 S_rot(sym)= 0.0000000 using= 0.1480761
The lowlevel G_mRRHO calculation @ c1 was successful for CONF51/rrho_part2: 0.14838215 S_rot(sym)= 0.0000000 using= 0.1483822
--------------------------------------------------
* Gibbs free energies of part2 *
--------------------------------------------------
CONF# E(GFNn-xTB) ΔE(GFNn-xTB) E [Eh] Gsolv [Eh] GmRRHO [Eh] Gtot ΔGtot Boltzmannweight
[a.u.] [kcal/mol] r2scan-3c COSMO-RS-normal GFN2 [Eh] [kcal/mol] % at 298.15 K
[r2scan-3c] [alpb]-bhess
CONF2 -43.3959511 0.18 -687.0929614 -0.0279381 0.1487102 -686.9721893 0.71 3.05
CONF3 -43.3956417 0.37 -687.0936616 -0.0276693 0.1487424 -686.9725885 0.46 4.66
CONF4 -43.3942646 1.23 -687.0900815 -0.0308741 0.1484927 -686.9724630 0.54 4.08
CONF5 -43.3940567 1.36 -687.0914932 -0.0299652 0.1486902 -686.9727681 0.35 5.64
CONF6 -43.3926837 2.23 -687.0897209 -0.0295924 0.1487988 -686.9705145 1.76 0.52
CONF7 -43.3926044 2.28 -687.0877613 -0.0329647 0.1483288 -686.9723972 0.58 3.81
CONF8 -43.3924830 2.35 -687.0892849 -0.0316395 0.1480630 -686.9728613 0.29 6.22
CONF9 -43.3924281 2.39 -687.0899520 -0.0299305 0.1490469 -686.9708357 1.56 0.73
CONF10 -43.3923958 2.41 -687.0862984 -0.0305477 0.1476347 -686.9692114 2.58 0.13
CONF11 -43.3922081 2.52 -687.0891138 -0.0322739 0.1486419 -686.9727457 0.36 5.50
CONF12 -43.3921384 2.57 -687.0901510 -0.0309210 0.1483711 -686.9727010 0.39 5.25
CONF13 -43.3920387 2.63 -687.0885671 -0.0326800 0.1482019 -686.9730452 0.17 7.56
CONF14 -43.3919912 2.66 -687.0887067 -0.0330700 0.1484551 -686.9733216 0.00 10.13 <------
CONF15 -43.3917659 2.80 -687.0911883 -0.0297275 0.1486001 -686.9723157 0.63 3.49
CONF16 -43.3916896 2.85 -687.0874486 -0.0300377 0.1476961 -686.9697903 2.22 0.24
CONF17 -43.3914378 3.01 -687.0869188 -0.0339085 0.1478995 -686.9729279 0.25 6.68
CONF18 -43.3911600 3.18 -687.0854168 -0.0338099 0.1481027 -686.9711240 1.38 0.99
CONF19 -43.3910782 3.23 -687.0881927 -0.0314174 0.1487991 -686.9708109 1.58 0.71
CONF21 -43.3908014 3.41 -687.0874576 -0.0334708 0.1480488 -686.9728796 0.28 6.34
CONF25 -43.3905959 3.54 -687.0876706 -0.0320147 0.1486154 -686.9710699 1.41 0.93
CONF26 -43.3905265 3.58 -687.0885027 -0.0286509 0.1480364 -686.9691172 2.64 0.12
CONF27 -43.3904368 3.64 -687.0831797 -0.0377266 0.1478323 -686.9730740 0.16 7.79
CONF29 -43.3902113 3.78 -687.0881583 -0.0296301 0.1477586 -686.9700298 2.07 0.31
CONF31 -43.3895276 4.21 -687.0887446 -0.0319721 0.1481248 -686.9725918 0.46 4.68
CONF43 -43.3886315 4.77 -687.0844193 -0.0365084 0.1476077 -686.9733201 0.00 10.11
CONF46 -43.3883061 4.97 -687.0889402 -0.0288259 0.1480761 -686.9696901 2.28 0.22
CONF51 -43.3863091 6.23 -687.0865276 -0.0309995 0.1483822 -686.9691449 2.62 0.12
Number of conformers observed within the following ΔG windows:
ΔG [kcal/mol] #CONF sum(Boltzmann_weights)
---------------------------------------------
0 - 0.5 12 0.81
0 - 1.0 16 0.95
0 - 1.5 18 0.97
0 - 2.0 21 0.99
0 - 2.5 24 1.00
0 - 3.0 27 1.00
---------------------------------------------
Calculating ALPB_Gsolv values for evaluation of the std. dev. of Gsolv.
Constructed folders!
Starting 27 ALPB-Gsolv calculations
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF2/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF3/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF4/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF5/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF6/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF7/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF8/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF9/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF10/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF11/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF12/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF13/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF14/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF15/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF16/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF17/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF18/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF19/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF21/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF25/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF26/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF27/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF29/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF31/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF43/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF46/alpb_gsolv
Running ALPB_GSOLV calculation in 3762289.majestix.thch.uni-bonn.de/CONF51/alpb_gsolv
Tasks completed!
ALPB_GSOLV calculation was successful for CONF2/alpb_gsolv: -0.03045398
ALPB_GSOLV calculation was successful for CONF3/alpb_gsolv: -0.03035216
ALPB_GSOLV calculation was successful for CONF4/alpb_gsolv: -0.03005437
ALPB_GSOLV calculation was successful for CONF5/alpb_gsolv: -0.02980082
ALPB_GSOLV calculation was successful for CONF6/alpb_gsolv: -0.03061040
ALPB_GSOLV calculation was successful for CONF7/alpb_gsolv: -0.03186123
ALPB_GSOLV calculation was successful for CONF8/alpb_gsolv: -0.03170442
ALPB_GSOLV calculation was successful for CONF9/alpb_gsolv: -0.02979975
ALPB_GSOLV calculation was successful for CONF10/alpb_gsolv: -0.03097779
ALPB_GSOLV calculation was successful for CONF11/alpb_gsolv: -0.03076229
ALPB_GSOLV calculation was successful for CONF12/alpb_gsolv: -0.03260893
ALPB_GSOLV calculation was successful for CONF13/alpb_gsolv: -0.03196657
ALPB_GSOLV calculation was successful for CONF14/alpb_gsolv: -0.03131471
ALPB_GSOLV calculation was successful for CONF15/alpb_gsolv: -0.03159527
ALPB_GSOLV calculation was successful for CONF16/alpb_gsolv: -0.03076693
ALPB_GSOLV calculation was successful for CONF17/alpb_gsolv: -0.03234002
ALPB_GSOLV calculation was successful for CONF18/alpb_gsolv: -0.03099869
ALPB_GSOLV calculation was successful for CONF19/alpb_gsolv: -0.02908210
ALPB_GSOLV calculation was successful for CONF21/alpb_gsolv: -0.03210804
ALPB_GSOLV calculation was successful for CONF25/alpb_gsolv: -0.03159858
ALPB_GSOLV calculation was successful for CONF26/alpb_gsolv: -0.03027935
ALPB_GSOLV calculation was successful for CONF27/alpb_gsolv: -0.03249734
ALPB_GSOLV calculation was successful for CONF29/alpb_gsolv: -0.03186747
ALPB_GSOLV calculation was successful for CONF31/alpb_gsolv: -0.03218616
ALPB_GSOLV calculation was successful for CONF43/alpb_gsolv: -0.03280407
ALPB_GSOLV calculation was successful for CONF46/alpb_gsolv: -0.02931210
ALPB_GSOLV calculation was successful for CONF51/alpb_gsolv: -0.03040050
SD of solvation models @ 298.15 K (all units in kcal/mol):
CONFX ΔG(COSMO-RS) ΔG(DCOSMO-RS_gsolv) ΔG(ALPB_gsolv) SD(COSMO-RS 40%, DCOSMO-RS_gsolv 40%, ALPB_gsolv 20%)
----------------------------------------------------------------------------------------------------
CONF2 3.22 3.36 0.54 1.35
CONF3 3.39 3.62 0.60 1.43
CONF4 1.38 1.46 0.79 0.31
CONF5 1.95 2.08 0.95 0.53
CONF6 2.18 2.45 0.44 0.93
CONF7 0.07 -0.39 -0.34 0.26
CONF8 0.90 0.72 -0.24 0.52
CONF9 1.97 2.56 0.95 0.72
CONF10 1.58 1.16 0.21 0.61
CONF11 0.50 0.54 0.35 0.09
CONF12 1.35 1.32 -0.81 1.05
CONF13 0.24 -0.10 -0.41 0.30
CONF14 0.00 0.00 0.00 0.00
CONF15 2.10 1.72 -0.18 1.04
CONF16 1.90 1.51 0.34 0.70
CONF17 -0.53 -0.64 -0.64 0.07
CONF18 -0.46 -0.11 0.20 0.31
CONF19 1.04 1.38 1.40 0.21
CONF21 -0.25 -0.57 -0.50 0.18
CONF25 0.66 1.61 -0.18 0.83
CONF26 2.77 2.91 0.65 1.08
CONF27 -2.92 -3.65 -0.74 1.31
CONF29 2.16 2.26 -0.35 1.25
CONF31 0.69 0.38 -0.55 0.56
CONF43 -2.16 -2.89 -0.93 0.87
CONF46 2.66 2.77 1.26 0.72
CONF51 1.30 2.23 0.57 0.78
----------------------------------------------------------------------------------------------------
Calculating Boltzmann averaged free energy of ensemble!
temperature /K: avE(T) /a.u. avGmRRHO(T) /a.u. avGsolv(T) /a.u. avG(T) /a.u.
----------------------------------------------------------------------------------------------------
273.15 -687.0878885 0.1524606 -0.0360408 -686.9714687
278.15 -687.0879917 0.1516315 -0.0353266 -686.9716868
283.15 -687.0880896 0.1507948 -0.0346304 -686.9719252
288.15 -687.0881823 0.1499508 -0.0339519 -686.9721834
293.15 -687.0882696 0.1490998 -0.0332915 -686.9724613
298.15 -687.0883521 0.1482412 -0.0326487 -686.9727596 <<==part2==
303.15 -687.0884301 0.1473758 -0.0320225 -686.9730769
308.15 -687.0885036 0.1465028 -0.0314139 -686.9734147
313.15 -687.0885727 0.1456227 -0.0308226 -686.9737726
318.15 -687.0886381 0.1447356 -0.0302482 -686.9741507
323.15 -687.0886992 0.1438415 -0.0296910 -686.9745487
328.15 -687.0887571 0.1429403 -0.0291501 -686.9749669
333.15 -687.0888112 0.1420323 -0.0286259 -686.9754048
338.15 -687.0888618 0.1411171 -0.0281181 -686.9758629
343.15 -687.0889091 0.1401951 -0.0276264 -686.9763405
348.15 -687.0889532 0.1392660 -0.0271506 -686.9768378
353.15 -687.0889947 0.1383302 -0.0266899 -686.9773544
358.15 -687.0890331 0.1373874 -0.0262447 -686.9778904
363.15 -687.0890691 0.1364376 -0.0258139 -686.9784453
368.15 -687.0891027 0.1354814 -0.0253972 -686.9790185
373.15 -687.0891334 0.1345182 -0.0249950 -686.9796102
----------------------------------------------------------------------------------------------------
--------------------------------------------------
Conformers considered further
--------------------------------------------------
Conformers that are below the Boltzmann threshold G_thr(2) of 99.0%:
CONF14, CONF43, CONF27, CONF13, CONF17, CONF21, CONF8, CONF5, CONF11, CONF12, CONF31
CONF3, CONF4, CONF7, CONF15, CONF2, CONF18, CONF25, CONF9, CONF19, CONF6
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>END of Part2<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
Ran part2 in 6556.6225 seconds
----------------------------------------------------------------------------------------------------
OPTICAL ROTATION MODE - PART5
----------------------------------------------------------------------------------------------------
Part5: on
frequency in [nm]: 589.0
functional for SCF: r2scan-3c
functional for optical rotation: pbe-d4
basis set for optical rotation: def2-SVPD
Boltzmann sum threshold employed: 99.0
solvent: h2o
solvation model: cosmo
Considering the following 21 conformers:
CONF2, CONF3, CONF4, CONF5, CONF6, CONF7, CONF8, CONF9, CONF11, CONF12, CONF13
CONF14, CONF15, CONF17, CONF18, CONF19, CONF21, CONF25, CONF27, CONF31, CONF43
--------------------------------------------------
* Gibbs free energies used in part5 *
--------------------------------------------------
CONF# E [Eh] Gsolv [Eh] GmRRHO [Eh] Gtot ΔGtot Boltzmannweight
r2scan-3c COSMO-RS-normal GFN2 [Eh] [kcal/mol] % at 298.15 K
[r2scan-3c] [alpb]-bhess
CONF2 -687.0929614 -0.0279381 0.1487102 -686.9721893 0.71 3.09
CONF3 -687.0936616 -0.0276693 0.1487424 -686.9725885 0.46 4.71
CONF4 -687.0900815 -0.0308741 0.1484927 -686.9724630 0.54 4.13
CONF5 -687.0914932 -0.0299652 0.1486902 -686.9727681 0.35 5.70
CONF6 -687.0897209 -0.0295924 0.1487988 -686.9705145 1.76 0.52
CONF7 -687.0877613 -0.0329647 0.1483288 -686.9723972 0.58 3.85
CONF8 -687.0892849 -0.0316395 0.1480630 -686.9728613 0.29 6.29
CONF9 -687.0899520 -0.0299305 0.1490469 -686.9708357 1.56 0.74
CONF11 -687.0891138 -0.0322739 0.1486419 -686.9727457 0.36 5.57
CONF12 -687.0901510 -0.0309210 0.1483711 -686.9727010 0.39 5.31
CONF13 -687.0885671 -0.0326800 0.1482019 -686.9730452 0.17 7.65
CONF14 -687.0887067 -0.0330700 0.1484551 -686.9733216 0.00 10.25 <------
CONF15 -687.0911883 -0.0297275 0.1486001 -686.9723157 0.63 3.53
CONF17 -687.0869188 -0.0339085 0.1478995 -686.9729279 0.25 6.75
CONF18 -687.0854168 -0.0338099 0.1481027 -686.9711240 1.38 1.00
CONF19 -687.0881927 -0.0314174 0.1487991 -686.9708109 1.58 0.72
CONF21 -687.0874576 -0.0334708 0.1480488 -686.9728796 0.28 6.42
CONF25 -687.0876706 -0.0320147 0.1486154 -686.9710699 1.41 0.94
CONF27 -687.0831797 -0.0377266 0.1478323 -686.9730740 0.16 7.88
CONF31 -687.0887446 -0.0319721 0.1481248 -686.9725918 0.46 4.73
CONF43 -687.0844193 -0.0365084 0.1476077 -686.9733201 0.00 10.23
Conformers that are below the Boltzmann-thr of 99.0:
CONF14, CONF43, CONF27, CONF13, CONF17, CONF21, CONF8, CONF5, CONF11, CONF12, CONF31
CONF3, CONF4, CONF7, CONF15, CONF2, CONF18, CONF25, CONF9, CONF19, CONF6
Constructed folders!
Optical-rotation is calculated at pbe-d4/def2-SVPD[COSMO]_[SCF=r2scan/def2-SVPD](tm) level.
Starting 21 optical-rotation calculations.
Running single-point in CONF43/OR
Running single-point in CONF14/OR
Running single-point in CONF27/OR
Running single-point in CONF13/OR
Running single-point in CONF21/OR
Running single-point in CONF17/OR
Running single-point in CONF8/OR
Running optical-rotation calculation in CONF27/OR
Running optical-rotation calculation in CONF8/OR
Running optical-rotation calculation in CONF43/OR
Running optical-rotation calculation in CONF17/OR
Running optical-rotation calculation in CONF21/OR
Running optical-rotation calculation in CONF13/OR
Running optical-rotation calculation in CONF14/OR
Running single-point in CONF5/OR
Running single-point in CONF11/OR
Running single-point in CONF12/OR
Running single-point in CONF31/OR
Running single-point in CONF3/OR
Running single-point in CONF4/OR
Running single-point in CONF7/OR
Running optical-rotation calculation in CONF11/OR
Running optical-rotation calculation in CONF12/OR
Running optical-rotation calculation in CONF5/OR
Running optical-rotation calculation in CONF31/OR
Running optical-rotation calculation in CONF3/OR
Running optical-rotation calculation in CONF7/OR
Running optical-rotation calculation in CONF4/OR
Running single-point in CONF15/OR
Running single-point in CONF2/OR
Running single-point in CONF18/OR
Running single-point in CONF25/OR
Running single-point in CONF9/OR
Running single-point in CONF19/OR
Running single-point in CONF6/OR
Running optical-rotation calculation in CONF15/OR
Running optical-rotation calculation in CONF2/OR
Running optical-rotation calculation in CONF18/OR
Running optical-rotation calculation in CONF25/OR
Running optical-rotation calculation in CONF9/OR
Running optical-rotation calculation in CONF19/OR
Running optical-rotation calculation in CONF6/OR
Tasks completed!
Optical-rotation calculation was successful for CONF2/OR at 589.0 nm: 102.7973792 populated to 3.09 %
Optical-rotation calculation was successful for CONF3/OR at 589.0 nm: 106.2937684 populated to 4.71 %
Optical-rotation calculation was successful for CONF4/OR at 589.0 nm: 111.1385990 populated to 4.13 %
Optical-rotation calculation was successful for CONF5/OR at 589.0 nm: 116.9873437 populated to 5.70 %
Optical-rotation calculation was successful for CONF6/OR at 589.0 nm: 43.9366079 populated to 0.52 %
Optical-rotation calculation was successful for CONF7/OR at 589.0 nm: 103.4236640 populated to 3.85 %
Optical-rotation calculation was successful for CONF8/OR at 589.0 nm: 91.2248593 populated to 6.29 %
Optical-rotation calculation was successful for CONF9/OR at 589.0 nm: 58.8846581 populated to 0.74 %
Optical-rotation calculation was successful for CONF11/OR at 589.0 nm: 116.0659851 populated to 5.57 %
Optical-rotation calculation was successful for CONF12/OR at 589.0 nm: 125.5155013 populated to 5.31 %
Optical-rotation calculation was successful for CONF13/OR at 589.0 nm: 111.9494751 populated to 7.65 %
Optical-rotation calculation was successful for CONF14/OR at 589.0 nm: 95.8030729 populated to 10.25 %
Optical-rotation calculation was successful for CONF15/OR at 589.0 nm: 99.9359637 populated to 3.53 %
Optical-rotation calculation was successful for CONF17/OR at 589.0 nm: 94.2968263 populated to 6.75 %
Optical-rotation calculation was successful for CONF18/OR at 589.0 nm: 65.0637755 populated to 1.00 %
Optical-rotation calculation was successful for CONF19/OR at 589.0 nm: 87.2072536 populated to 0.72 %
Optical-rotation calculation was successful for CONF21/OR at 589.0 nm: 105.8620382 populated to 6.42 %
Optical-rotation calculation was successful for CONF25/OR at 589.0 nm: 56.2008664 populated to 0.94 %
Optical-rotation calculation was successful for CONF27/OR at 589.0 nm: 85.5830106 populated to 7.88 %
Optical-rotation calculation was successful for CONF31/OR at 589.0 nm: 89.2702544 populated to 4.73 %
Optical-rotation calculation was successful for CONF43/OR at 589.0 nm: 79.3215642 populated to 10.23 %
Averaged specific rotation at 589.0 nm : 98.902 in deg*[dm(g/cc)]^(-1)
SD based on SD of Gsolv (part2) : 6.464 in deg*[dm(g/cc)]^(-1)
SD based on SD in G of 0.4 kcal/mol: 1.145 in deg*[dm(g/cc)]^(-1)
frequency min(OR) CONF max(OR) CONF
--------------------------------------------------
589.0 43.94 CONF6 125.52 CONF12
--------------------------------------------------
* min and max values are not Boltzmann weighted.
>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>END of Part5<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
Ran part5 in 307.7719 seconds
Part : #conf time time (including restarts)
-----------------------------------------------------------------------
Input : 58 - -
Part0_all : 32 42.12 s 42.12 s
Part1_initial_sort : 29 207.00 s 207.00 s
Part1_all : 29 214.08 s 214.08 s
Part2_opt : 27 6374.21 s 6374.21 s
Part2_all : 21 6556.62 s 6556.62 s
Part5 : 21 307.77 s 307.77 s
-----------------------------------------------------------------------
All parts : - 7120.59 s 7120.59 s
CENSO all done!
$CENSO global configuration file: .censorc
$VERSION:1.1.2
ORCA: /home/$USER/orca_5_0_1_linux_x86-64_openmpi411
ORCA version: 5.0.1
GFN-xTB: /home/$USER/bin/xtb
CREST: /home/$USER/bin/crest
mpshift: /home/$USER/TURBOMOLE.7.5/bin/em64t-unknown-linux-gnu/mpshift
escf: /home/$USER/TURBOMOLE.7.5/bin/em64t-unknown-linux-gnu/escf
#COSMO-RS
ctd = BP_TZVP_C30_1601.ctd cdir = "/home/$USER/COSMOthermX16/COSMOtherm/CTDATA-FILES" ldir = "/home/$USER/COSMOthermX16/COSMOtherm/CTDATA-FILES"
$ENDPROGRAMS
$CRE SORTING SETTINGS:
$GENERAL SETTINGS:
nconf: all # ['all', 'number e.g. 10 up to all conformers']
charge: 0 # ['number e.g. 0']
unpaired: 0 # ['number e.g. 0']
solvent: gas # ['gas', 'acetone', 'acetonitrile', 'aniline', 'benzaldehyde', 'benzene', 'ccl4', '...']
prog_rrho: xtb # ['xtb']
temperature: 298.15 # ['temperature in K e.g. 298.15']
trange: [273.15, 378.15, 5] # ['temperature range [start, end, step]']
multitemp: on # ['on', 'off']
evaluate_rrho: on # ['on', 'off']
consider_sym: on # ['on', 'off']
bhess: on # ['on', 'off']
imagthr: automatic # ['automatic or e.g., -100 # in cm-1']
sthr: automatic # ['automatic or e.g., 50 # in cm-1']
scale: automatic # ['automatic or e.g., 1.0 ']
rmsdbias: off # ['on', 'off']
sm_rrho: alpb # ['alpb', 'gbsa']
progress: off # possibilities
check: on # ['on', 'off']
prog: tm # ['tm', 'orca']
func: r2scan-3c # ['b3-lyp', 'b3lyp', 'b3lyp-3c', 'b3lyp-d3', 'b3lyp-d3(0)', 'b3lyp-d4', 'b3lyp-nl', '...']
basis: automatic # ['automatic', 'def2-TZVP', 'def2-mSVP', 'def2-mSVP', 'def2-mSVP', 'def2-mSVP', '...']
maxthreads: 7 # ['number of threads e.g. 2']
omp: 4 # ['number cores per thread e.g. 4']
balance: off # possibilities
cosmorsparam: automatic # ['automatic', '12-fine', '12-normal', '13-fine', '13-normal', '14-fine', '...']
$PART0 - CHEAP-PRESCREENING - SETTINGS:
part0: on # ['on', 'off']
func0: b97-d # ['b3-lyp', 'b3lyp', 'b3lyp-3c', 'b3lyp-d3', 'b3lyp-d3(0)', 'b3lyp-d4', '...']
basis0: def2-SV(P) # ['automatic', 'def2-SV(P)', 'def2-TZVP', 'def2-mSVP', 'def2-mSVP', 'def2-mSVP', '...']
part0_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
part0_threshold: 4.0 # ['number e.g. 4.0']
$PART1 - PRESCREENING - SETTINGS:
# func and basis is set under GENERAL SETTINGS
part1: on # ['on', 'off']
smgsolv1: cosmors # ['alpb_gsolv', 'cosmo', 'cosmors', 'cosmors-fine', 'cpcm', 'dcosmors', '...']
part1_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
part1_threshold: 3.5 # ['number e.g. 5.0']
$PART2 - OPTIMIZATION - SETTINGS:
# func and basis is set under GENERAL SETTINGS
part2: on # ['on', 'off']
opt_limit: 2.5 # ['number e.g. 4.0']
sm2: default # ['cosmo', 'cpcm', 'dcosmors', 'default', 'smd']
smgsolv2: cosmors # ['alpb_gsolv', 'cosmo', 'cosmors', 'cosmors-fine', 'cpcm', 'dcosmors', '...']
part2_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
ancopt: on # ['on']
hlow: 0.01 # ['lowest force constant in ANC generation, e.g. 0.01']
opt_spearman: on # ['on', 'off']
part2_threshold: 99 # ['Boltzmann sum threshold in %. e.g. 95 (between 1 and 100)']
optlevel2: automatic # ['crude', 'sloppy', 'loose', 'lax', 'normal', 'tight', 'vtight', 'extreme', '...']
optcycles: 8 # ['number e.g. 5 or 10']
spearmanthr: -4.0 # ['value between -1 and 1, if outside set automatically']
radsize: 10 # ['number e.g. 8 or 10']
crestcheck: off # ['on', 'off']
$PART3 - REFINEMENT - SETTINGS:
part3: off # ['on', 'off']
prog3: prog # ['tm', 'orca', 'prog']
func3: pw6b95 # ['b3-lyp', 'b3lyp', 'b3lyp-3c', 'b3lyp-d3', 'b3lyp-d3(0)', 'b3lyp-d4', 'b3lyp-nl', '...']
basis3: def2-TZVPD # ['DZ', 'QZV', 'QZVP', 'QZVPP', 'SV(P)', 'SVP', 'TZVP', 'TZVPP', 'aug-cc-pV5Z', '...']
smgsolv3: cosmors # ['alpb_gsolv', 'cosmo', 'cosmors', 'cosmors-fine', 'cpcm', 'dcosmors', '...']
part3_gfnv: gfn2 # ['gfn1', 'gfn2', 'gfnff']
part3_threshold: 99 # ['Boltzmann sum threshold in %. e.g. 95 (between 1 and 100)']
$NMR PROPERTY SETTINGS:
$PART4 SETTINGS:
part4: off # ['on', 'off']
couplings: on # ['on', 'off']
progJ: prog # ['tm', 'orca', 'adf', 'prog']
funcJ: pbe0 # ['b3-lyp', 'b3lyp', 'b3lyp-3c', 'b3lyp-d3', 'b3lyp-d3(0)', 'b3lyp-d4', 'b3lyp-nl', '...']
basisJ: def2-TZVP # ['DZ', 'QZV', 'QZVP', 'QZVPP', 'SV(P)', 'SVP', 'TZVP', 'TZVPP', 'aug-cc-pV5Z', '...']
sm4J: default # ['cosmo', 'cpcm', 'dcosmors', 'smd']
shieldings: on # ['on', 'off']
progS: prog # ['tm', 'orca', 'adf', 'prog']
funcS: pbe0 # ['b3-lyp', 'b3lyp', 'b3lyp-3c', 'b3lyp-d3', 'b3lyp-d3(0)', 'b3lyp-d4', 'b3lyp-nl', '...']
basisS: def2-TZVP # ['DZ', 'QZV', 'QZVP', 'QZVPP', 'SV(P)', 'SVP', 'TZVP', 'TZVPP', 'aug-cc-pV5Z', '...']
sm4S: default # ['cosmo', 'cpcm', 'dcosmors', 'smd']
reference_1H: TMS # ['TMS']
reference_13C: TMS # ['TMS']
reference_19F: CFCl3 # ['CFCl3']
reference_29Si: TMS # ['TMS']
reference_31P: TMP # ['TMP', 'PH3']
1H_active: on # ['on', 'off']
13C_active: on # ['on', 'off']
19F_active: off # ['on', 'off']
29Si_active: off # ['on', 'off']
31P_active: off # ['on', 'off']
resonance_frequency: 300.0 # ['MHz number of your experimental spectrometer setup']
$OPTICAL ROTATION PROPERTY SETTINGS:
$PART5 SETTINGS:
optical_rotation: off # ['on', 'off']
funcOR: pbe # ['functional for opt_rot e.g. pbe']
funcOR_SCF: r2scan-3c # ['functional for SCF in opt_rot e.g. r2scan-3c']
basisOR: def2-SVPD # ['basis set for opt_rot e.g. def2-SVPD']
frequency_optical_rot: [589.0] # ['list of freq in nm to evaluate opt rot at e.g. [589, 700]']
$END CENSORC
#label #unmod_alpha #%pop #pop_alpha
CONF2 102.7973792 3.09 3.1746247
CONF3 106.2937684 4.71 5.0098495
CONF4 111.1385990 4.13 4.5864760
CONF5 116.9873437 5.70 6.6696538
CONF6 43.9366079 0.52 0.2302506
CONF7 103.4236640 3.85 3.9809478
CONF8 91.2248593 6.29 5.7404428
CONF9 58.8846581 0.74 0.4335951
CONF11 116.0659851 5.57 6.4618040
CONF12 125.5155013 5.31 6.6645412
CONF13 111.9494751 7.65 8.5591455
CONF14 95.8030729 10.25 9.8160203
CONF15 99.9359637 3.53 3.5283330
CONF17 94.2968263 6.75 6.3671226
CONF18 65.0637755 1.00 0.6502028
CONF19 87.2072536 0.72 0.6255631
CONF21 105.8620382 6.42 6.7913141
CONF25 56.2008664 0.94 0.5303495
CONF27 85.5830106 7.88 6.7461188
CONF31 89.2702544 4.73 4.2223559
CONF43 79.3215642 10.23 8.1137383
ENSEMBLE - 100.00 98.9024494
#label #unmod_alpha #%pop #pop_alpha
CONF18 62.0011571 1.20 0.7415879
CONF37 77.3646553 8.16 6.3099117
CONF2 104.2469597 1.97 2.0581874
CONF3 106.1874492 4.56 4.8374369
CONF4 112.4111334 3.78 4.2486734
CONF5 111.6304190 3.43 3.8342079
CONF7 104.7549480 3.59 3.7561630
CONF8 91.8417680 5.94 5.4558852
CONF10 122.6012719 2.44 2.9913932
CONF11 115.8694122 5.27 6.1059079
CONF12 112.9598654 7.34 8.2949892
CONF13 96.7248433 10.52 10.1792607
CONF14 103.5470026 3.94 4.0822709
CONF15 112.1247398 6.01 6.7386684
CONF17 95.6710169 4.46 4.2714917
CONF22 106.9330550 15.21 16.2623453
CONF23 106.7655902 6.78 7.2357336
CONF28 86.4144602 5.40 4.6648660
ENSEMBLE - 100.00 102.0689805
#label #unmod_alpha #%pop #pop_alpha
CONF2 104.9669754 2.80 2.9419140
CONF3 106.0120235 5.30 5.6222722
CONF4 110.9329463 3.93 4.3572913
CONF6 104.4255264 4.29 4.4798153
CONF7 93.5973398 5.00 4.6773696
CONF8 58.3051609 0.76 0.4441435
CONF9 122.1148522 3.79 4.6283274
CONF10 116.2003884 6.79 7.8874111
CONF11 127.1443760 5.17 6.5735045
CONF12 110.2302123 9.07 9.9979829
CONF13 96.0502078 11.63 11.1667826
CONF14 101.3493071 4.20 4.2537923
CONF15 137.1625824 17.03 23.3628110
CONF16 93.9188928 7.22 6.7768989
CONF17 39.3066203 1.39 0.5459943
CONF18 63.6041283 1.11 0.7040302
CONF19 85.7532225 0.68 0.5855706
CONF36 76.9580458 9.85 7.5790273
ENSEMBLE - 100.00 106.5849390
The mean value is 102.5 °[dm(g/cm³)]¹ and in good agreement with the experimental value of 112.2 °[dm(g/cm³)]¹.
The same procedure is performed for \({\beta}\)-D-glucopyranose:
#label #unmod_alpha #%pop #pop_alpha
CONF36 71.8436660 6.91 4.9674147
CONF61 -33.7762603 1.43 -0.4833508
CONF62 -73.4341103 1.97 -1.4430452
CONF2 14.5921787 5.71 0.8326308
CONF3 -2.2178727 8.06 -0.1787735
CONF8 39.6099777 1.52 0.6033281
CONF10 20.4043013 9.21 1.8798166
CONF18 44.1015730 1.36 0.5976118
CONF19 24.0564442 8.31 1.9979496
CONF22 -27.4858229 6.61 -1.8161625
CONF32 32.5573942 2.09 0.6810443
CONF33 -22.3070859 8.96 -1.9986714
CONF35 -31.3889925 1.17 -0.3676520
CONF38 -1.7718248 14.31 -0.2535221
CONF45 17.7717916 15.84 2.8149573
CONF47 -17.4698219 1.23 -0.2150347
CONF51 -20.7939267 5.32 -1.1056746
ENSEMBLE - 100.00 6.5128666
#label #unmod_alpha #%pop #pop_alpha
CONF2 11.3134566 8.16 0.9234435
CONF3 -2.9269891 13.22 -0.3870561
CONF15 42.3742219 2.58 1.0931013
CONF19 -28.2196786 12.37 -3.4905616
CONF27 44.0426779 4.36 1.9194166
CONF28 -5.3044713 1.32 -0.0702160
CONF30 -31.1999822 2.02 -0.6293121
CONF31 -8.2853209 10.51 -0.8710484
CONF33 -44.3695905 1.48 -0.6579142
CONF38 11.0792758 31.38 3.4768954
CONF39 48.1934298 10.54 5.0782686
CONF49 22.2853984 2.05 0.4570863
ENSEMBLE - 100.00 6.8421032
#label #unmod_alpha #%pop #pop_alpha
CONF2 12.2430597 3.75 0.4589710
CONF3 -2.8935102 6.77 -0.1959881
CONF8 39.9385168 1.37 0.5470013
CONF10 16.7667327 7.85 1.3161104
CONF17 42.6955530 1.30 0.5534156
CONF18 8.6413723 14.85 1.2836111
CONF20 -32.9571222 6.69 -2.2044944
CONF21 29.8935830 4.50 1.3441909
CONF27 43.5091601 1.92 0.8336047
CONF29 -22.5893260 8.04 -1.8168235
CONF32 -32.7355186 1.03 -0.3377336
CONF33 70.4098794 6.26 4.4050269
CONF35 31.0303288 5.07 1.5737914
CONF37 -6.6000456 9.34 -0.6161968
CONF45 -11.3040279 7.47 -0.8443406
CONF51 -12.7692834 5.03 -0.6419296
CONF58 21.9406646 1.09 0.2395658
CONF59 -7.6250365 7.68 -0.5855640
ENSEMBLE - 100.00 5.3122187
The mean value is 6.2 °[dm(g/cm³)]¹ and in good agreement with the experimental value of 17.5°[dm(g/cm³)]¹. \({\alpha}\) and \({\beta}\) forms of D-glucopyranose could clearly be identified.
Hint
By default, CENSO uses the gauge-invariant velocity representation for the calculation of the OR. This option is currently not available for hybrid DFAs in TURBOMOLE and the lenght representation is used instead when using a hybrid DFA for the calculation of the OR.
Restarting calculations
CENSO keeps track of all performed calculations, if they succeed, fail or are pending. The data is stored in enso.json and enables restarting of CENSO runs. Restarting a calculation is useful if you only considered few conformers in the first run and want to increase the number of investigated conformers (e.g., start from 30 and go to 100 conformers). In the case of difficult molecules (e.g., transition metal compounds) where large differences between the low-cost methods in the sorting parts and the high-level method in part 2 are found, the default thresholds may be increased. Or if after a previous calculation of a free ensemble energy, a property, e.g. OR, is to be calculated. This can be done by first calculating the free energy, checking the results and the final ensemble and then restart to calculate the property (OR). Some restart limitations are given though: You can not change anything that concerns the geometry optimization, since all results have to be created with the same method-combination.
When restarting CENSO, files with information of the previous calculation (enso.json, enso_ensemble_partx.xyz, …) are automatically backed up.
Restarting a calculation is performed by calling censo and adjusting the new settings by command line:
# increase the number of considered conformers to 200
$ censo -restart -nc 200 > censo-restart.out &
# increase the thresholds for parts 0 and 1
# if relevant conformers have been sorted out
$ censo --restart -thrpart0 5.0 -thrpart1 3.0 > censo_restart.out
# calculate OR after previous free energy calculation
$ censo -restart -OR on > censo-restart-OR.out &